Abstract
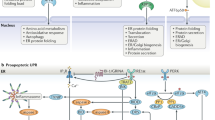
The ultimate mechanism that cells use to ensure the quality of intracellular proteins is the selective destruction of misfolded or damaged polypeptides. In eukaryotic cells, the large ATP-dependent proteolytic machine, the 26S proteasome, prevents the accumulation of non-functional, potentially toxic proteins. This process is of particular importance in protecting cells against harsh conditions (for example, heat shock or oxidative stress) and in a variety of diseases (for example, cystic fibrosis and the major neurodegenerative diseases). A full understanding of the pathogenesis of the protein-folding diseases will require greater knowledge of how misfolded proteins are recognized and selectively degraded.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Glickman, M. H. & Ciechanover, A. The ubiquitin–proteasome proteolytic pathway: Destruction for the sake of construction. Physiol. Rev. 82, 373–428 (2002).
Goldberg, A. L. & Dice, J. F. Intracellular protein degradation in mammalian and bacterial cells. Annu. Rev. Biochem. 43, 835–869 (1974).
Sherman, M. & Goldberg, A. L. Cellular defenses against unfolded proteins: a cell biologist thinks about neurodegenerative diseases. Neuron 29, 15–32 (2001).
Goldberg, A. L. Degradation of abnormal proteins in E. coli. Proc. Natl Acad. Sci. USA 69, 422–426 (1972).
Zwickl, P., Goldberg, A. L. & Baumeister, W. in Proteasomes: The World of Regulatory Proteolysis (eds Wolf, D. H. & Hilt, W.) 8–20 (Landes Bioscience, Georgetown, Texas, 2000).
Etlinger, J. & Goldberg, A. L. A soluble, ATP-dependent proteolytic system responsible for the degradation of abnormal proteins in reticulocytes. Proc. Natl Acad. Sci. USA. 74, 54–58 (1977).
Klemes, Y., Etlinger, J. D. & Goldberg, A. L. Properties of proteins degraded rapidly in reticulocytes: intracellular aggregation of the globin molecules prior to hydrolysis. J. Biol. Chem. 256, 8436–8444 (1981).
Bunn, H. F. et al. Hemoglobin: Molecular, Genetic, and Clinical Aspects (Saunders, Philadelphia, 1986).
Goldberg, A. L. & Goff, S. A. in Maximizing Gene Expression Ch. 9 (eds Reznikoff, W. & Gold, L.) 287–314 (Butterworths, Stoneham, Massachusetts, 1986).
Kostova, Z. & Wolf, D. H. For whom the bell tolls: protein quality control of the endoplasmic reticulum and the ubiquitin–proteasome connection. EMBO J. 22, 2309–2317 (2003).
Goff, S. A., Voellmy, R. & Goldberg, A. L. in Ubiquitin Ch. 8 (ed. Rechsteiner, M.) 207–238 (Plenum, New York, 1988).
Roche, E. & Sauer, R. T. SsrA-mediated peptide tagging caused by rare codons and tRNA scarcity. EMBO J. 18, 4579–4589 (1999).
Frydman, J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu. Rev. Biochem. 70, 603–647 (2001).
Hartl, F. U. & Mayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Schubert, U. et al. Rapid degradation of a large fraction of newly synthesized proteins by proteasomes. Nature 44, 770–774 (2000).
Gronostajski, R., Pardee, A. B. & Goldberg, A. L. The ATP-dependence of the degradation of short- and long-lived proteins in growing fibroblasts. J. Biol. Chem. 260, 3344–3349 (1985).
Berlett, B. S. & Stadtman, E. R. Protein oxidation in aging, disease, and oxidative stress. J. Biol. Chem. 272, 20313–20316 (1997).
Grune, T., Reinheckel, T., Davies, K. J. Degradation of oxidized proteins in mammalian cells. FASEB J. 11, 536–534 (1997).
Tarcsa, E., Szymanska, G., Lecker, S., O'Connor, C. M. & Goldberg, A. L. Ca2+-free calmodulin and calmodulin damaged by in vitro aging are selectively degraded by 26S proteasomes without ubiquitylation. J. Biol. Chem. 275, 20295–20301 (2000).
Prouty, W. F. & Goldberg, A. L. Fate of abnormal proteins in E. coli: accumulation in intracellular granules before catabolism. Nature New Biol. 240, 147–150 (1972).
Glover, J. R. & Lindquist, S. Hsp104, Hsp70, and Hsp40: A novel chaperone system that rescues previously aggregated proteins. Cell 94, 73–82 (1998).
Johnston, J. A., Ward, C. L. & Kopito, R. R. Aggresomes: a cellular response to misfolded proteins. J. Cell Biol. 1998. 143, 1883–1898.
Yamamoto, A., Lucas, J. J. & Hen, R. Reversal of neuropathology and motor dysfunction in a conditional model of Huntington's disease. Cell 101, 57–66 (2000).
Ciechanover, A., Heller, H., Elias, S., Haas, A. L. & Hershko A. ATP-dependent conjugation of reticulocyte proteins with the polypeptide required for protein degradation. Proc. Natl Acad. Sci. USA 77, 1365–1368 (1980).
Hough, R., Pratt, G. & Rechsteiner, M. Purification of two high molecular weight proteases from rabbit reticulocyte lysate. J. Biol. Chem. 262, 8303–8313 (1987).
Waxman, L., Fagan, J. M. & Goldberg, A. L. Demonstration of two distinct high molecular weight proteases in rabbit reticulocytes, one of which degrades ubiquitin conjugates. J. Biol. Chem. 262, 2451–2457 (1987).
Voges, D., Zwickl, P., Baumeister, W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (2000).
Ananthan, J., Goldberg, A. L. & Voellmy, R. Abnormal proteins serve as eukaryotic stress signals and trigger the activation of heat-shock genes. Science 232, 522–524 (1986).
Meacham, G. C., Patterson, C., Zhang, W., Younger, J. M. & Cyr, D. M. The Hsc70 co-chaperone CHIP targets immature CFTR for proteasomal degradation. Nature Cell Biol. 3, 100–105 (2001).
Murata, S., Minami, Y., Minami, M., Chiba, T. & Tanaka, K. CHIP is a chaperone-dependent E3 ligase that ubiquitylates unfolded protein. EMBO Rep. 2, 1133–1138. (2001).
Wickner, S., Maurizi, M. & Gottesman, S. Posttranslational quality control: folding, refolding, and degrading proteins. Science 286, 1888–1893 (1999).
Huang, H.C., Sherman, M. Y., Kandror, O. & Goldberg, A. L. The molecular chaperone DnaJ is required for the degradation of a soluble abnormal protein in E. coli. J. Biol. Chem. 276, 3920–3928 (2001).
Goldberg, A. L. The mechanisms and functions of ATP-dependent proteases in bacterial and animal cells. Eur. J. Biochem. 203, 9–23 (1992).
Chung, C. H. Proteases in Escherichia coli. Science 262, 372–374 (1993).
Maurizi, M. R. Proteases and protein degradation in Escherichia coli. Experientia 48, 178–201 (1992).
Langer, T., Kaser, M., Klanner, C., Leonhard, K. AAA proteases of mitochondria: quality control of membrane proteins and regulatory functions during mitochondrial biogenesis. Biochem. Soc. Trans. 29, 431–436 (2001).
Suzuki, C. K., Rep, M., van Dijl, J. M., Suda, K., Grivell, L. A., Schatz, G. ATP-dependent proteases that also chaperone protein biogenesis. Trends Biochem. Sci. 22, 118–123 (1997).
Kandror, O., Sherman, M. Y. & Goldberg, A. L. Rapid degradation of an abnormal protein in E. coli proceeds through repeated cycles of association with GroEL. J. Biol. Chem. 274, 37743–37749 (1999).
Groll, M. et al. A gated channel into the proteasome core particle. Nature Struct. Biol. 7, 1062–1067 (2000).
Benaroudj, N., Zwickl, P., Seemüller, E., Baumeister, W. & Goldberg, A. L. ATP hydrolysis by the proteasome regulatory complex PAN serves multiple functions in protein degradation. Mol. Cell 11, 69–78 (2003).
Lee, D. H. & Goldberg, A. L. Proteasome inhibitors cause rapid induction of heat-shock proteins and trehalose, which together confer thermotolerance in Saccharomyces cerevisiae. Mol. Cell. Biol. 18, 30–38 (1998).
Kisselev, A. F. & Goldberg, A. L. Proteasome inhibitors: from research tools to drug candidates. Chemy Biol. 8, 739–758 (2001).
Meiners, S. et al. Inhibition of proteasome activity induces concerted expression of proteasome genes and de novo formation of mammalian proteasomes. J. Biol. Chem. 278, 21517–21525 (2003).
Meriin, A. B. et al. Protein-damaging stresses activate c-Jun N-terminal kinase via inhibition of its dephosphorylation: a novel pathway controlled by HSP72. Mol. Cell. Biol. 19, 2547–2555 (1999).
Hideshima, T. et al. The proteasome inhibitor pS-341 inhibits growth, induces apoptosis, and overcomes drug resistance in human multiple myeloma cells. Cancer Res. 61, 3071–3076 (2001).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Goldberg, A. Protein degradation and protection against misfolded or damaged proteins. Nature 426, 895–899 (2003). https://doi.org/10.1038/nature02263
Issue Date:
DOI: https://doi.org/10.1038/nature02263
This article is cited by
-
Key role of K+ and Ca2+ in high-yield ethanol production by S. Cerevisiae from concentrated sugarcane molasses
Microbial Cell Factories (2024)
-
Post-ischemic ubiquitination at the postsynaptic density reversibly influences the activity of ischemia-relevant kinases
Communications Biology (2024)
-
Systematic identification of 20S proteasome substrates
Molecular Systems Biology (2024)
-
MiR-383 sensitizes osteosarcoma cells to bortezomib treatment via down-regulating PSMB5
Molecular Biology Reports (2024)
-
An overview on glycation: molecular mechanisms, impact on proteins, pathogenesis, and inhibition
Biophysical Reviews (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.