Abstract
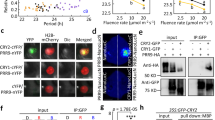
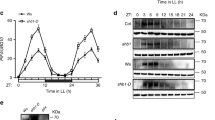
Adaptation to seasonal change is a crucial component of an organism's survival strategy. To monitor seasonal variation, organisms have developed the capacity to measure day length (photoperiodism). Day-length assessment involves the photoperiodic control of flowering in Arabidopsis thaliana, whereby the coincidence of light and high expression of CONSTANS (CO) induces the expression of FLOWERING LOCUS T (FT), leading to flowering in long-day conditions1. Although controlling CO expression is clearly a key step in day-length discrimination, the mechanism that generates day-length-dependent CO expression remains unknown. Here we show that the clock-controlled FLAVIN-BINDING, KELCH REPEAT, F-BOX (FKF1)2 protein has an essential role in generating the diurnal CO peak and that this function is dependent on light. We show that a recombinant FKF1 LIGHT, OXYGEN OR VOLTAGE (LOV)3 domain binds the chromophore flavin mononucleotide and undergoes light-induced photochemistry, indicating that FKF1 may function as a photoperiodic blue-light receptor. It is likely that the circadian control of FKF1 expression and the light regulation of FKF1 function coincide to control the daytime CO waveform precisely, which in turn is crucial for day-length discrimination by Arabidopsis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yanovsky, M. J. & Kay, S. A. Molecular basis of seasonal time measurement in Arabidopsis. Nature 419, 308–312 (2002)
Nelson, D. C., Lasswell, J., Rogg, L. E., Cohen, M. A. & Bartel, B. FKF1, a clock-controlled gene that regulates the transition to flowering in Arabidopsis. Cell 101, 331–340 (2000)
Christie, J. M., Salomon, M., Nozue, K., Wada, M. & Briggs, W. R. LOV (light, oxygen, or voltage) domains of the blue-light photoreceptor phototropin (nph1): binding sites for the chromophore flavin mononucleotide. Proc. Natl Acad. Sci. USA 96, 8779–8783 (1999)
Yanovsky, M. J. & Kay, S. A. Living by the calendar: how plants know when to flower. Nature Rev. Mol. Cell Biol. 4, 265–275 (2003)
Somers, D. E., Schultz, T. F., Milnamow, M. & Kay, S. A. ZEITLUPE encodes a novel clock-associated PAS protein from Arabidopsis. Cell 101, 319–329 (2000)
Schultz, T. F., Kiyosue, T., Yanovsky, M., Wada, M. & Kay, S. A. A role for LKP2 in the circadian clock of Arabidopsis. Plant Cell 13, 2659–2670 (2001)
Millar, A. J., Short, S. R., Chua, N. H. & Kay, S. A. A novel circadian phenotype based on firefly luciferase expression in transgenic plants. Plant Cell 4, 1075–1087 (1992)
Strayer, C. et al. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science 289, 768–771 (2000)
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotechnol 17, 1030–1032 (1999)
Harmer, S. L. et al. Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290, 2110–2113 (2000)
Alabadi, D. et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880–883 (2001)
Alabadi, D., Yanovsky, M. J., Mas, P., Harmer, S. L. & Kay, S. A. Critical role for CCA1 and LHY in maintaining circadian rhythmicity in Arabidopsis. Curr. Biol. 12, 757–761 (2002)
Mizoguchi, T. et al. LHY and CCA1 are partially redundant genes required to maintain circadian rhythms in Arabidopsis. Dev. Cell 2, 629–641 (2002)
Suarez-Lopez, P. et al. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410, 1116–1120 (2001)
Christie, J. M. et al. Arabidopsis NPH1: a flavoprotein with the properties of a photoreceptor for phototropism. Science 282, 1698–1701 (1998)
Sakai, T. et al. Arabidopsis nph1 and npl1: blue light receptors that mediate both phototropism and chloroplast relocation. Proc. Natl Acad. Sci. USA 98, 6969–6974 (2001)
Salomon, M., Christie, J. M., Knieb, E., Lempert, U. & Briggs, W. R. Photochemical and mutational analysis of the FMN-binding domains of the plant blue light receptor, phototropin. Biochemistry 39, 9401–9410 (2000)
Crosson, S. & Moffat, K. Structure of a flavin-binding plant photoreceptor domain: insights into light-mediated signal transduction. Proc. Natl Acad. Sci. USA 98, 2995–3000 (2001)
Lin, C. Blue light receptors and signal transduction. Plant Cell 14 (suppl.), S207–S225 (2002)
Yanovsky, M. J., Mazzella, M. A., Whitelam, G. C. & Casal, J. J. Resetting of the circadian clock by phytochromes and cryptochromes in Arabidopsis. J. Biol. Rhythms 16, 523–530 (2001)
Kinoshita, T. et al. phot1 and phot2 mediate blue light regulation of stomatal opening. Nature 414, 656–660 (2001)
Sessions, A. et al. A high-throughput Arabidopsis reverse genetics system. Plant Cell 14, 2985–2994 (2002)
Onouchi, H., Igeno, M. I., Perilleux, C., Graves, K. & Coupland, G. Mutagenesis of plants overexpressing CONSTANS demonstrates novel interactions among Arabidopsis flowering-time genes. Plant Cell 12, 885–900 (2000)
Carrington, J. C., Freed, D. D. & Leinicke, A. J. Bipartite signal sequence mediates nuclear translocation of the plant potyviral NIa protein. Plant Cell 3, 953–962 (1991)
Puig, O. et al. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24, 218–229 (2001)
Hajdukiewicz, P., Svab, Z. & Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol. Biol. 25, 989–994 (1994)
Blazquez, M. A. & Weigel, D. Independent regulation of flowering by phytochrome B and gibberellins in Arabidopsis. Plant Physiol. 120, 1025–1032 (1999)
Swartz, T. E. et al. The photocycle of a flavin-binding domain of the blue light photoreceptor phototropin. J. Biol. Chem. 276, 36493–36500 (2001)
Acknowledgements
We thank T. Schultz, P. Más, F. Harmon and S. Hazen for critically reading the manuscript; Syngenta for the T-DNA insertion line; B. Bartel for fkf1 and the pGEX FKF1 LOV construct; T. Kagawa and M. Wada for phot1 phot2; M. Yanovsky for cry1 cry2; G. Coupland for 35S::CO; and J. Harper for the TAP tag construct. This work was supported by grants from the NIH (to S.A.K. and H.G.T.); a grant from the NSF (to W.R.B.); and a grant from JSPS Postdoctoral Fellowships for Research Abroad (to T.I.). This is manuscript 15868-CB of The Scripps Research Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Imaizumi, T., Tran, H., Swartz, T. et al. FKF1 is essential for photoperiodic-specific light signalling in Arabidopsis. Nature 426, 302–306 (2003). https://doi.org/10.1038/nature02090
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02090
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.