Abstract
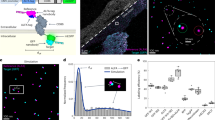
Protein folding is inherently a heterogeneous process because of the very large number of microscopic pathways that connect the myriad unfolded conformations to the unique conformation of the native structure. In a first step towards the long-range goal of describing the distribution of pathways experimentally, Förster resonance energy transfer1 (FRET) has been measured on single, freely diffusing molecules2,3,4. Here we use this method to determine properties of the free-energy surface for folding that have not been obtained from ensemble experiments. We show that single-molecule FRET measurements of a small cold-shock protein expose equilibrium collapse of the unfolded polypeptide and allow us to calculate limits on the polypeptide reconfiguration time. From these results, limits on the height of the free-energy barrier to folding are obtained that are consistent with a simple statistical mechanical model, but not with the barriers derived from simulations using molecular dynamics. Unlike the activation energy, the free-energy barrier includes the activation entropy and thus has been elusive to experimental determination for any kinetic process in solution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Van Der Meer, B. W., Coker, G. III & Chen, S. S.-Y. Resonance Energy Transfer. Theory and Data (VHC, New York, 1994)
Jia, Y. W. et al. Folding dynamics of single GCN4 peptides by fluorescence energy transfer confocal microscopy. Chem. Phys. 247, 69–83 (1999)
Talaga, D. S. et al. Dynamics and folding of single two-stranded coiled-coil peptides studied by fluorescent energy transfer confocal microscopy. Proc. Natl Acad. Sci. USA 97, 13021–13026 (2000)
Deniz, A. A. et al. Single-molecule protein folding: diffusion fluorescence resonance energy transfer studies of the denaturation of chymotrypsin inhibitor 2. Proc. Natl Acad. Sci. USA 97, 5179–5184 (2000)
Perl, D. et al. Conservation of rapid two-state folding in mesophilic, thermophilic and hyperthermophilic cold shock proteins. Nature Struct. Biol. 5, 229–235 (1998)
Mandelkern, L. in Poly-α-amino Acids. Protein Models for Conformational Studies (ed. Fasman, G. D.) 675–724 (Marcel Dekker, New York, 1967)
Jacob, J., Baker, B., Bryant, R. G. & Cafiso, D. S. Distance estimates from paramagnetic enhancements of nuclear relaxation in linear and flexible model peptides. Biophys. J. 77, 1086–1092 (1999)
Stryer, L. & Haugland, R. P. Energy transfer: a spectroscopic ruler. Proc. Natl Acad. Sci. USA 58, 719–726 (1967)
Wassenberg, D., Welker, C. & Jaenicke, R. Thermodynamics of the unfolding of the cold-shock protein from Thermotoga maritima. J. Mol. Biol. 289, 187–193 (1999)
Fersht, A. Structure and Mechanism in Protein Science: a Guide to Enzyme Catalysis and Protein Folding (W. H. Freeman, New York, 1999)
Alonso, D. O. V. & Dill, K. A. Solvent denaturation and stabilization of globular proteins. Biochemistry 30, 5974–5985 (1991)
Chan, C. K. et al. Submillisecond protein folding kinetics studied by ultrarapid mixing. Proc. Natl Acad. Sci. USA 94, 1779–1784 (1997)
Millett, I. S., Doniach, S. & Plaxco, K. W. Towards a taxonomy of the denatured state: small angle scattering studies of unfolded proteins. Adv. Prot. Chem. (in the press)
Allen, M. P. & Tildesley, D. J. Computer Simulation of Liquids 192 (Clarendon, Oxford, 1987)
Dahan, M. et al. Ratiometric measurement and identification of single diffusing molecules. Chem. Phys. 247, 85–106 (1999)
Deniz, A. A. et al. Single-pair fluorescence resonance energy transfer on freely diffusing molecules: observation of Förster distance dependence and subpopulations. Proc. Natl Acad. Sci. USA 96, 3670–3675 (1999)
Bryngelson, J. D. & Wolynes, P. G. Spin glasses and the statistical mechanics of protein folding. Proc. Natl Acad. Sci. USA 84, 7524–7528 (1987)
Bryngelson, J. D., Onuchic, J. N., Socci, N. D. & Wolynes, P. G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins Struct. Funct. Genet. 21, 167–195 (1995)
Dobson, C. M., Sali, A. & Karplus, M. Protein folding: a perspective from theory and experiment. Angew. Chem. Int. Edit. 37, 868–893 (1998)
Socci, N. D., Onuchic, J. N. & Wolynes, P. G. Diffusive dynamics of the reaction coordinate for protein folding funnels. J. Chem. Phys. 104, 5860–5868 (1996)
Klimov, D. K. & Thirumalai, D. Viscosity dependence of the folding rates of proteins. Phys. Rev. Lett. 79, 317–320 (1997)
Nymeyer, H., Socci, N. D. & Onuchic, J. N. Landscape approaches for determining the ensemble of folding transition states: Success and failure hinge on the degree of frustration. Proc. Natl Acad. Sci. USA 97, 634–639 (2000)
Kramers, H. A. Brownian motion in a field of force and the diffusion model of chemical reactions. Physica 7, 284–304 (1940)
Lapidus, L. J., Steinbach, P. J., Eaton, W. A., Szabo, A. & Hofrichter, J. Effects of chain stiffness on the dynamics of loop formation in polypeptides. J. Phys. Chem. B (in the press)
Jäger, M., Nguyen, H., Crane, J. C., Kelley, J. W. & Gruebele, M. The folding mechanism of a β-sheet: the WW domain. J. Mol. Biol. 311, 373–393 (2001)
Hagen, S. J., Hofrichter, J. & Eaton, W. A. The rate of intrachain diffusion of unfolded cytochrome c. J. Phys. Chem. B 101, 2352–2365 (1997)
Portman, J. J., Takada, S. & Wolynes, P. G. Microscopic theory of protein folding rates. II. Local reaction coordinates and chain dynamics. J. Chem. Phys. 114, 5082–5096 (2001)
Muñoz, V. & Eaton, W. A. A simple model for calculating the kinetics of protein folding from three-dimensional structure. Proc. Natl Acad. Sci. USA 96, 11311–11316 (1999)
Shea, J. E. & Brooks, C. L. From folding theories to folding proteins: A review and assessment of simulation studies of protein folding and unfolding. Annu. Rev. Phys. Chem. 52, 499–535 (2001)
Reid, K. L., Rodriguez, H. M., Hillier, B. J. & Gregoret, L. M. Stability and folding properties of a model β-sheet protein, Escherichia coli CspA. Protein Sci. 7, 470–479 (1998)
Kremer, W. et al. Solution NMR structure of the cold-shock protein from the hyperthermophilic bacterium Thermotoga maritima. Eur. J. Biochem. 268, 2527–2539 (2001)
Acknowledgements
We thank A. Szabo and I. Gopich for suggestions and guidance on theoretical issues; S. Doniach, J. Hofrichter and G. Hummer for discussion and comments on the manuscript; L. Davis and R. Zare for advice regarding the single-molecule instrument; and L. Pannell for mass spectroscopy measurements. B.S. was supported by an Emmy Noether fellowship from the Deutsche Forschungsgemeinschaft.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Schuler, B., Lipman, E. & Eaton, W. Probing the free-energy surface for protein folding with single-molecule fluorescence spectroscopy. Nature 419, 743–747 (2002). https://doi.org/10.1038/nature01060
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01060
This article is cited by
-
Folding pathway of a discontinuous two-domain protein
Nature Communications (2024)
-
FRETpredict: a Python package for FRET efficiency predictions using rotamer libraries
Communications Biology (2024)
-
An automated single-molecule FRET platform for high-content, multiwell plate screening of biomolecular conformations and dynamics
Nature Communications (2023)
-
Single-molecule FRET unmasks structural subpopulations and crucial molecular events during FUS low-complexity domain phase separation
Nature Communications (2023)
-
Direct observation of chaperone-modulated talin mechanics with single-molecule resolution
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.