Abstract
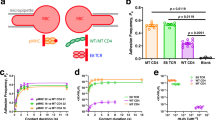
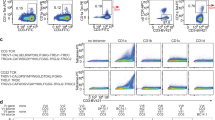
T cells probe a diverse milieu of peptides presented by molecules of the major histocompatibility complex (MHC) by using the T-cell receptor (TCR) to scan these ligands with high sensitivity and specificity1. Here we describe a physical basis for this scanning process by studying the residues involved in both the initial association and the stable binding of TCR to peptide–MHC, using the well-characterized TCR and peptide–MHC pair of 2B4 and MCC-IEk (moth cytochrome c, residues 88–103)2. We show that MHC contacts dictate the initial association, guiding TCR docking in a way that is mainly independent of the peptide. Subsequently, MCC-IEk peptide contacts dominate stabilization, imparting specificity and influencing T-cell activation by modulating the duration of binding. This functional subdivision of the peptide–MHC ligand suggests that a two-step process for TCR recognition facilitates the efficient scanning of diverse peptide–MHC complexes on the surface of cells and also makes TCRs inherently crossreactive towards different peptides bound by the same MHC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Germain, R. N. & Stefanova, I. The dynamics of T cell receptor signalling: complex orchestration and the key roles of tempo and cooperation. Annu. Rev. Immunol. 17, 467–522 (1999)
Davis, M. M. et al. Ligand recognition by αβ T cell receptors. Annu. Rev. Immunol. 16, 523–544 (1998)
Rudolph, M. G. & Wilson, I. A. The specificity of TCR/pMHC interaction. Curr. Opin. Immunol. 14, 52–65 (2002)
Fremont, D. H. et al. Structural basis of cytochrome c presentation by IEk. J. Exp. Med. 195, 1043–1052 (2002)
Baker, B. M., Turner, R. V., Gagnon, S. J., Wiley, D. C. & Biddison, W. E. Identification of a crucial energetic footprint on the α1 helix of human histocompatibility leukocyte antigen (HLA)-A2 that provides functional interactions for recognition by Tax peptide/HLA-A2-specific T cell receptors. J. Exp. Med. 193, 551–562 (2001)
Ehrich, E. W. et al. T cell receptor interactions with peptide/major histocompatibility complex (MHC) and superantigen/MHC ligands is dominated by antigen. J. Exp. Med. 178, 713–722 (1993)
Reay, P., Kantor, R. M. & Davis, M. M. Use of global amino acid replacements to define the requirements for MHC binding and T cell recognition of moth cytochrome c (93–103). J. Immunol. 152, 3946–3957 (1994)
Manning, T. C. et al. Alanine scanning mutagenesis of an αβ T cell receptor: mapping the energy of antigen recognition. Immunity 8, 413–425 (1998)
Lee, P. U., Churchill, H. R., Daniels, M., Jameson, S. C. & Kranz, D. M. Role of 2C T cell receptor residues in the binding of self- and allo-major histocompatibility complexes. J. Exp. Med. 191, 1355–1364 (2000)
Degano, M. et al. A functional hot spot for antigen recognition in a superagonist TCR/MHC complex. Immunity 12, 251–261 (2000)
Garcia, K. C. et al. Structural basis of plasticity in T cell receptor recognition of a self peptide–MHC antigen. Science 279, 1166–1172 (1998)
Kersh, G. J., Kersh, E. N., Fremont, D. H. & Allen, P. M. High- and low-potency ligands with similar affinities for the TCR: the importance of kinetics in TCR signaling. Immunity 9, 817–826 (1998)
Baker, B. M., Ding, Y.-H., Garboczi, D. N., Biddison, W. E. & Wiley, D. C. Structural, biochemical, and biophysical studies of HLA-A2/altered peptide ligands binding to viral-peptide-specific human T-cell receptors. Cold Spring Harb. Symp. Quant. Biol. 64, 235–241 (1999)
Fersht, A. R. Nucleation steps in protein folding. Curr. Opin. Struct. Biol. 7, 3–9 (1997)
Schreiber, G. & Fersht, A. R. Rapid electrostatically assisted association of proteins. Nature Struct. Biol. 3, 427–431 (1996)
Davis, S. J., Davies, E. A., Tucknott, M. G., Jones, E. Y. & van der Merwe, P. A. The role of charged residues mediating low affinity protein-protein recognition at the cell surface by CD2. Proc. Natl Acad. Sci. USA 95, 5490–5494 (1998)
Hare, B. J. et al. Structure, specificity, and CDR mobility of a class II restricted single-chain T cell receptor. Nature Struct. Biol. 6, 574–581 (1999)
Reiser, J. B. et al. A T cell receptor CDR3β loop undergoes conformational changes of unprecedented magnitude upon binding to a peptide/MHC class I complex. Immunity 16, 345–354 (2002)
Boniface, J. J., Reich, Z., Lyons, D. S. & Davis, M. M. Thermodynamics of T cell receptor binding to peptide-MHC: evidence for a general mechanism of molecular scanning. Proc. Natl Acad. Sci. USA 96, 11446–11451 (1999)
Willcox, B. E. et al. TCR binding to peptide-MHC stabilizes a flexible recognition interface. Immunity 10, 357–365 (1999)
Mason, D. A very high level of cross-reactivity is an essential feature of the T-cell receptor. Immunol. Today 19, 395–404 (1998)
Clackson, T. & Wells, J. A. A hot spot of binding energy in a hormone-receptor interface. Science 267, 383–386 (1995)
Bevan, M. J. In a radiation chimaera, host H-2 antigens determine immune responsiveness of donor cytotoxic cells. Nature 269, 417–418 (1977)
Zinkernagel, R. M. et al. On the thymus in the differentiation of ‘H-2 self-recognition’ by T cells: evidence for dual recognition? J. Exp. Med. 147, 882–896 (1978)
Zerrahn, J., Held, W. & Raulet, D. H. The MHC reactivity of the T cell repertoire prior to positive and negative selection. Cell 88, 627–636 (1997)
Hennecke, J. & Wiley, D. C. Structure of a complex of the human α/β T cell receptor (TCR) HA1. (7), influenze hemagglutinin peptide, and major histocompatibility complex class II molecule, HLA-DR4 (DRA*0101 and DRB1*0401): insight into TCR cross-restriction and alloreactivity. J. Exp. Med. 195, 571–581 (2002)
Luz, J. G. et al. Structural comparison of allogeneic and syngeneic T cell receptor-peptide-major histocompatibility complex complexes: a buried alloreactive mutation subtly alters peptide presentation substantially increasing Vβ interactions. J. Exp. Med. 195, 1175–1186 (2002)
Savage, P. A. & Davis, M. M. A kinetic window constricts the T cell receptor repertoire in the thymus. Immunity 14, 243–252 (2001)
Tanchot, C., Lemonnier, F. A., Pérarnau, B., Freitas, A. A. & Rocha, B. Differential requirements for survival and proliferation of CD8 naïve or memory T cells. Science 276, 2057–2058 (1997)
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991)
Acknowledgements
We thank J. Boniface, C. Gerke, S. Hedrick, D. Herschlag, J. Huppa, M. Krogsgaard, M. Kuhns, Z. Reich, R. Sciammas and members of the Davis lab for discussions and comments on the manuscript; B. Malissen and D. Fremont for communicating results before publication; N. Prado for assistance with TCR production; and A. Cochran and H. Lowman for use of their BIAcore 2000. L.C.W. is a postdoctoral fellow of the Cancer Research Fund of the Damon Runyon Walter Winchell Foundation. D.S.L. was a Howard Hughes Medical Institute predoctoral fellow. This work was supported by the NIH (M.M.D. and K.C.G.), the Howard Hughes Medical Institute (M.M.D.), the Multiple Sclerosis Society (K.C.G.), and the Cancer Research Institute (K.C.G.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Wu, L., Tuot, D., Lyons, D. et al. Two-step binding mechanism for T-cell receptor recognition of peptide–MHC. Nature 418, 552–556 (2002). https://doi.org/10.1038/nature00920
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature00920
This article is cited by
-
Facile repurposing of peptide–MHC-restricted antibodies for cancer immunotherapy
Nature Biotechnology (2023)
-
Structural understanding of T cell receptor triggering
Cellular & Molecular Immunology (2020)
-
Characteristics of T cell receptor repertoires of patients with acute myocardial infarction through high-throughput sequencing
Journal of Translational Medicine (2019)
-
Global survey of the immunomodulatory potential of common drugs
Nature Chemical Biology (2017)
-
Differential utilization of binding loop flexibility in T cell receptor ligand selection and cross-reactivity
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.