Abstract
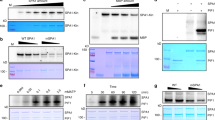
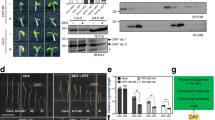
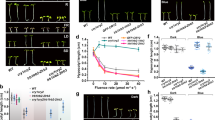
Far-red light regulates many aspects of seedling development, such as inhibition of hypocotyl elongation and the promotion of greening1, acting in part through phytochrome A (phyA). The RING motif protein COP1 is also important because cop1 mutants exhibit constitutive photomorphogenesis in darkness2,3. COP1 is present in the nucleus in darkness but is gradually relocated to the cytoplasm upon illumination4. Here we show that COP1 functions as an E3 ligase ubiquitinating both itself and the myb transcription activator LAF1, which is required for complete phyA responses5. In transgenic plants, inducible COP1 overexpression leads to a decrease in LAF1 concentrations, but is blocked by the proteasome inhibitor MG132. The coiled-coil domain of SPA1, a negative regulator of phyA signalling6, has no effect on COP1 auto-ubiquitination but facilitates LAF1 ubiquitination at low COP1 concentrations. These results indicate that, in darkness, COP1 functions as a repressor of photomorphogenesis by promoting the ubiquitin-mediated proteolysis of a subset of positive regulators, including LAF1. After the activation of phyA, SPA1 stimulates the E3 activity of residual nuclear COP1 to ubiquitinate LAF1, thereby desensitizing phyA signals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neff, M. M., Fankhauser, C. & Chory, J. Light: an indicator of time and place. Genes Dev. 14, 257–271 (2000)
Deng, X. W., Caspar, T. & Quail, P. H. cop1: a regulatory locus involved in light controlled development and gene expression in Arabidopsis. Genes Dev. 5, 1172–1182 (1991)
von Arnim, A. G. & Deng, X. W. Ring finger motif of Arabidopsis thaliana COP1 defines a new class of zinc-binding domain. J. Biol. Chem. 268, 19626–19631 (1993)
von Arnim, A. G., Osterlund, M. T., Kwok, S. F. & Deng, X. W. Genetic and developmental control of nuclear accumulation of COP1, a repressor of photomorphogenesis in Arabidopsis. Plant Physiol. 114, 779–788 (1997)
Ballesteros, M. L. et al. LAF1, a MYB transcription activator for phytochrome A signaling. Genes Dev. 15, 2613–2625 (2001)
Hoecker, U., Xu, Y. & Quail, P. H. SPA1: a new genetic locus involved in phytochrome A-specific signal transduction. Plant Cell 10, 19–33 (1998)
Ma, L. et al. Light control of Arabidopsis development entails coordinated regulation of genome expression and cellular pathways. Plant Cell 13, 2589–2607 (2001)
Quail, P. H. et al. Phytochromes: photosensory perception and signal transduction. Science 268, 675–680 (1995)
Quail, P. H. Phytochrome photosensory signalling networks. Nature Rev. Mol. Cell Biol. 3, 85–93 (2002)
Nagy, F. & Schafer, E. Phytochromes control photomorphogenesis by differentially regulated, interacting signaling pathways in higher plants. Annu. Rev. Plant Biol. 53, 329–355 (2002)
Hardtke, C. S. & Deng, X. W. The cell biology of the COP/DET/FUS proteins. Regulating proteolysis in photomorphogenesis and beyond? Plant Physiol. 124, 1548–1557 (2000)
Ang, L. H. et al. Molecular interaction between COP1 and HY5 defines a regulatory switch for light control of Arabidopsis development. Mol. Cell 1, 213–222 (1998)
Hoecker, U. & Quail, P. H. The phytochrome A-specific signaling intermediate SPA1 interacts directly with COP1, a constitutive repressor of light signaling in Arabidopsis. J. Biol. Chem. 276, 38173–38178 (2001)
Stacey, M. G. & von Arnim, A. G. A novel motif mediates the targeting of the Arabidopsis COP1 protein to subnuclear foci. J. Biol. Chem. 274, 27231–27236 (1999)
Torii, K. U. et al. The RING finger motif of photomorphogenic repressor COP1 specifically interacts with the RING-H2 motif of a novel Arabidopsis protein. J. Biol. Chem. 274, 27674–27681 (1999)
Osterlund, M. T., Hardtke, C. S., Wei, N. & Deng, X. W. Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405, 462–466 (2000)
Hardtke, C. S., Okamoto, H., Stoop-Myer, C. & Deng, X. W. Biochemical evidence for ubiquitin ligase activity of the Arabidopsis COP1 interacting protein 8 (CIP8). Plant J. 30, 385–394 (2002)
Torii, K. U., McNellis, T. W. & Deng, X. W. Functional dissection of Arabidopsis COP1 reveals specific roles of its three structural modules in light control of seedling development. EMBO J. 17, 5577–5587 (1998)
Xie, Q. et al. SINAT5 promotes ubiquitin-related degradation of NAC1 to attenuate auxin signals. Nature 419, 167–170 (2002)
Zuo, J., Niu, Q. W. & Chua, N. H. An estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000)
Dieterle, M., Zhou, Y. C., Schafer, E., Funk, M. & Kretsch, T. EID1, an F-box protein involved in phytochrome A-specific light signaling. Genes Dev. 15, 939–944 (2001)
Hoecker, U., Tepperman, J. M. & Quail, P. H. SPA1, a WD-repeat protein specific to phytochrome A signal transduction. Science 284, 496–499 (1999)
Holley, C. L., Olson, M. R., Colon-Ramos, D. A. & Kornbluth, S. Reaper eliminates IAP proteins through stimulated IAP degradation and generalized translational inhibition. Nature Cell Biol. 4, 439–444 (2002)
Ryoo, H. D., Bergmann, A., Gonen, H., Ciechanover, A. & Steller, H. Regulation of Drosophila IAP1 degradation and apoptosis by reaper and ubcD1. Nature Cell Biol. 4, 432–438 (2002)
Yoo, S. J. et al. Hid, Rpr and Grim negatively regulate DIAP1 levels through distinct mechanisms. Nature Cell Biol. 4, 416–424 (2002)
Wilson, R. et al. The DIAP1 RING finger mediates ubiquitination of Dronc and is indispensable for regulating apoptosis. Nature Cell Biol. 4, 445–450 (2002)
Lafarga, M. et al. Clastosome: A subtype of nuclear body enriched in 19S and 20S proteasomes, ubiquitin, and protein substrates of proteasome. Mol. Biol. Cell 13, 2771–2782 (2002)
Xie, Q., Frugis, G., Colgan, D. & Chua, N. H. Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev. 14, 3024–3036 (2000)
Kost, B., Spielhofer, P. & Chua, N. H. A GFP-mouse talin fusion protein labels plant actin filaments in vivo and visualizes the actin cytoskeleton in growing pollen tubes. Plant J. 16, 393–401 (1998)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Acknowledgements
We thank X.-W. Deng for COP1 cDNA, and P. Hare for discussions. This work was supported by an NIH grant to N.-H.C. J.-Y.Y. is a graduate student on leave from Chung Hsing University, Taiwan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Seo, H., Yang, JY., Ishikawa, M. et al. LAF1 ubiquitination by COP1 controls photomorphogenesis and is stimulated by SPA1. Nature 423, 995–999 (2003). https://doi.org/10.1038/nature01696
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01696
This article is cited by
-
Arabidopsis COP1 guides stomatal response in guard cells through pH regulation
Communications Biology (2024)
-
OAF is a DAF-like gene that controls ovule development in plants
Communications Biology (2023)
-
Light signaling as cellular integrator of multiple environmental cues in plants
Physiology and Molecular Biology of Plants (2023)
-
Shade avoidance syndrome in soybean and ideotype toward shade tolerance
Molecular Breeding (2023)
-
The Arabidopsis Phytoglobin 2 mediates phytochrome B (phyB) light signaling responses during somatic embryogenesis
Planta (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.