Abstract
Knowledge of the elastic properties of DNA is required to understand the structural dynamics of cellular processes such as replication and transcription. Measurements of force and extension on single molecules of DNA1,2,3 have allowed direct determination of the molecule's mechanical properties, provided rigorous tests of theories of polymer elasticity4, revealed unforeseen structural transitions induced by mechanical stresses3,5,6,7, and established an experimental and conceptual framework for mechanical assays of enzymes that act on DNA8. However, a complete description of DNA mechanics must also consider the effects of torque, a quantity that has hitherto not been directly measured in micromanipulation experiments. We have measured torque as a function of twist for stretched DNA—torsional strain in over- or underwound molecules was used to power the rotation of submicrometre beads serving as calibrated loads. Here we report tests of the linearity of DNA's twist elasticity, direct measurements of the torsional modulus (finding a value ∼40% higher than generally accepted), characterization of torque-induced structural transitions, and the establishment of a framework for future assays of torque and twist generation by DNA-dependent enzymes. We also show that cooperative structural transitions in DNA can be exploited to construct constant-torque wind-up motors and force–torque converters.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Smith, S. B., Finzi, L. & Bustamante, C. Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science 258, 1122–1126 (1992)
Strick, T. R., Allemand, J. F., Bensimon, D., Bensimon, A. & Croquette, V. The elasticity of a single supercoiled DNA molecule. Science 271, 1835–1837 (1996)
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: The elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996)
Bustamante, C., Marko, J. F., Siggia, E. D. & Smith, S. Entropic elasticity of lambda-phage DNA. Science 265, 1599–1600 (1994)
Cluzel, P. et al. DNA: An extensible molecule. Science 271, 792–794 (1996)
Allemand, J. F., Bensimon, D., Lavery, R. & Croquette, V. Stretched and overwound DNA forms a Pauling-like structure with exposed bases. Proc. Natl Acad. Sci. USA 95, 14152–14157 (1998)
Leger, J. F. et al. Structural transitions of a twisted and stretched DNA molecule. Phys. Rev. Lett. 83, 1066–1069 (1999)
Bustamante, C., Bryant, Z. & Smith, S. B. Ten years of tension: Single-molecule DNA mechanics. Nature 421, 423–427 (2003)
Bouchiat, C. & Mezard, M. Elasticity model of a supercoiled DNA molecule. Phys. Rev. Lett. 80, 1556–1559 (1998)
Sarkar, A., Leger, J. F., Chatenay, D. & Marko, J. F. Structural transitions in DNA driven by external force and torque. Phys. Rev. E 63, 051903 (2001)
Smith, S. B., Cui, Y. & Bustamante, C. Optical-trap force transducer that operates by direct measurement of light momentum. Methods Enzymol. 361, 134–162 (2003)
Strick, T. R., Bensimon, D. & Croquette, V. Micro-mechanical measurement of the torsional modulus of DNA. Genetica 106, 57–62 (1999)
Selvin, P. R. et al. Torsional rigidity of positively and negatively supercoiled DNA. Science 255, 82–85 (1992)
Millar, D. P., Robbins, R. J. & Zewail, A. H. Direct observation of the torsional dynamics of DNA and RNA by picosecond spectroscopy. Proc. Natl Acad. Sci. USA 77, 5593–5597 (1980)
Heath, P. J., Clendenning, J. B., Fujimoto, B. S. & Schurr, J. M. Effect of bending strain on the torsion elastic constant of DNA. J. Mol. Biol. 260, 718–730 (1996)
Horowitz, D. S. & Wang, J. C. Torsional rigidity of DNA and length dependence of the free energy of DNA supercoiling. J. Mol. Biol. 173, 75–91 (1984)
Shore, D. & Baldwin, R. L. Energetics of DNA twisting. II. Topoisomer analysis. J. Mol. Biol. 170, 983–1007 (1983)
Crothers, D. M., Drak, J., Kahn, J. D. & Levene, S. D. DNA bending, flexibility, and helical repeat by cyclization kinetics. Methods Enzymol. 212, 3–29 (1992)
Vologodskii, A. V. & Marko, J. F. Extension of torsionally stressed DNA by external force. Biophys. J. 73, 123–132 (1997)
Moroz, J. D. & Nelson, P. Entropic elasticity of twist-storing polymers. Macromolecules 31, 6333–6347 (1998)
Yasuda, R., Miyata, H. & Kinosita, K. Jr Direct measurement of the torsional rigidity of single actin filaments. J. Mol. Biol. 263, 227–236 (1996)
Williams, M. C., Rouzina, I. & Bloomfield, V. A. Thermodynamics of DNA interactions from single molecule stretching experiments. Acc. Chem. Res. 35, 159–166 (2002)
Soong, R. K. et al. Powering an inorganic nanodevice with a biomolecular motor. Science 290, 1555–1558 (2000)
Yasuda, R., Noji, H., Kinosita, K. Jr & Yoshida, M. F1-ATPase is a highly efficient molecular motor that rotates with discrete 120 degree steps. Cell 93, 1117–1124 (1998)
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003)
Harada, Y. et al. Direct observation of DNA rotation during transcription by Escherichia coli RNA polymerase. Nature 409, 113–115 (2001)
Wobbe, C. R., Dean, F., Weissbach, L. & Hurwitz, J. In vitro replication of duplex circular DNA containing the simian virus 40 DNA origin site. Proc. Natl Acad. Sci. USA 82, 5710–5714 (1985)
Davenport, R. J., Wuite, G. J., Landick, R. & Bustamante, C. Single-molecule study of transcriptional pausing and arrest by E. coli RNA polymerase. Science 287, 2497–2500 (2000)
Davis, M. H. The slow translation and rotation of two unequal spheres in a viscous fluid. Chem. Eng. Sci. 24, 1769–1776 (1969)
Acknowledgements
We thank E. Watson and Y. Inclán for technical assistance, E. Nogales for microscope time, and A. Vologodskii, V. Croquette, D. Bensimon, D. Collin, N. Pokala and Y. Chemla for critical readings of the manuscript and/or discussions. Z.B. is an HHMI predoctoral fellow, M.D.S. is supported by a PMMB training grant, and J.G. holds a fellowship from the Hertz Foundation. This work was supported by the NIH and DOE.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2003_BFnature01810_MOESM1_ESM.avi
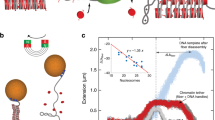
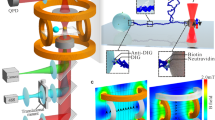
Supplementary Movie 1: Assembly of the experimental system. The molecular construct is stretched between an anti-DIG bead on the micropipette and an anti-fluorescein bead held in buffer flow (right to left). A 520 nm streptavidin-coated bead is held in the optical trap in the lower portion of the screen. The stage is moved to position the biotinylated portion of the molecule near the streptavidin bead, which becomes attached to the DNA and is pulled out of the trap. The lower bead is then placed in the trap to complete the assembly of the system (Fig. 1b). In the final half of the movie, the tension on the molecule is repeatedly increased to ~65 pN. Overstretching occurs in the lower DNA segment but not the upper segment, confirming the existence of a nick below the bead (allowing the DNA to unwind and enter the overstretched state) but not above it. Bead rotation is inhibited by flow. (AVI 2627 kb)
41586_2003_BFnature01810_MOESM2_ESM.avi
Supplementary Movie 2: 520 nm bead rotation during relaxation of an overwound molecule. A molecule with a 520 nm rotor bead was overwound as in Figure 1b. The movie begins subsequent to release of the rotor. The bead rotates at constant velocity for the first ~35 s, due to constant-torque conversion of P-DNA to B-DNA. Next, the angular velocity decays as torque and twist are removed from the molecule. Constant tension was maintained at 45 pN. (AVI 10402 kb)
41586_2003_BFnature01810_MOESM3_ESM.avi
Supplementary Movie 3: Constant-torque rotation powered by the P-B transition is shown for a 760 nm rotor bead. Constant tension was maintained at 45 pN. Off-axis alignment of outer beads is due to an eccentric DNA attachment point; DNA is held vertically as judged by the absence of horizontal force. (AVI 6085 kb)
41586_2003_BFnature01810_MOESM4_ESM.avi
Supplementary Movie 4: Overstretching and rewinding. Overstretching and rewinding is shown for a molecule with a 400 nm rotor bead (see Figure 3a). During the first ~26 s of the movie, the molecule is held at 85 pN and the bead rotation reflects unwinding of the molecule during the B-S transition. The separation of the beads increases at constant tension as the molecule converts to the longer S form. During the last part of the movie, the molecule is relaxed to 15 pN and the bead rotates in the opposite direction as the molecule rewinds. The movie is 2X actual speed. (AVI 12270 kb)
Rights and permissions
About this article
Cite this article
Bryant, Z., Stone, M., Gore, J. et al. Structural transitions and elasticity from torque measurements on DNA. Nature 424, 338–341 (2003). https://doi.org/10.1038/nature01810
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01810
This article is cited by
-
Exploring TRF2-Dependent DNA Distortion Through Single-DNA Manipulation Studies
Communications Biology (2024)
-
The energy landscape for R-loop formation by the CRISPR–Cas Cascade complex
Nature Structural & Molecular Biology (2023)
-
Storage of mechanical energy in DNA nanorobotics using molecular torsion springs
Nature Physics (2023)
-
Probing the microscopic structure and flexibility of oxidized DNA by molecular simulations
Indian Journal of Physics (2022)
-
Supercoiling and looping promote DNA base accessibility and coordination among distant sites
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.