Abstract
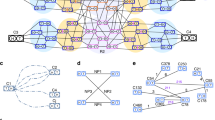
A physical map of a genome is an essential guide for navigation, allowing the location of any gene or other landmark in the chromosomal DNA. We have constructed a physical map of the mouse genome that contains 296 contigs of overlapping bacterial clones and 16,992 unique markers. The mouse contigs were aligned to the human genome sequence on the basis of 51,486 homology matches, thus enabling use of the conserved synteny (correspondence between chromosome blocks) of the two genomes to accelerate construction of the mouse map. The map provides a framework for assembly of whole-genome shotgun sequence data, and a tile path of clones for generation of the reference sequence. Definition of the human–mouse alignment at this level of resolution enables identification of a mouse clone that corresponds to almost any position in the human genome. The human sequence may be used to facilitate construction of other mammalian genome maps using the same strategy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Fleischmann, R. D. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–512 (1995)
Churcher, C. et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome IX. Nature 387, 84–87 (1997)
The yeast genome directory. Nature 387 (suppl.), 5 (1997)
The C. elegans Sequencing Consortium Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998)
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000)
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Collins, J. & Hohn, B. Cosmids: a type of plasmid gene-cloning vector that is packageable in vitro in bacteriophage lambda heads. Proc. Natl Acad. Sci. USA 75, 4242–4246 (1978)
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl Acad. Sci. USA 89, 8794–8797 (1992)
Coulson, A., Sulston, J., Brenner, S. & Karn, J. Towards a physical map of the genome of the nematode Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 83, 7821–7825 (1986)
Olson, M. V. et al. Random-clone strategy for genomic restriction mapping in yeast. Proc. Natl Acad. Sci. USA 83, 7826–7830 (1986)
Bentley, D. R. et al. The physical maps for sequencing human chromosomes 1, 6, 9, 10, 13, 20 and X. Nature 409, 942–943 (2001)
McPherson, J. D. et al. A physical map of the human genome. Nature 409, 934–941 (2001)
Green, E. D. Strategies for the systematic sequencing of complex genomes. Nature Rev. Genet. 2, 573–583 (2001)
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999)
Waterston, R. & Sulston, J. E. The Human Genome Project: reaching the finish line. Science 282, 53–54 (1998)
Nadeau, J. H. & Taylor, B. A. Lengths of chromosomal segments conserved since divergence of man and mouse. Proc. Natl Acad. Sci. USA 81, 814–818 (1984)
DeBry, R. W. & Seldin, M. F. Human/mouse homology relationships. Genomics 33, 337–351 (1996)
Nadeau, J. H. & Sankoff, D. Counting on comparative maps. Trends Genet. 14, 495–501 (1998)
Thomas, J. W. et al. Comparative genome mapping in the sequence-based era: early experience with human chromosome 7. Genome Res. 10, 624–633 (2000)
Kim, J. et al. Homology-driven assembly of a sequence-ready mouse BAC contig map spanning regions related to the 46-Mb gene-rich euchromatic segments of human chromosome 19. Genomics 74, 129–141 (2001)
Osoegawa, K. et al. Bacterial artificial chromosome libraries for mouse sequencing and functional analysis. Genome Res. 10, 116–128 (2000)
Hardison, R. C., Oeltjen, J. & Miller, W. Long human–mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997)
Koop, B. F. et al. The human T-cell receptor TCRAC/TCRDC (Cα/Cδ) region: organization, sequence, and evolution of 97.6 kb of DNA. Genomics 19, 478–493 (1994)
Ashworth, L. K. et al. An integrated metric physical map of human chromosome 19. Nature Genet. 11, 422–427 (1995)
Kent, W. J. The BLAST-like alignment tool. Genome Res. 12, 656–664 (2002)
Copeland, N. G. et al. A genetic linkage map of the mouse: current applications and future prospects. Science 262, 57–66 (1993)
Dietrich, W. F. et al. A comprehensive genetic map of the mouse genome. Nature 380, 149–152 (1996)
Cai, W. W. et al. An SSLP marker-anchored BAC framework map of the mouse genome. Nature Genet. 29, 133–134 (2001)
Hudson, T. J. et al. A radiation hybrid map of mouse genes. Nature Genet. 29, 201–205 (2001)
Schuler, G. D. Sequence mapping by electronic PCR. Genome Res. 7, 541–550 (1997)
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
Ross, M. T., LaBrie, S., McPherson, J. P. & Stanton, V. P. in Current Protocols in Human Genetics (eds Dracopoli, N. C. et al.) 5.6.1–5.6.5 (Wiley, New York, 1999)
Puttagunta, R. et al. Comparative maps of human 19p13.3 and mouse chromosome 10 allow identification of sequences at evolutionary breakpoints. Genome Res. 10, 1369–1380 (2000)
Carver, E. A. & Stubbs, L. Zooming in on the human–mouse comparative map: genome conservation re-examined on a high-resolution scale. Genome Res. 7, 1123–1137 (1997)
Watkins-Chow, D. E. et al. Genetic mapping of 21 genes on mouse chromosome 11 reveals disruptions in linkage conservation with human chromosome 5. Genomics 40, 114–122 (1997)
Stubbs, L. et al. Detailed comparative map of human chromosome 19q and related regions of the mouse genome. Genomics 35, 499–508 (1996)
Dehal, P. et al. Human chromosome 19 and related regions in mouse: conservative and lineage-specific evolution. Science 293, 104–111 (2001)
Pletcher, M. T., Wiltshire, T., Cabin, D. E., Villanueva, M. & Reeves, R. H. Use of comparative physical and sequence mapping to annotate mouse chromosome 16 and human chromosome 21. Genomics 74, 45–54 (2001)
Oeltjen, J. C. et al. Large-scale comparative sequence analysis of the human and murine Bruton's tyrosine kinase loci reveals conserved regulatory domains. Genome Res. 7, 315–329 (1997)
Batzoglou, S., Pachter, L., Mesirov, J. P., Berger, B. & Lander, E. S. Human and mouse gene structure: comparative analysis and application to exon prediction. Genome Res. 10, 950–958 (2000)
Marra, M. A. et al. High throughput fingerprint analysis of large-insert clones. Genome Res. 7, 1072–1084 (1997)
Soderlund, C., Humphray, S., Dunham, A. & French, L. Contigs built with fingerprints, markers, and FPC V4. 7. Genome Res. 10, 1772–1787 (2000)
Zhao, S. et al. Mouse BAC ends quality assessment and sequence analyses. Genome Res. 11, 1736–1745 (2001)
Altschul, S. F. & Gish, W. Local alignment statistics. Methods Enzymol. 266, 460–480 (1996)
Nusbaum, C. et al. A YAC-based physical map of the mouse genome. Nature Genet. 22, 388–393 (1999)
Evans, E. P. in Genetic Variants of the Laboratory Mouse (eds Lyon, M. F., Rastan, S. & Brown, S. D. M.) 1446 (Oxford Univ. Press, New York, 1996)
Pletcher, M. T. et al. Chromosome evolution: the junction of mammalian chromosomes in the formation of mouse chromosome 10. Genome Res. 10, 1463–1467 (2000)
Martindale, D. W. et al. Comparative genomic sequence analysis of the Williams syndrome region (LIMK1-RFC2) of human chromosome 7q11.23. Mamm. Genome 11, 890–898 (2000)
DeSilva, U. et al. Generation and comparative analysis of approximately 3.3 Mb of mouse genomic sequence orthologous to the region of human chromosome 7q11.23 implicated in Williams syndrome. Genome Res. 12, 3–15 (2002)
Acknowledgements
The authors acknowledge the support of the Wellcome Trust, the National Institutes of Health and the US Department of Energy. We are grateful to the web team at the Sanger Institute for assistance with developing map displays, to P. Deloukas for RH map analysis, and E. Arnold-Berkowits, S. Lo, J. Gill and all present and past members of the Institute for Genomic Research BAC end sequencing team for the sequencing work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2002_BFnature00957_MOESM1_ESM.zip
This file contains a static version of the mouse fingerprint contig (FPC) map; it is an archive representation of the data at the time of publication. (Start with the file bac.1.0.html). A live, updated version of this data can be seen in CytoView at Ensembl (choose 'MapViewer'), and also at the NCBI (use the 'Jump to chr' box to select chromosome; if searching specific features, select 'cytoview' option to view the map). The FPC map was previous displayed in Ensembl before the availability of sequence covering most of the genome. Since then, Ensembl displays have switched to being sequence based, with the FPC data mapped onto it and visible through the CytoView interface. Both the sequence and the FPC map are being refined as the mouse genome is finished. (ZIP 3564 kb)
This file contains a static version of the map derived from the synteny between the mouse and human genomes (Fig. 5); it is an archive representation of the data at the time of publication. (Start with the file hm.1.1.html). A live, updated version of this data can be seen in SyntenyView at Ensembl.
Copies of these data, plus comparisons and discussion of genetic, RH and clone maps, and an interactive version of the synteny displays of Fig. 5, are available at the Wellcome Trust Sanger Institute. (ZIP 1541 kb)
Updated views of the map are available from the authors' websites (http://www.ensembl.org/Mus_musculus/cytoview and http://www.ncbi.nlm.nih.gov/genome/guide/mouse), as is an archive version for this publication, plus comparisons and discussion of genetic, RH and clone maps, and an interactive version of the synteny displays of Fig. 5 (http://www.sanger.ac.uk/Projects/M_musculus/publications/fpcmap-2002).
Rights and permissions
About this article
Cite this article
Gregory, S., Sekhon, M., Schein, J. et al. A physical map of the mouse genome. Nature 418, 743–750 (2002). https://doi.org/10.1038/nature00957
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature00957
This article is cited by
-
The plasticitome of cortical interneurons
Nature Reviews Neuroscience (2023)
-
Construction of stable mouse artificial chromosome from native mouse chromosome 10 for generation of transchromosomic mice
Scientific Reports (2021)
-
Features of the organization of bread wheat chromosome 5BS based on physical mapping
BMC Genomics (2018)
-
An alignment-free method to find and visualise rearrangements between pairs of DNA sequences
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.