Abstract
Neurospora crassa is a central organism in the history of twentieth-century genetics, biochemistry and molecular biology. Here, we report a high-quality draft sequence of the N. crassa genome. The approximately 40-megabase genome encodes about 10,000 protein-coding genes—more than twice as many as in the fission yeast Schizosaccharomyces pombe and only about 25% fewer than in the fruitfly Drosophila melanogaster. Analysis of the gene set yields insights into unexpected aspects of Neurospora biology including the identification of genes potentially associated with red light photobiology, genes implicated in secondary metabolism, and important differences in Ca2+ signalling as compared with plants and animals. Neurospora possesses the widest array of genome defence mechanisms known for any eukaryotic organism, including a process unique to fungi called repeat-induced point mutation (RIP). Genome analysis suggests that RIP has had a profound impact on genome evolution, greatly slowing the creation of new genes through genomic duplication and resulting in a genome with an unusually low proportion of closely related genes.
Similar content being viewed by others
Main
Research on Neurospora in the early part of the twentieth century paved the way for modern genetics and molecular biology. First documented in 1843 as a contaminant of bakeries in Paris1, Neurospora was developed as an experimental organism in the 1920s2,3. Subsequent work on Neurospora by Beadle and Tatum4 in the 1940s established the relationship between genes and proteins, summarized in the ‘one-gene-one-enzyme’ hypothesis. In the latter half of the century, Neurospora had a central role as a model organism, contributing to the fundamental understanding of genome defence systems, DNA methylation, mitochondrial protein import, circadian rhythms, post-transcriptional gene silencing and DNA repair5. Because Neurospora is a multicellular filamentous fungus, it has also provided a system to study cellular differentiation and development as well as other aspects of eukaryotic biology6.
The legacy of over 70 years of research7, coupled with the availability of molecular and genetic tools, offers enormous potential for continued discovery. The sequencing of the N. crassa genome was undertaken to maximize this potential. Here, we report an initial sequence and analysis of the Neurospora genome.
Neurospora genome sequence
The Neurospora genome is much larger (greater than 40 megabases (Mb)) than that of S. pombe and Saccharomyces cerevisiae (both about 12 Mb). Accordingly, first we sought to produce and analyse a high-quality draft sequence en route to a finished sequence.
The genome sequence was assembled from deep whole-genome shotgun (WGS) coverage obtained by paired-end sequencing from a variety of clone types (Supplementary Information). In all, the data provided an average of >20-fold sequence coverage and >98-fold physical coverage of the genome. The Arachne package8 was used to assemble the draft genome sequence. The resulting assembly consists of 958 sequence contigs with a total length of 38.6 Mb (Table 1) and an N50 length of 114.5 kilobases (kb) (that is, 50% of all bases are contained in contigs of at least 114.5 kb). Contigs were assembled into 163 scaffolds with a total length of 39.9 Mb (including gaps between contigs) and an N50 length of 1.56 Mb.
Most of the assembly (97%) is contained in the 44 largest scaffolds, and there are 38 tiny scaffolds with lengths <4 kb. Forty-two of the large scaffolds (and one of the smaller ones) could be anchored readily to the Neurospora genetic map7 by virtue of their containing genetic markers with sequence.
The assembly has long-range continuity, with the N50 scaffold size being nearly 1,000-fold larger than the average gene size. The assembly represents the vast majority of the genome, as assessed by comparison with available finished sequence and genetic markers. It contains 99.13% of available finished sequence (17 Mb from linkage groups II and V9) and all of the 252 genetic markers with sequence. This estimate, however, does not account for unusual genomic regions such as the ribosomal DNA repeats, centromeres and telomeres; such regions may contain about 1.7 Mb of additional sequence10, corresponding to 2–3% of the genome that cannot be assembled readily with available techniques. The long-range continuity of the assembly was also confirmed by comparisons with previously described bacterial artificial chromosome (BAC) physical maps for linkage groups II and V11, as only one discrepancy was noted.
The assembly also has high accuracy, with 99.5% of the sequence having Arachne quality scores ≥30. Comparison with the 17 Mb of finished sequence confirms the sequence accuracy, with a discrepancy rate for this subset of less than 10-5. The comparison also largely confirms the assembly, as only 12 minor discrepancies were identified (Supplementary Information).
Genes
Gene count and basic characteristics
A total of 10,082 protein-coding genes (9,200 longer than 100 amino acids) were predicted (Table 1 and Supplementary S0). This constitutes nearly twice as many genes as in S. pombe (about 4,800) and S. cerevisiae (about 6,300), and nearly as many as in D. melanogaster (about 14,300). Genes cover at least 44% of the genome sequence with an average gene density of one gene per 3.7 kb. The average gene length of 1.67 kb is slightly longer than the 1.4-kb average gene length for both S. cerevisiae and S. pombe. The difference in gene length is due to the greater number of introns in Neurospora genes—an average of 1.7 introns per gene with an average intron size of 134 nucleotides. Notably, most predicted Neurospora introns lack a polypyrimidine tract, which is common in other eukaryotic introns, but do contain a strong branchpoint sequence (Supplementary Information).
Comparative analysis
A total of 4,140 (41%) Neurospora proteins lack significant matches to known proteins from public databases (Table 1), reflecting the early stage of fungal genome exploration and the diversity of fungal genes remaining to be described. Furthermore, 5,805 (57%) Neurospora proteins do not have significant matches to genes in either of the sequenced yeast species (Supplementary Information). When compared to sequenced eukaryotes, a total of 1,421 (14%) Neurospora genes display best BLASTP matches to proteins in either plants or animals (Supplementary Information). Of these, 584 lack high-scoring hits to either sequenced yeast species. These data reflect the biology shared by filamentous fungi and higher eukaryotes, which in a number of cases is absent in the yeasts.
Epigenetics, genome defence and genome evolution
Neurospora is an important model for the study of epigenetic phenomena, possessing a wide variety of epigenetic mechanisms and related genome defence mechanisms. The most remarkable of these mechanisms is repeat-induced point mutation (RIP), a process unique to fungi.
Repeat-induced point mutation
First discovered in Neurospora12,13, RIP is a process that efficiently detects and mutates both copies of a sequence duplication. RIP acts during the haploid dikaryotic stage of the Neurospora sexual reproductive cycle, causing numerous C•G to T•A mutations within duplicated sequences. In a single passage through the sexual cycle, up to 30% of the C•G pairs in duplicated sequences can be mutated, with a strong preference for C to T mutations occurring at CpA dinucleotides14. The pattern of mutations produces a characteristic skewing of dinucleotide frequencies that allows RIP-mutated sequences to be detected accurately15. RIP requires a minimal duplicated sequence length of about 400 base pairs (bp)16 and greater than roughly 80% sequence identity between duplicates17. In addition to suffering mutations, RIP-mutated sequences are frequent targets for DNA methylation. As with mammals, DNA methylation has been shown to cause gene silencing in Neurospora18. RIP thus mutates and epigenetically silences repetitive DNA.
RIP has been proposed to act as a defence against selfish or mobile DNA13. However, because RIP mutation and methylation can extend beyond the bounds of duplicated sequences19, RIP can have both mutational and epigenetic effects on neighbouring unique sequences. Furthermore, RIP acts on all duplicated sequences, including those arising from large-scale chromosomal duplications as well as gene duplications20. The presence of RIP thus has profound consequences for the evolution of the Neurospora genome. Indeed, it has been proposed that RIP might prevent gene innovation through gene duplication13,21. With the availability of the Neurospora genome sequence, we were able to address this hypothesis.
Multigene families
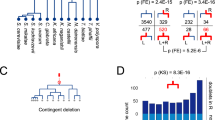
To investigate the impact of RIP on protein families in Neurospora, genes were clustered into ‘multigene families’ on the basis of an all versus all comparison of protein sequences (see Methods). As shown in Fig. 1, the percentage of genes in multigene families in selected sequenced eukaryotes is correlated with genome size. However, in marked contrast to the other analysed organisms, Neurospora possesses many fewer genes in multigene families than expected. When the analysis is expanded to include an additional 17 sequenced prokaryotes (Supplementary Information), only Mycoplasma genitalium, Mycoplasma pulminus, Ureaplasma urealyticum and Vibrio cholerae display a correspondingly small proportion of genes in families. This is noteworthy considering that the Mycoplasma genus is thought to have undergone reductive evolution and represent minimal life forms22.
Our analysis reveals another characteristic of Neurospora gene families. Unlike other sequenced eukaryotes, Neurospora possesses only a handful of highly similar gene pairs. Figure 2 displays histograms of amino acid and nucleotide similarities between each gene in the six organisms analysed and the best-matching gene in that organism. A significant proportion of genes have best matches with greater than 80% amino acid and nucleotide identity in all the organisms considered except Neurospora. Neurospora contains only eight genes with top matches of greater than 80% amino acid or coding sequence identity. This value is significant because, as described above, RIP mutates duplicated sequences that display greater than about 80% nucleotide similarity. Thus, the small proportion of genes in multigene families and the near absence of highly similar genes are consistent with the actions of RIP.
a, b, Histogram of amino acid (a) and nucleotide (b) per cent identity of top-scoring self-matches for genes in selected sequenced eukaryotic genomes. For each organism, the protein and coding regions for each gene (not including pseudogenes) were compared to those of every other gene in the same genome using BLASTX. Top-scoring matches were aligned using ClustalW and per cent identities calculated. In contrast to other eukaryotes, Neurospora possesses only eight genes with a top match of greater than 80% amino acid or nucleotide identity.
An example of the lack of highly similar genes in multigene families is revealed in an analysis of predicted major facilitator superfamily (MFS) sugar transporters (Fig. 3). Neurospora has about the same number of predicted MFS sugar transporters as S. cerevisiae. However, a phylogenetic analysis of fungal sugar transporters indicates that the Neurospora proteins are substantially more divergent than those of S. cerevisiae as well as those of S. pombe. Furthermore, the Neurospora transporters contain no apparent instances of recent duplication. In contrast, most of the S. cerevisiae HXT hexose and S. pombe GHT transporters represent two relatively recent and independent expansions and include very recently duplicated genes. Thus, despite a diversity of MFS sugar transporters, Neurospora seems to lack close paralogues in this gene family, consistent with the results of the genome-wide multigene family analysis.
Phylogenetic tree of major facilitator superfamily (MFS) sugar transporters from S. cerevisiae, S. pombe and Neurospora. Coloured dots represent branching points between predicted paralogous genes in Neurospora (red), S. cerevisiae (blue) and S. pombe (green). In contrast to both yeast species, Neurospora transporters contain no predicted instances of recent paralogous duplication.
Analyses of other gene families yielded similar results (data not shown). Furthermore, the paucity of closely related sequences is evident not only at the level of complete genes, but even at the level of individual exons, protein domains and protein architectures (Supplementary S4).
Gene evolution through gene duplication
The above results suggest that RIP has had a powerful impact in suppressing the creation of new genes or partial genes through genomic duplication. This is consistent with the large number of mutations induced in duplicated sequences by RIP. Computer simulations (see Methods) indicate that after a gene duplication, each copy has an 80% probability of acquiring an in-frame stop codon after only a single round of RIP and a 99.5% probability by the point that RIP has mutated the copies to less than 85% nucleotide similarity. The high frequency of stop codons reflects the preference of RIP for mutating CpA to TpA, increasing the prevalence of the stop codons TAA and TAG.
These results raise the critical question of whether any significant gene duplication has occurred in Neurospora subsequent to the acquisition of RIP. We searched for empirical evidence of duplicated genes that have survived RIP by analysing the set of Neurospora coding sequences using two different measures15 for detecting RIP-mutated sequences (see Methods). These measures use the characteristic skewing of dinucleotides produced by RIP to detect mutated sequences. According to these measures, only 59 of the 9,200 predicted genes encoding proteins ≥100 amino acids show evidence of mutation by RIP. Of these, only eight consist of pairs of predicted duplicated genes (genes in the same multigene family) in which both copies are predicted to be RIP-mutated. Thus, few pairs of duplicated genes display evidence of having both survived RIP (Supplementary Information).
Gene duplication is thought to have a primary role in the innovation of new genes23. However, taken together, our data support the conclusion that most, if not all, paralogous genes in Neurospora duplicated and diverged before the emergence of RIP, and since that point the evolution of new genes through gene duplication has been virtually arrested. This conclusion raises the question of whether and how Neurospora is able to evolve new genes. A number of mechanisms that do not involve gene duplication are conceivable, although ultimately a conclusive analysis may be possible only by comparing the genome of Neurospora with the genomes of closely related species to illuminate recent evolutionary history. Nonetheless, our results indicate that the cost to Neurospora of increased genome security through RIP is a significant impact on the evolution of new gene functions through gene duplication.
Repetitive DNA
An analysis of repeat sequences longer than 200 bp and with greater than 65% similarity (see Methods) revealed that 10% of the Neurospora assembly consists of repeat sequences, consistent with previously reported estimates21.
The repeat sequence of Neurospora provides a testament to the efficiency of RIP. Applying the measures of RIP mentioned above to the Neurospora genome revealed that most of the repetitive sequences (81%) in Neurospora have been mutated by RIP. Conversely, only 18% of predicted RIP-mutated sequence is non-repetitive, potentially reflecting loss of the corresponding duplicated sequence. As described above, duplications greater than about 400 bp are susceptible to RIP16. In keeping with this, we observe that over 97% of genomic repeats greater than 400 bp in length are RIP-mutated. Moreover, repeats longer than 400 bp clustered by sequence similarity display an average sequence identity within clusters of 78%, with 93% of clusters displaying an average identity of less than 85%. This corresponds to previous estimates indicating that RIP requires greater than about 80% sequence identity to detect duplicated sequences.
Consistent with the hypothesis that RIP acts as a defence mechanism against selfish DNA13, no intact mobile elements were identified. Furthermore, a significant proportion of the Neurospora RIP-mutated sequence (46% of repetitive nucleotides) can be identified as relics of mobile elements (Supplementary Information).
Ribosomal RNA
The only large repetitive sequences known to have survived RIP in Neurospora are the approximately 175–200 copies10 of the large rDNA tandem repeat containing the 17S, 5.8S and 25S rRNA genes. As in higher eukaryotes, these tandem repeats occur within the nucleolar organizer region (NOR), and their resistance to RIP seems to stem from this localization13. Within the genome sequence we found several copies of the rDNA repeat outside the NOR. In every case, they display evidence of mutation by RIP, consistent with previous observations13. Thus, the sequence of the rDNA repeat does not in itself seem to confer resistance to RIP.
The 5S rRNA genes in Neurospora have survived RIP in a different manner. In contrast to most higher eukaryotes in which the 5S rRNA genes form tandem repeats, the 5S genes are dispersed throughout the genome in Neurospora24. A total of 74 copies comprising several different subtypes of 5S rDNA are dispersed through all seven chromosomes. This dispersal coupled with their small size (approximately 120 nucleotides) ensures that they are not recognized by RIP.
DNA methylation
Neurospora has been used extensively as a model for studying DNA methylation in eukaryotes25. The Neurospora genome includes two potential cytosine DNA methyltransferase genes. One, called dim-2, is required for all known DNA methylation26. The other, called rid, is required for RIP and is a member of a family found thus far only in filamentous fungi27. In Neurospora, an estimated 1.5% of cytosines are methylated28,29, and it has been suggested that nearly all DNA methylation is a result of RIP1,30,31.
Plasmid reads for Neurospora were sequenced from libraries cloned separately in methylation-tolerant and methylation-intolerant strains of Escherichia coli. Although not intended for this purpose, these libraries provided a basis for predicting DNA methylation by comparing the representation of regions in sequence obtained from each library (see Methods). Testing the accuracy of such predictions, we found that 8 of 10 regions predicted to be methylated were experimentally confirmed as such. The predictions thus have good specificity—although they lack sensitivity (see Methods).
The specificity of the predictions provides insight into the pattern of methylation in the Neurospora genome. Regions predicted to be methylated show a marked correspondence to regions predicted to be repetitive and RIP-mutated (Fig. 4). Fully 85% correspond to predicted RIP-mutated sequences. However, a small proportion (10%) corresponds to predicted non-repetitive and non-RIP-mutated sequence. In two out of ten such cases, both the methylation and the non-repetitive nature of these sequences were experimentally verified. This raises the possibility that methylation in Neurospora may also have non-defence roles, as proposed for higher organisms.
Prediction of RIP, methylation and repeat sequence in 1-kb windows for selected contigs. Red lines plot the TpA/ApT RIP index (see Methods); red dots indicate windows predicted to be RIP-mutated (TpA/ApT > 1.2). Blue lines plot the proportion of reads from the methyl-tolerant library; blue dots indicate windows predicted to be methylated based on >70% methyl-tolerant reads (see Methods). Black lines plot repeat content as a fraction of nucleotides in each window that is in repetitive sequence; black dots indicate windows with >50% repeat sequence. Contigs were selected to illustrate regions predicted as methylated.
RNA silencing
Post-transcriptional gene silencing (PTGS), or RNA silencing, is widespread among organisms and is increasingly being recognized as a principal switch for controlling eukaryotic gene expression32. RNA-silencing pathways are thought to be derived from ancestral natural defence systems directed against invading nucleic acids33. Consistent with this, all known PTGS mechanisms share similar components34.
Neurospora possesses two RNA-silencing pathways. The first, called quelling, acts during vegetative growth. This pathway was uncovered through the study of three genes, qde-1, qde-2 and qde-3, coding respectively for an RNA-dependent RNA polymerase (RdRP), an argonaute and a RecQ helicase35. The second pathway, called meiotic silencing, acts during sexual reproduction36,37. Before our analysis, a gene called sad-1, encoding an RdRP, had been identified for this pathway38.
Our analysis of the Neurospora genome sequence uncovered several additional genes implicated in RNA silencing (Table 2). These include one RdRP, one argonaute-like protein and one RecQ-like helicase, as well as two dicer-like ribonucleases. A phylogenetic analysis (Supplementary S7) of the predicted RdRPs, argonaute-like proteins and dicer-like proteins indicates that the Neurospora genes comprise two paralogous sets. One set includes the three qde genes and is thus predicted to correspond to the quelling pathway. The other set includes sad-1, and in phylogenetic trees these genes branch consistently with those of the single pathway observed in S. pombe37,39. On the basis of this analysis, we predict that one of the identified dicers, Sms-3, belongs phylogenetically to the meiotic silencing pathway, whereas the other, dcl-2, belongs to the quelling pathway (Table 2). In addition, we predicted that the identified argonaute, Sms-2, also belongs phylogenetically to the meiotic silencing pathway. Subsequent experimental work has supported roles for Sms-2 (ref. 40) and Sms-3 (M. McLaughlin, D. W. Lee, R. Pratt and R. Aramayo, manuscript in preparation) in meiotic silencing. Taken together, these results suggest that meiotic silencing and quelling represent two phylogenetically distinct RNA-dependent silencing pathways. We further hypothesize that both might have evolved from a single ancestral RNA-silencing pathway.
Fungal biology and evolution
The Neurospora genome sequence provides an opportunity to study the genetic basis underlying the extraordinary biochemical and metabolic diversity exhibited by a filamentous fungus. Our analysis of the genome sequence has resulted in a number of surprising insights into the biology and evolution of Neurospora and other filamentous fungi.
Cell signalling and environmental responses
Discovery of putative red-light-sensing genes
Blue light is an important regulator of Neurospora growth and development, affecting the circadian rhythm of conidiation, carotenogenesis of hyphae and numerous facets of sexual development41. Although Neurospora photobiology has been studied intensively for more than two decades, the genome sequence has nonetheless revealed a number of previously uncharacterized sequences with similarity to blue-light-sensing genes, including both a cryptochrome homologue and a gene whose product contains a single PAS/LOV-type domain associated with light sensing.
Furthermore, Neurospora possesses two putative phytochrome homologues most similar to bacteriophytochromes—genes known for their role in red light sensing in prokaryotes—and a putative homologue of the Aspergillus nidulans velvet gene implicated in the regulation of both red and blue light responses. The presence of these genes is unexpected given that no red light photobiology has been described for Neurospora so far. It has been shown recently that in addition to red light sensing, some Arabidopsis phytochromes associate with cryptochromes to have a role in blue light sensing and signalling42. Therefore, the two phytochromes and the velvet homologue may also regulate this aspect of Neurospora photobiology.
Importance of two-component signalling in filamentous fungi
Mitogen-activated protein kinase (MAPK) pathways integrate signals from multiple receptor pathways including two-component signalling systems43. The basic two-component system consists of a histidine kinase and a cognate response regulator. The nine MAPK pathway proteins identified in the Neurospora genome sequence (Fig. 5) correspond to those found in S. pombe and S. cerevisiae, indicating that the basic MAPK machinery is conserved between these species. In contrast, Neurospora has a significantly expanded complement of 11 histidine kinases, as compared with one in S. cerevisiae and three in S. pombe. Two of the 11 genes have been characterized previously in Neurospora44, whereas a third is similar to proteins in Aspergillus fumigatus and A. nidulans that affect conidiation (L. A. Alex and M. I. Simon, unpublished observations; see also ref. 44). Functions for the remaining genes are unknown, although seven (including the two phytochromes discussed above) contain PAS/PAC domains, implicating them in oxygen and light responses. This number of histidine kinases suggests a larger role than previously expected, and reveals filamentous fungi to be more similar in this regard to plants, where two-component systems are abundant, than to animals, where these systems are absent.
The numbers identified for each gene are in parentheses. An asterisk indicates that the location in the plasma membrane and/or organelle membranes is not determined. AC, adenylyl cyclase; C, Ca2+ channel protein; CaM, calmodulin; Ca2+/CaM-reg, calcium- and calmodulin-regulated protein; CAP, cyclase-associated protein; DAG diacylglycerol; GPCR, G-protein-coupled receptor; Gα, G protein α-subunit; Gβ, G protein β-subunit; Gγ, G protein γ-subunit; HPT, histidine-containing phosphotransfer domain protein; MAPK, MAP kinase; MAPKK, MAPK kinase; MAPKKK, MAPKK kinase; PKA-C, protein kinase A catalytic subunit; PKA-R, protein kinase A regulatory subunit; PLC, phospholipase C; PKC, protein kinase C; T, Ca2+ transport protein (P-type Ca2+ ATPase, H+/Ca2+ exchanger, or Na+/Ca2+ exchanger).
A new family of G-protein-coupled receptors
Eukaryotic cells sense many environmental stimuli through seven-transmembrane-helix, G-protein-coupled receptors (GPCRs)45. Our analysis indicates that Neurospora possesses ten predicted seven-transmembrane-helix proteins (Fig. 5), three of which belong to a new class not previously identified in any fungus. These three genes encode proteins similar to cyclic AMP GPCRs from the protists Dictyostelium discoideum46 and Polysphodylium pallidum, and also to predicted proteins from Arabidopsis thaliana47 and Caenorhabditis elegans. The D. discoideum proteins sense cAMP levels during chemotaxis and multicellular development48. This suggests a possible analogous function in Neurospora. The existence of an extracellular cAMP signalling pathway has never been demonstrated previously in any fungal system.
In support of this hypothesis, along with the presence of putative cAMP receptors, Neurospora was found to possess the full complement of proteins required for the synthesis and degradation of cAMP. Furthermore, Neurospora wild-type strains accumulate cAMP in the extracellular medium49, although a role in extracellular signalling has not been established. Taken together, these data suggest the possibility that cAMP or a related molecule may serve as an extracellular signal in Neurospora.
Ca2+ sensory transduction in filamentous fungi
A considerable body of evidence, primarily from pharmacological studies, indicates that Ca2+ signalling regulates numerous processes in filamentous fungi50. However, the identification of the main components of even one Ca2+-mediated response pathway in filamentous fungi has remained elusive. The genome sequence of Neurospora has provided over 25 of the proteins likely to be necessary for Ca2+ signalling in filamentous fungi (Fig. 5).
A notable difference between Ca2+ signalling in Neurospora as compared with plants and animals was revealed by the genome sequence. An important aspect of Ca2+ signalling in plant and animal cells involves Ca2+ release from internal stores. This is commonly mediated by the second messengers inositol-1,4,5-trisphosphate (InsP3) and cADP ribose, or by Ca2+-induced Ca2+ release51. InsP3 is present within Neurospora hyphae52, and physiological evidence including intracellular membrane-associated, InsP3-activated Ca2+ channel activity supports a role in Ca2+ signalling53,54. In spite of this, Neurospora (and S. cerevisiae) lacks recognizable InsP3 receptors. In addition, neither ADP ribosyl cyclase nor ryanodine receptor proteins, principal components of Ca2+ release mechanisms in plant and animal cells, are found in Neurospora. These observations raise the question of whether other second messenger systems responsible for Ca2+ release from internal stores remain to be discovered in filamentous fungi.
Growth and development
Hyphal growth
True hyphae produced by filamentous fungi are tubular structures consisting of cellular compartments that are usually delineated by incomplete septa5. In contrast, the pseudohyphae produced by yeasts consist of chains of uninucleate elongated cells55 with no apparent cytoplasmic continuity. The molecular mechanisms underlying these two modes of growth are not well understood.
The two signalling pathways that regulate pseudohyphal growth in S. cerevisiae—the MAPK and cAMP modules—are both conserved in the Neurospora genome. In Candida albicans, capable of pseudohyphal, true hyphal and budding growth, both pathways are required for true hyphal production, suggesting a similar role in Neurospora56. However, at least three transcription factors—Tec1p, Flo8p and Sfl1p—specifically required for regulating pseudohyphal development in S. cerevisiae56 were not found in Neurospora. Conversely, Neurospora possesses a gene with similarity to a transcription factor necessary for hyphal growth in C. albicans36 (Efg1). This transcription factor is not required for pseudohyphal growth in C. albicans, nor is the homologous protein in S. cerevisiae (Phd1p)56. More study of the complex pathways underlying these modes of growth is required. Nonetheless, these data clarify in part the distinctions and similarities between the signalling pathways and regulatory components of hyphal and pseudohyphal growth.
The macroconidiation pathway differs from that in A. nidulans
Macroconidia are asexual spores common to filamentous fungi but absent from yeast5,57. Components of the macroconidiation pathway have been identified in both Neurospora and the filamentous fungus A. nidulans, and known upstream signalling proteins seem to be conserved in both species58. In contrast, there is little conservation of downstream components between the two fungi. In Neurospora, the acon-2, acon-3, fld and fl genes are essential for conidiation5, whereas in A. nidulans, the FlbC, FlbD, BrlA, AbaA and WetA gene products are required. Our analysis of the genome sequence revealed that Neurospora possesses no FlbC, BrlA or AbaA homologues, and a protein with only very weak similarity to approximately 100 amino acids at the carboxy terminus of WetA. These data make clear that the molecular machinery underlying macroconidiation in Neurospora differs significantly from that in A. nidulans.
Secondary metabolism
The fungal kingdom produces a vast array of small, bioactive compounds termed secondary metabolites that are best known for their roles as pigments, antibiotics and mycotoxins. With the exception of carotenoid and melanin pigment synthesis, Neurospora has not been shown to possess secondary metabolism. It was thus surprising that the Neurospora genome sequence revealed a number of putative genes for secondary metabolite production.
Non-ribosomal peptide synthetases
Three predicted non-ribosomal peptide synthetase (NRPS) genes and one NRPS-related gene were identified in the Neurospora genome sequence (Fig. 6). By phylogenomic analysis, one NRPS gene is orthologous to an Aureobasidium pullulans NRPS. The most closely related NRPS of known function is sid2 of Ustilago maydis, which is responsible for production of hydroxamate siderophores59. The remaining two are of unknown function, although one is orthologous to an NRPS in Magnaporthe grisea, and the other is orthologous to an NRPS found in all other filamentous ascomycetes with genome sequence (see Methods). The NRPS-related gene shares 66% amino acid identity with the CPS1 gene product that contributes to the virulence of Cochliobolus heterostrophus, C. victoriae and Gibberella zeae60.
Polyketide synthases
Seven polyketide synthase (PKS) genes were identified in the Neurospora genome, which could be classified into three groups on the basis of domain structure (Fig. 6). The first class contains genes similar to DHN-melanin PKS genes of the fungi Exophila dermatitidis61, Colletotrichum lagenarium62 and Alternaria alternata63. Sequence identity to numerous expressed sequence tag (EST) sequences from sexual and perithecial libraries suggest a role in melanin pigment synthesis during sexual development64. The genes in the second class are similar in structure to several fungal PKSs, including the Aspergillus terreus lovF gene required for lovastatin synthesis. The genes in the third class resemble other fungal genes, including the A. terreus lovB gene, which is also required for lovastatin synthesis.
Diterpene metabolism
Diterpenes comprise a diverse group of compounds, primarily in plants and fungi, with roles in defence, pathogenicity and regulation of plant growth. The genome sequence revealed several genes associated with diterpene biosynthesis in other organisms, including a terpene synthase, several genes related to gibberellin oxidases, and a member of the cytochrome P450 mono-oxygenase gene family. These genes include at least one member of each of the three enzyme classes required for the biosynthesis of gibberellic acid. Gibberellic acid, a normal growth regulator in plants, was first identified as a metabolic product of the plant pathogen Gibberella fujikuroi, a relative of Neurospora that causes ‘foolish seedling’ disease in rice65. The presence of these genes in Neurospora suggests that many components necessary for gibberellic acid production were present in the ancestors of Neurospora and G. fujikuroi.
We speculate that the secondary metabolism genes identified may have roles in morphogenesis and chemotropism66, interspecies communication and possibly even chemical defence. The identification of these genes in Neurospora suggests that apparent major differences in lifestyles among related fungi, such as pathogenicity, may derive in part from minor modifications of gene function and expression.
Plant pathogenicity and Neurospora
The ability to parasitize living plants is widespread throughout the fungal kingdom. Although Neurospora is a saprotroph (that is, it feeds on dead or decaying matter), the genome sequence contains numerous genes similar to those required for plant pathogenesis identified in fungal pathogens. In particular, a number of genes were identified that have no known function in other organisms except in pathogenesis (Supplementary S8). Neurospora also possesses a wide range of extracellular enzymes capable of digesting plant cell wall polymers, although there is no clear cutinase homologue. Cutin is one of the main layers protecting the epidermis of the leaves of plants, and many, but not all, plant pathogens have cutinase activity. Neurospora has a wide range of cytochrome P450 enzymes that are important in some host–pathogen systems for detoxification of plant anti-fungal compounds. In addition, a large number of identified ABC (ATP-binding cassette) and MFS drug efflux systems could have a role in combating toxic plant compounds. The capability to form secondary metabolite members of the PKS, NRPS and terpenoid families, as described above, is present. Also, Neurospora contains all signal transduction components implicated in ascomycete pathogenesis that have been described so far. Thus, although Neurospora is not known to be a pathogen, the genome sequence has revealed many genes with similarity to those required for pathogenesis.
Discussion
Although Neurospora has been studied intensely for over 70 years, the analysis of the genome sequence has provided many new insights into a variety of cellular processes, including cell signalling, growth and differentiation, secondary metabolism and genome defence. The analysis has also uncovered surprising similarities between the saprotrophic Neurospora and pathogenic fungi, providing a new perspective on the molecular underpinnings of these lifestyles. Finally, the genome sequence has revealed the remarkable impact of RIP on the evolution of genes in Neurospora. Recent reports indicating the apparent presence of RIP in other fungi67,68 broaden the implications of our findings. The apparent lack of functional gene duplication in Neurospora provides a unique opportunity to study other modes of evolution in this experimentally tractable organism.
The genome sequence of Neurospora provides only a first glimpse into the genomic basis of the biological diversity of the filamentous fungi. Fungal genome sequences from the many ongoing69 and planned70 projects will expand this view as well as provide extraordinary opportunities for comparative analyses. This new era in fungal biology promises to yield insight into this important group of organisms, as well as to provide a deeper understanding of the fundamental cellular processes common to all eukaryotes.
Methods
Strain and growth conditions
Twenty 5-ml cultures of N. crassa wild-type strain N150 (74-OR23-1VA; Fungal Genetics Stock Center 2489) were grown on a shaker in Vogel's minimal medium5 for 3 days at 32 °C. Tissues were collected, freeze-dried overnight and DNA was extracted as previously described71. DNA from the 20 samples was mixed and used for library construction.
Sequencing and assembly
The genome was sequenced by the WGS method. Plasmid (4-kb inserts) and fosmid (40-kb inserts) libraries were generated as described at http://www-genome.wi.mit.edu/. Jumping clone (subclone) libraries with 50-kb inserts were generated as described elsewhere72. Neurospora cosmid and BAC clones were obtained from previously constructed libraries11,21. Sequencing methods for all clone types are described at http://www-genome.wi.mit.edu/. All inserts were sequenced from both ends to generate paired reads. The sequence coverage generated is shown in Supplementary Information. The sequence was assembled using Arachne8. Finished sequence from linkage groups II and V was provided by MIPS and is available at http://mips.gsf.de/proj/neurospora/.
Annotation and analysis
We annotated the Neurospora genome using the Calhoun annotation system. The genome sequence was searched against the public protein databases using BLASTX with a threshold value of E ≤ 10-5. Genes were predicted using a combination of FGENESH, FGENESH + and Genewise (Supplementary Information). The gene calling programs were validated against a test set of 191 previously characterized Neurospora proteins. Predicted genes were validated against ESTs aligned to the genome using SIM4. All predicted genes were searched against the PFAM set of hidden Markov models using the HMMER program and the public protein databases using BLASTP. Transfer RNAs were identified using the tRNAScan-SE program. Multigene families were constructed by searching each annotated gene against every other gene using BLASTP, requiring matches with E ≤ 10-5 over 60% of the longer gene length, and clustering genes based on single linkage transitive closure. Repeat sequences were detected by searching the genome sequence against itself using CrossMatch, filtering for alignments longer than 200 bp in length, and clustering pairs based on region overlap. Relics of RIP-mutated mobile elements were annotated by manual inspection.
The tree of MFS sugar transporters was created by aligning amino acid sequences using ClustalW, manually trimming to remove ambiguously aligned regions, and applying a neighbour-joining algorithm using PAUP*. RIP-mutated regions were detected by calculating one or both of two different dinucleotide ratios for sequence regions15. Regions with TpA/ApT > 2 or (CpA + TpG)/(ApC + GpT) < 0.7 were predicted as RIP-mutated. Prediction of RIP sequence across the genome used only the TpA/ApT ratio, whereas the analysis of coding sequences used both (with a positive prediction by either measure taken as a prediction of RIP). RIP simulations were implemented in Matlab and were based on parameters derived from Table 2 of ref. 16. The simulation code is available on request. During each round of simulated RIP, every cytosine-containing dinucleotide on one strand (selected with equal probabilities) was mutated according to the probabilities: (CA = 0.3, CT = 0.05, CG = 0.01, CC = 0.009). DNA methylation was predicted by calculating the proportion of plasmid reads overlapping 1-kb windows from both the methylation-tolerant and methylation-intolerant libraries. Regions with greater than 70% of reads derived from the methylation-tolerant library were predicted to be methylated. Specificity was estimated as described in the text. Methylation was experimentally assessed using Southern analyses as described elsewhere30. Sensitivity was estimated by testing 19 repetitive and RIP-mutated 1-kb regions that were not predicted to be methylated. Of the 19 regions, 14 were in fact methylated. Thus the data provide good specificity but poor sensitivity.
Predicted RNA-silencing genes were aligned with homologues from plants, animals and other fungi using T-Coffee v1.37. C-terminal and amino-terminal regions of low homology were removed and the sequences realigned until alignments started and stopped at regions of unambiguous similarity. Both neighbour-joining trees, using ClustalX, and maximum posterior probability trees, using MrBayes 2.01, were generated and analysed.
Analysis of predicted non-ribosomal peptide synthetases and polyketide synthases made use of genome data for C. heterostrophus, Botryotinia fuckeliana, G. verticillioides and G. zeae provided by the Torrey Mesa Research Institute/Syngenta. Additional details, analysis results and the genome sequence are available at http://www-genome.wi.mit.edu/.
References
Payen, A. (rapporteur) Extrait d'un rapport adressé à M. Le Maréchal Duc de Dalmatie, Ministre de la Guerre, Président du Conseil, sur une altération extraordinaire du pain de munition. Ann. Chim. Phys. 3 Ser. 9, 5–21 (1843)
Shear, C. L. & Dodge, B. O. Life histories and heterothallism of the red bread-mold fungi of the Monilia sitophila group. J. Agric. Res. 34, 1019–1042 (1927)
Lindegren, C. C. A six-point map of the sex chromosome of Neurospora crassa. J. Genet. 32, 243–256 (1936)
Beadle, G. W. & Tatum, E. L. Genetic control of biochemical reactions in Neurospora. Proc. Natl Acad. Sci. USA 27, 499–506 (1941)
Davis, R. H. Neurospora: Contributions of a Model Organism (Oxford Univ. Press, New York, 2000)
Davis, R. H. & Perkins, D. D. Neurospora: a model of model microbes. Nature Rev. Genet. 3, 397–403 (2002)
Perkins, D. D., Radford, A. & Sachs, M. S. The Neurospora Compendium: Chromosomal Loci (Academic, San Diego, California, 2001)
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002)
Schulte, U., Becker, I., Mewes, H. W. & Mannhaupt, G. Large scale analysis of sequences from Neurospora crassa. J. Biotechnol. 94, 3–13 (2002)
Free, S. J., Rice, P. W. & Metzenberg, R. L. Arrangement of the genes coding for ribosomal ribonucleic acids in Neurospora crassa. J. Bacteriol. 137, 1219–1226 (1979)
Aign, V., Schulte, U. & Hoheisel, J. D. Hybridization-based mapping of Neurospora crassa linkage groups II and V. Genetics 157, 1015–1020 (2001)
Selker, E. U., Cambareri, E. B., Jensen, B. C. & Haack, K. R. Rearrangement of duplicated DNA in specialized cells of Neurospora. Cell 51, 741–752 (1987)
Selker, E. U. Premeiotic instability of repeated sequences in Neurospora crassa. Annu. Rev. Genet. 24, 579–613 (1990)
Cambareri, E. B., Jensen, B. C., Schabtach, E. & Selker, E. U. Repeat-induced G-C to A-T mutations in Neurospora. Science 244, 1571–1575 (1989)
Margolin, B. S. et al. A methylated Neurospora 5S rRNA pseudogene contains a transposable element inactivated by repeat-induced point mutation. Genetics 149, 1787–1797 (1998)
Watters, M. K., Randall, T. A., Margolin, B. S., Selker, E. U. & Stadler, D. R. Action of repeat-induced point mutation on both strands of a duplex and on tandem duplications of various sizes in Neurospora. Genetics 153, 705–714 (1999)
Cambareri, E. B., Singer, M. J. & Selker, E. U. Recurrence of repeat-induced point mutation (RIP) in Neurospora crassa. Genetics 127, 699–710 (1991)
Rountree, M. R. & Selker, E. U. DNA methylation inhibits elongation but not initiation of transcription in Neurospora crassa. Genes Dev. 11, 2383–2395 (1997)
Irelan, J. T., Hagemann, A. T. & Selker, E. U. High frequency repeat-induced point mutation (RIP) is not associated with efficient recombination in Neurospora. Genetics 138, 1093–1103 (1994)
Perkins, D. D., Margolin, B. S., Selker, E. U. & Haedo, S. D. Occurrence of repeat induced point mutation in long segmental duplications of Neurospora. Genetics 147, 125–136 (1997)
Kelkar, H. S. et al. The Neurospora crassa genome: cosmid libraries sorted by chromosome. Genetics 157, 979–990 (2001)
Hutchison, C. A. et al. Global transposon mutagenesis and a minimal Mycoplasma genome. Science 286, 2165–2169 (1999)
Ohno, S. Evolution by Gene Duplication (Springer, New York, 1970)
Selker, E. U. et al. Dispersed 5S RNA genes in N. crassa: structure, expression and evolution. Cell 24, 819–828 (1981)
Selker, E. U. et al. The methylated component of the Neurospora crassa genome. Nature 422, 893–897 (2003)
Kouzminova, E. & Selker, E. U. dim-2 encodes a DNA methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J. 20, 4309–4323 (2001)
Freitag, M., Williams, R. L., Kothe, G. O. & Selker, E. U. A cytosine methyltransferase homologue is essential for repeat-induced point mutation in Neurospora crassa. Proc. Natl Acad. Sci. USA 99, 8802–8807 (2002)
Russell, P. J., Rodland, K. D., Rachlin, E. M. & McCloskey, J. A. Differential DNA methylation during the vegetative life cycle of Neurospora crassa. J. Bacteriol. 169, 2902–2905 (1987)
Foss, H. M., Roberts, C. J., Claeys, K. M. & Selker, E. U. Abnormal chromosome behavior in Neurospora mutants defective in DNA methylation. Science 262, 1737–1741 (1993)
Selker, E. U., Fritz, D. Y. & Singer, M. J. Dense nonsymmetrical DNA methylation resulting from repeat-induced point mutation in Neurospora. Science 262, 1724–1728 (1993)
Singer, M. J., Marcotte, B. A. & Selker, E. U. DNA methylation associated with repeat-induced point mutation in Neurospora crassa. Mol. Cell. Biol. 15, 5586–5597 (1995)
Wolffe, A. P. & Matzke, M. A. Epigenetics: regulation through repression. Science 286, 481–486 (1999)
Zamore, P. D. Ancient pathways programmed by small RNAs. Science 296, 1265–1269 (2002)
Hutvagner, G. & Zamore, P. D. RNAi: nature abhors a double-strand. Curr. Opin. Genet. Dev. 12, 225–232 (2002)
Catalanotto, C., Azzalin, G., Macino, G. & Cogoni, C. Involvement of small RNAs and role of the qde genes in the gene silencing pathway in Neurospora. Genes Dev. 16, 790–795 (2002)
Aramayo, R. & Metzenberg, R. L. Meiotic transvection in fungi. Cell 86, 103–113 (1996)
Shiu, P. K., Raju, N. B., Zickler, D. & Metzenberg, R. L. Meiotic silencing by unpaired DNA. Cell 107, 905–916 (2001)
Shiu, P. K. & Metzenberg, R. L. Meiotic silencing by unpaired DNA. Properties, regulation and suppression. Genetics 161, 1483–1495 (2002)
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002)
Lee, D., Pratt, R. J., McLaughlin, M., Aramayo, R. An Argonaute-like protein is required for meiotic silencing. Genetics (in the press)
Linden, H., Ballario, P., Arpaia, G. & Macino, G. Seeing the light: news in Neurospora blue light signal transduction. Adv. Genet. 41, 35–54 (1999)
Devlin, P. F. & Kay, S. A. Circadian photoperception. Annu. Rev. Physiol. 63, 677–694 (2001)
Gustin, M. C., Albertyn, J., Alexander, M. & Davenport, K. MAP kinase pathways in the yeast Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 62, 1264–1300 (1998)
Pott, G. B., Miller, T. K., Bartlett, J. A., Palas, J. S. & Selitrennikoff, C. P. The isolation of FOS-1, a gene encoding a putative two-component histidine kinase from Aspergillus fumigatus. Fungal Genet. Biol. 31, 55–67 (2000)
Dohlman, H. G., Thorner, J., Caron, M. G. & Lefkowitz, R. J. Model systems for the study of seven-transmembrane-segment receptors. Annu. Rev. Biochem. 60, 653–688 (1991)
Klein, P. S. et al. A chemoattractant receptor controls development in Dictyostelium discoideum. Science 241, 1467–1472 (1988)
Josefsson, L. G. & Rask, L. Cloning of a putative G-protein-coupled receptor from Arabidopsis thaliana. Eur. J. Biochem. 249, 415–420 (1997)
Aubry, L. & Firtel, R. Integration of signalling networks that regulate Dictyostelium differentiation. Annu. Rev. Cell Dev. Biol. 15, 469–517 (1999)
Ivey, F. D., Yang, Q. & Borkovich, K. A. Positive regulation of adenylyl cyclase activity by a Gαi homolog in Neurospora crassa. Fungal Genet. Biol. 26, 48–61 (1999)
Gadd, G. M. in The Growing Fungus (eds Gow, N. A. R. & Gadd, G. M.) 183–210 (Chapman & Hall, London, 1994)
Bootman, M. D. et al. Calcium signalling—an overview. Semin. Cell Dev. Biol. 12, 3–10 (2001)
Lakin-Thomas, P. L. Effects of inositol starvation on the levels of inositol phosphates and inositol lipids in Neurospora crassa. Biochem. J. 292, 805–811 (1993)
Silverman-Gavrila, L. B. & Lew, R. R. Regulation of the tip-high [Ca2+] gradient in growing hyphae of the fungus Neurospora crassa. Eur. J. Cell Biol. 80, 379–390 (2001)
Cornelius, G., Gebauer, G. & Techel, D. Inositol trisphosphate induces calcium release from Neurospora crassa vacuoles. Biochem. Biophys. Res. Commun. 162, 852–856 (1989)
Gimeno, C. J., Ljungdahl, P. O., Styles, C. A. & Fink, G. R. Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell 68, 1077–1090 (1992)
Lengeler, K. B. et al. Signal transduction cascades regulating fungal development and virulence. Microbiol. Mol. Biol. Rev. 64, 746–785 (2000)
Adams, T. H., Wieser, J. K. & Yu, J. H. Asexual sporulation in Aspergillus nidulans. Microbiol. Mol. Biol. Rev. 62, 35–54 (1998)
Ivey, F. D., Kays, A. M. & Borkovich, K. A. Shared and independent roles for a Gαi protein and adenylyl cyclase in regulating development and stress responses in Neurospora crassa. Eukaryot. Cell 1, 634–642 (2002)
Yuan, W., Gentil, G., Budde, A. & Leong, S. Characterization of the Ustilago maydis sid2 gene encoding a multidomain peptide synthetase in the ferrichrome biosynthetic gene cluster. J. Bacteriol. 183, 4040–4051 (2001)
Lu, S.-W., Yoder, O. & Turgeon, B. G. Highly conserved virulence-related CPS1 homologs from plant and human pathogenic fungi. Fungal Genet. News. 48, A246 (2001)
Feng, B. et al. Molecular cloning and characterization of WdPKS1, a gene involved in dihydroxynaphthalene melanin biosynthesis and virulence in Wangiella (Exophiala) dermatitidis. Infect. Immun. 69, 1781–1794 (2001)
Takano, Y. et al. Structural analysis of PKS1, a polyketide synthase gene involved in melanin biosynthesis in Colletotrichum lagenarium. Mol. Gen. Genet. 249, 162–167 (1995)
Takano, Y., Kubo, Y., Kawamura, C., Tsuge, T. & Furusawa, I. The Alternaria alternata melanin biosynthesis gene restores appressorial melanization and penetration of cellulose membranes in the melanin-deficient albino mutant of Colletotrichum lagenarium. Fungal Genet. Biol. 21, 131–140 (1997)
Nelson, M. A. et al. Expressed sequences from conidial, mycelial, and sexual stages of Neurospora crassa. Fungal Genet. Biol. 21, 348–363 (1997)
Kurosawa, E. Experimental studies on the nature of the substance excreted by the ‘bakanae’ fungus. Trans. Nat. Hist. Soc. Formosa 16, 213–227 (1926) (in Japanese)
Vigfusson, N. & Cano, R. Artificial induction of the sexual cycle of Neurospora crassa. Nature 249, 383–385 (1974)
Graia, F. et al. Genome quality control: RIP (repeat-induced point mutation) comes to Podospora. Mol. Microbiol. 40, 586–595 (2001)
Ikeda, K. et al. Repeat-induced point mutation (RIP) in Magnaporthe grisea: implications for its sexual cycle in the natural field context. Mol. Microbiol. 45, 1355–1364 (2002)
TIGR Microbial Database 〈http://www.tigr.org/tdb/mdb/mdbinprogress.html〉 (2003).
Fungal Genome Initiative 〈http://www-genome.wi.mit.edu/seq/fgi/〉 (2002).
Luo, Z., Freitag, M. & Sachs, M. S. Translational regulation in response to changes in amino acid availability in Neurospora crassa. Mol. Cell. Biol. 15, 5235–5245 (1995)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Pickford, A., Braccini, L., Macino, G. & Cogoni, C. The QDE-3 homologue RecQ-2 co-operates with QDE-3 in DNA repair in Neurospora crassa. Curr. Genet. 42, 220–227 (2003)
Acknowledgements
The authors would like to thank J. Arnold, H. Inoue, G. Turner, B. Bowman, P. Harriman and P. Youderian for their support. We also thank Lion Bioscience, V. Aign and J. Hoheisel for making available the BAC libraries used during sequencing; M. Karasz for developing the web pages hosting the Neurospora Interpro analysis at MIPS; the Torrey Mesa Research Institute/Syngenta for providing genome data for C. heterostrophus, B. fuckeliana, G. verticillioides and G. zeae; and all members of the Whitehead Institute/MIT Center for Genome Research sequencing group. Funding for the Neurospora genome project was provided by the National Science Foundation. Additional funding was provided by the Deutsche Forschungsgemeinschaft, The Israel Science Foundation and the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2003_BFnature01554_MOESM7_ESM.doc
Figure S6.1: The proportion of genes in multigene families for selected sequenced eukaryotes and prokaryotes (DOC 25 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Galagan, J., Calvo, S., Borkovich, K. et al. The genome sequence of the filamentous fungus Neurospora crassa. Nature 422, 859–868 (2003). https://doi.org/10.1038/nature01554
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01554
This article is cited by
-
Unveiling a classical mutant in the context of the GH3 β-glucosidase family in Neurospora crassa
AMB Express (2024)
-
The transcriptional factor Clr-5 is involved in cellulose degradation through regulation of amino acid metabolism in Neurospora crassa
BMC Biotechnology (2023)
-
The cell functions of phospholipase C-1, Ca2+/H+ exchanger-1, and secretory phospholipase A2 in tolerance to stress conditions and cellulose degradation in Neurospora crassa
Archives of Microbiology (2023)
-
Diversity of biologically active secondary metabolites in the ascomycete order Sordariales
Mycological Progress (2022)
-
Neurospora crassa is a potential source of anti-cancer agents against breast cancer
Breast Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.