Abstract
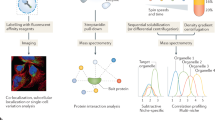
A fundamental goal of cell biology is to define the functions of proteins in the context of compartments that organize them in the cellular environment. Here we describe the construction and analysis of a collection of yeast strains expressing full-length, chromosomally tagged green fluorescent protein fusion proteins. We classify these proteins, representing 75% of the yeast proteome, into 22 distinct subcellular localization categories, and provide localization information for 70% of previously unlocalized proteins. Analysis of this high-resolution, high-coverage localization data set in the context of transcriptional, genetic, and protein–protein interaction data helps reveal the logic of transcriptional co-regulation, and provides a comprehensive view of interactions within and between organelles in eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Goffeau, A. et al. Life with 6000 genes. Science 274, 546, 563–567 (1996)
Velculescu, V. E. et al. Characterization of the yeast transcriptome. Cell 88, 243–251 (1997)
Wang, Y. et al. Precision and functional specificity in mRNA decay. Proc. Natl Acad. Sci. USA 99, 5860–5865 (2002)
Martzen, M. R. et al. A biochemical genomics approach for identifying genes by the activity of their products. Science 286, 1153–1155 (1999)
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001)
Uetz, P. et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000)
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl Acad. Sci. USA 98, 4569–4574 (2001)
Gavin, A.-C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002)
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002)
Lee, T. I. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002)
Ross-Macdonald, P. et al. Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413–418 (1999)
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Tong, A. H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001)
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002)
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998)
Dolinski, K. et al. Saccharomyces Genome Database 〈http://www-genome.stanford.edu/Saccharomyces〉 (2003).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998)
Campbell, R. E. et al. A monomeric red fluorescent protein. Proc. Natl Acad. Sci. USA 99, 7877–7882 (2002)
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E. S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003)
Cliften, P. et al. Finding functional features in Saccharomyces genomes by phylogenetic footprinting. Science 301, 71–76 (2003)
Rout, M. P. et al. The yeast nuclear pore complex: composition, architecture, and transport mechanism. J. Cell Biol. 148, 635–651 (2000)
Wigge, P. A. et al. Analysis of the Saccharomyces spindle pole by matrix-assisted laser desorption/ionization (MALDI) mass spectrometry. J. Cell Biol. 141, 967–977 (1998)
Bhattacharya, S., Chen, L., Broach, J. R. & Powers, S. Ras membrane targeting is essential for glucose signaling but not for viability in yeast. Proc. Natl Acad. Sci. USA 92, 2984–2988 (1995)
van Berkel, M. A., Caro, L. H., Montijn, R. C. & Klis, F. M. Glucosylation of chimeric proteins in the cell wall of Saccharomyces cerevisiae. FEBS Lett. 349, 135–138 (1994)
Gould, S. J. et al. Peroxisomal protein import is conserved between yeast, plants, insects and mammals. EMBO J. 9, 85–90 (1990)
Pelham, H. R., Hardwick, K. G. & Lewis, M. J. Sorting of soluble ER proteins in yeast. EMBO J. 7, 1757–1762 (1988)
Taylor, S. W., Fahy, E. & Ghosh, S. S. Global organellar proteomics. Trends Biotechnol. 21, 82–88 (2003)
Shou, W. et al. Exit from mitosis is triggered by Tem1-dependent release of the protein phosphatase Cdc14 from nucleolar RENT complex. Cell 97, 233–244 (1999)
Visintin, R., Hwang, E. S. & Amon, A. Cfi1 prevents premature exit from mitosis by anchoring Cdc14 phosphatase in the nucleolus. Nature 398, 818–823 (1999)
San-Segundo, P. A. & Roeder, G. S. Pch2 links chromatin silencing to meiotic checkpoint control. Cell 97, 313–324 (1999)
Rabitsch, K. P. et al. Kinetochore recruitment of two nucleolar proteins is required for homolog segregation in meiosis I. Dev. Cell 4, 535–548 (2003)
Andersen, J. S. et al. Directed proteomic analysis of the human nucleolus. Curr. Biol. 12, 1–11 (2002)
Hodges, P. E. et al. Annotating the human proteome: the Human Proteome Survey Database (HumanPSD) and an in-depth target database for G protein-coupled receptors (GPCR-PD) from Incyte Genomics. Nucleic Acids Res. 30, 137–141 (2002)
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997)
Holstege, F. C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998)
Ihmels, J. et al. Revealing modular organization in the yeast transcriptional network. Nature Genet. 31, 370–377 (2002)
Drawid, A., Jansen, R. & Gerstein, M. Genome-wide analysis relating expression level with protein subcellular localization. Trends Genet. 16, 426–430 (2000)
Bergmann, S., Ihmels, J. & Barkai, N. Iterative signature algorithm for the analysis of large-scale gene expression data. Phys. Rev. E 67, 031902 (2003)
Koc, E. C. et al. The large subunit of the mammalian mitochondrial ribosome. J. Biol. Chem. 276, 43958–43969 (2001)
von Mering, C. et al. Comparative assessment of large-scale data sets of protein–protein interactions. Nature 417, 399–403 (2002)
Breitkreutz, B.-J., Stark, C. & Tyers, M. The GRID: the General Repository for Interaction Datasets. Genome Biol. 4, R23 (2003)
Madden, K. & Snyder, M. Cell polarity and morphogenesis in budding yeast. Annu. Rev. Microbiol. 52, 687–744 (1998)
Trilla, J. A., Duran, A. & Roncero, C. Chs7p, a new protein involved in the control of protein export from the endoplasmic reticulum that is specifically engaged in the regulation of chitin synthesis in Saccharomyces cerevisiae. J. Cell Biol. 145, 1153–1163 (1999)
DeMarini, D. J. et al. A septin-based hierarchy of proteins required for localized deposition of chitin in the Saccharomyces cerevisiae cell wall. J. Cell Biol. 139, 75–93 (1997)
Cope, M. J., Yang, S., Shang, C. & Drubin, D. G. Novel protein kinases Ark1p and Prk1p associate with and regulate the cortical actin cytoskeleton in budding yeast. J. Cell Biol. 144, 1203–1218 (1999)
Breitkreutz, B.-J., Stark, C. & Tyers, M. Osprey: a network visualization system. Genome Biol. 4, R22 (2003)
Acknowledgements
We thank F. Lam for designing the original relational data model and for assistance with database organization; M. Springer for sharing microscopy expertise; J. Newman for assistance with FACS analysis; N. Barkai, S. Ghaemmaghami and K. Bower for discussions and sharing results; R. Tsien for providing the pRSET–mRFP1 plasmid; M. Levin for assistance with preparation of RFP-tagged strains; R. Marion for contributing GFP-tagged strains of CAK1, PHO4, RTS2, SET1 and STP4; A. DePace for preparing the illustration in Fig. 4c; D. Ahern, F. Sanchez, A. Belle and M. Liku for primer synthesis and other technical assistance; and S. Emr and members of the O'Shea and Weissman laboratories for critical discussion of the work. This work was supported by the Howard Hughes Medical Institute and the David and Lucile Packard Foundation (J.S.W. and E.K.O.). J.V.F. is the recipient of a Ruth L. Kirschstein National Research Service Award. The database and strain collection information is accessible at http://yeastgfp.ucsf.edu.Authors' contributions Strain construction and analysis was performed by W.-K.H. and J.V.F., bioinformatics analysis and database management by L.C.G., with additional database management by A.S.C., and oligonucleotide primer design by R.W.H.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Huh, WK., Falvo, J., Gerke, L. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003). https://doi.org/10.1038/nature02026
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02026
This article is cited by
-
From beer to breadboards: yeast as a force for biological innovation
Genome Biology (2024)
-
Nuclear Hsp104 safeguards the dormant translation machinery during quiescence
Nature Communications (2024)
-
Engineering stringent genetic biocontainment of yeast with a protein stability switch
Nature Communications (2024)
-
PIFiA: self-supervised approach for protein functional annotation from single-cell imaging data
Molecular Systems Biology (2024)
-
A nanobody-based strategy for rapid and scalable purification of human protein complexes
Nature Protocols (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.