Abstract
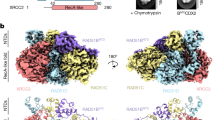
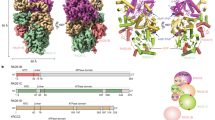
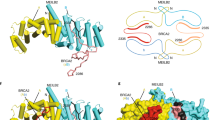
The breast cancer susceptibility protein BRCA2 controls the function of RAD51, a recombinase enzyme, in pathways for DNA repair by homologous recombination. We report here the structure of a complex between an evolutionarily conserved sequence in BRCA2 (the BRC repeat) and the RecA-homology domain of RAD51. The BRC repeat mimics a motif in RAD51 that serves as an interface for oligomerization between individual RAD51 monomers, thus enabling BRCA2 to control the assembly of the RAD51 nucleoprotein filament, which is essential for strand-pairing reactions during DNA recombination. The RAD51 oligomerization motif is highly conserved among RecA-like recombinases, highlighting a common evolutionary origin for the mechanism of nucleoprotein filament formation, mirrored in the BRC repeat. Cancer-associated mutations that affect the BRC repeat disrupt its predicted interaction with RAD51, yielding structural insight into mechanisms for cancer susceptibility.
This is a preview of subscription content, access via your institution
Access options





Similar content being viewed by others
References
Nathanson, K. N., Wooster, R. & Weber, B. L. Breast cancer genetics: what we know and what we need. Nature Med. 7, 552–556 (2001)
Bork, P., Blomberg, N. & Nilges, M. Internal repeats in the BRCA2 protein sequence. Nature Genet. 13, 22–23 (1996)
Bignell, G., Micklem, G., Stratton, M. R., Ashworth, A. & Wooster, R. The BRC repeats are conserved in mammalian BRCA2 proteins. Hum. Mol. Genet. 6, 53–58 (1997)
Warren, M. et al. Structural analysis of the chicken BRCA2 gene facilitates identification of functional domains and disease causing mutations. Hum. Mol. Genet. 11, 841–851 (2002)
Wong, A. K. C., Pero, R., Ormonde, P. A., Tavtigian, S. V. & Bartel, P. L. RAD51 interacts with the evolutionarily conserved BRC motifs in the human breast cancer susceptibility gene brca2. J. Biol. Chem. 272, 31941–31944 (1997)
Chen, P. L. et al. The BRC repeats in BRCA2 are critical for RAD51 binding and resistance to methyl methanesulfonate treatment. Proc. Natl Acad. Sci. USA 95, 5287–5292 (1998)
Chen, C. F., Chen, P. L., Zhong, Q., Sharp, Z. D. & Lee, W. H. Expression of BRC repeats in breast cancer cells disrupts the BRCA2-Rad51 complex and leads to radiation hypersensitivity and loss of G(2)/M checkpoint control. J. Biol. Chem. 274, 32931–32935 (1999)
Scully, R. & Livingston, D. M. In search of the tumour-suppressor functions of BRCA1 and BRCA2. Nature 408, 429–432 (2000)
Venkitaraman, A. R. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell 108, 171–182 (2002)
Chen, G., Yuan, S. S., Liu, W. et al. Radiation-induced assembly of Rad51 and Rad52 recombination complex requires ATM and c-Abl. J. Biol. Chem. 274, 12748–12752 (1999)
Yu, V. P. C. C. et al. Gross chromosomal rearrangements and genetic exchange between non-homologous chromosomes following BRCA2 inactivation. Genes Dev. 14, 1400–1406 (2000)
Moynahan, M. E., Pierce, A. J. & Jasin, M. BRCA2 is required for homology-directed repair of chromosomal breaks. Mol. Cell 7, 263–272 (2001)
Lim, D. S. & Hasty, P. A mutation in mouse rad51 results in an early embryonic lethal that is suppressed by a mutation in p53. Mol. Cell Biol. 16, 7133–7143 (1996)
Tsuzuki, T. et al. Targeted disruption of the Rad51 gene leads to lethality in embryonic mice. Proc. Natl Acad. Sci. USA 93, 6236–6240 (1996)
Patel, K. J. et al. Involvement of Brca2 in DNA repair. Mol. Cell 1, 347–357 (1998)
Tutt, A. et al. Absence of brca2 causes genome instability by chromosome breakage and loss associated with centrosome amplification. Curr. Biol. 9, 1107–1110 (1999)
Davies, A. A. et al. Role of BRCA2 in control of the RAD51 recombination and DNA repair protein. Mol. Cell 7, 273–282 (2001)
Story, R. M., Weber, I. T. & Steitz, T. A. The structure of the E. coli recA protein monomer and polymer. Nature 355, 318–325 (1992)
Sibanda, B. L. & Thornton, J. M. Beta-hairpin families in globular proteins. Nature 316, 170–174 (1985)
Story, R. M. & Steitz, T. A. Structure of the recA protein-ADP complex. Nature 355, 374–376 (1992)
Sung, P. & Robberson, D. L. DNA strand exchange mediated by a RAD51-ssDNA nucleoprotein filament with polarity opposite to that of RecA. Cell 82, 453–461 (1995)
Baumann, P., Benson, F. E. & West, S. C. Human Rad51 protein promotes ATP-dependent homologous pairing and strand transfer reactions in vitro. Cell 87, 757–766 (1996)
Ogawa, T., Yu, X., Shinohara, A. & Egelman, E. H. Similarity of the yeast RAD51 filament to the bacterial RecA filament. Science 259, 1896–1899 (1993)
Yu, X., Jacobs, S. A., West, S. C., Ogawa, T. & Egelman, E. H. Domain structure and dynamics in the helical filaments formed by RecA and Rad51 on DNA. Proc. Natl Acad. Sci. USA 98, 8419–8424 (2001)
Haaf, T., Golub, E. I., Reddy, G., Radding, C. M. & Ward, D. C. Nuclear foci of mammalian Rad51 recombination protein in somatic cells after DNA damage and its localization in synaptonemal complexes. Proc. Natl Acad. Sci. USA 92, 2298–2302 (1995)
Scully, R. et al. Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 88, 265–275 (1997)
Peranen, J., Rikkonen, M., Hyvonen, M. & Kaariainen, L. T7 vectors with modified T7lac promoter for the expression of proteins in E. coli. Anal. Biochem. 236, 371–373 (1996)
Boggon, T. J. & Shapiro, L. Screening for phasing atoms in protein crystallography. Structure Fold Des. 8, R143–R149 (2000)
Deacon, A. M., Weeks, C. M., Miller, R. & Ealick, S. E. The Shake-and-Bake structure determination of triclinic lysozyme. Proc. Natl Acad. Sci. USA 95, 9284–9289 (1998)
La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement in the MIR and MAD methods. Methods Enzymol. 276, 472–494 (1997)
Perrakis, A., Morris, R. & Lamzin, V. S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 54, 905–921 (1997)
Bruenger, A. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991)
Merritt, E. A. & Bacon, D. J. Raster3D photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997)
Acknowledgements
We thank A. Gupta for early work on the purification of the RAD51–BRCA2 complex; A. Thompson for technical assistance at the ESRF beamline ID29; M. Symmons for help with the figures; and R. Laskey for comments on this manuscript. This research was supported in the laboratory of T.L.B. by the Wellcome Trust, and in the laboratory of A.R.V. by the Medical Research Council and Cancer Research UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Pellegrini, L., Yu, D., Lo, T. et al. Insights into DNA recombination from the structure of a RAD51–BRCA2 complex. Nature 420, 287–293 (2002). https://doi.org/10.1038/nature01230
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature01230
This article is cited by
-
In Silico design and characterization of RAD51 protein inhibitors targeting homologous recombination repair for cancer therapy
Genome Instability & Disease (2023)
-
Repair of DNA double-strand breaks in plant meiosis: role of eukaryotic RecA recombinases and their modulators
Plant Reproduction (2023)
-
ZFP281-BRCA2 prevents R-loop accumulation during DNA replication
Nature Communications (2022)
-
Distinct pathways of homologous recombination controlled by the SWS1–SWSAP1–SPIDR complex
Nature Communications (2021)
-
A complex of BRCA2 and PP2A-B56 is required for DNA repair by homologous recombination
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.