Abstract
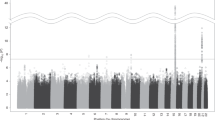
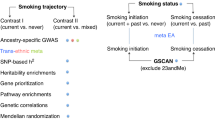
Cigarette smoking is a leading cause of preventable mortality worldwide. Nicotine dependence, which reduces the likelihood of quitting smoking, is a heritable trait with firmly established associations with sequence variants in nicotine acetylcholine receptor genes and at other loci. To search for additional loci, we conducted a genome-wide association study (GWAS) meta-analysis of nicotine dependence, totaling 38,602 smokers (28,677 Europeans/European Americans and 9925 African Americans) across 15 studies. In this largest-ever GWAS meta-analysis for nicotine dependence and the largest-ever cross-ancestry GWAS meta-analysis for any smoking phenotype, we reconfirmed the well-known CHRNA5-CHRNA3-CHRNB4 genes and further yielded a novel association in the DNA methyltransferase gene DNMT3B. The intronic DNMT3B rs910083-C allele (frequency=44–77%) was associated with increased risk of nicotine dependence at P=3.7 × 10−8 (odds ratio (OR)=1.06 and 95% confidence interval (CI)=1.04–1.07 for severe vs mild dependence). The association was independently confirmed in the UK Biobank (N=48,931) using heavy vs never smoking as a proxy phenotype (P=3.6 × 10−4, OR=1.05, and 95% CI=1.02–1.08). Rs910083-C is also associated with increased risk of squamous cell lung carcinoma in the International Lung Cancer Consortium (N=60,586, meta-analysis P=0.0095, OR=1.05, and 95% CI=1.01–1.09). Moreover, rs910083-C was implicated as a cis-methylation quantitative trait locus (QTL) variant associated with higher DNMT3B methylation in fetal brain (N=166, P=2.3 × 10−26) and a cis-expression QTL variant associated with higher DNMT3B expression in adult cerebellum from the Genotype-Tissue Expression project (N=103, P=3.0 × 10−6) and the independent Brain eQTL Almanac (N=134, P=0.028). This novel DNMT3B cis-acting QTL variant highlights the importance of genetically influenced regulation in brain on the risks of nicotine dependence, heavy smoking and consequent lung cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
World Health Organization. WHO report on the global tobacco epidemic, 2011: Warning about the dangers of tobacco. World Health Organization: Geneva, Switzerland, 2011.
U.S. Department of Health and Human Services. The health consequences of smoking—50 years of progress: a Report of the Surgeon General. U.S. Department of Health and Human Services: Atlanta, GA, 2014.
Centers for Disease Control and Prevention (CDC). Vital signs: current cigarette smoking among adults aged >18 years—United States, 2005–2010. MMWR Morb Mortal Wkly Rep 2011; 60: 1207–1212.
Centers for Disease Control and Prevention (CDC). Quitting smoking among adults—United States, 2001–2010. MMWR Morb Mortal Wkly Rep 2011; 60: 1513–1519.
Shiffman S, Tindle H, Li X, Scholl S, Dunbar M, Mitchell-Miland C. Characteristics and smoking patterns of intermittent smokers. Exp Clin Psychopharmacol 2012; 20: 264–277.
Breslau N, Johnson EO. Predicting smoking cessation and major depression in nicotine-dependent smokers. Am J Public Health 2000; 90: 1122–1127.
Baker TB, Piper ME, McCarthy DE, Bolt DM, Smith SS, Kim SY et al. Time to first cigarette in the morning as an index of ability to quit smoking: implications for nicotine dependence. Nicotine Tob Res 2007; 9(Suppl 4): S555–S570.
Sullivan PF, Kendler KS. The genetic epidemiology of smoking. Nicotine Tob Res 1999; 1: S51–S57.
Vink JM, Willemsen G, Boomsma DI. Heritability of smoking initiation and nicotine dependence. Behav Genet 2005; 35: 397–406.
Baker TB, Piper ME, Schlam TR, Cook JW, Smith SS, Loh WY et al. Are tobacco dependence and withdrawal related amongst heavy smokers? Relevance to conceptualizations of dependence. J Abnorm Psychol 2012; 121: 909–921.
Zelman DC, Brandon TH, Jorenby DE, Baker TB. Measures of affect and nicotine dependence predict differential response to smoking cessation treatments. J Consult Clin Psychol 1992; 60: 943–952.
Gu F, Wacholder S, Kovalchik S, Panagiotou OA, Reyes-Guzman C, Freedman ND et al. Time to smoke first morning cigarette and lung cancer in a case-control study. J Natl Cancer Inst 2014; 106: dju118.
Guertin KA, Gu F, Wacholder S, Freedman ND, Panagiotou OA, Reyes-Guzman C et al. Time to first morning cigarette and risk of chronic obstructive pulmonary disease: smokers in the PLCO Cancer Screening Trial. PLoS One 2015; 10: e0125973.
Bierut LJ, Madden PA, Breslau N, Johnson EO, Hatsukami D, Pomerleau OF et al. Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Thorgeirsson TE, Geller F, Sulem P, Rafnar T, Wiste A, Magnusson KP et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 2008; 452: 638–642.
Liu JZ, Tozzi F, Waterworth DM, Pillai SG, Muglia P, Middleton L et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat Genet 2010; 42: 436–440.
Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat Genet 2010; 42: 441–447.
Thorgeirsson TE, Gudbjartsson DF, Surakka I, Vink JM, Amin N, Geller F et al. Sequence variants at CHRNB3-CHRNA6 and CYP2A6 affect smoking behavior. Nat Genet 2010; 42: 448–453.
Rice JP, Hartz SM, Agrawal A, Almasy L, Bennett S, Breslau N et al. CHRNB3 is more strongly associated with Fagerstrom Test for Cigarette Dependence-based nicotine dependence than cigarettes per day: phenotype definition changes genome-wide association studies results. Addiction 2012; 107: 2019–2028.
David SP, Hamidovic A, Chen GK, Bergen AW, Wessel J, Kasberger JL et al. Genome-wide meta-analyses of smoking behaviors in African Americans. Transl Psychiatry 2012; 2: e119.
Loukola A, Wedenoja J, Keskitalo-Vuokko K, Broms U, Korhonen T, Ripatti S et al. Genome-wide association study on detailed profiles of smoking behavior and nicotine dependence in a twin sample. Mol Psychiatry 2014; 19: 615–624.
Gelernter J, Kranzler HR, Sherva R, Almasy L, Herman AI, Koesterer R et al. Genome-wide association study of nicotine dependence in American populations: identification of novel risk loci in both African-Americans and European-Americans. Biol Psychiatry 2015; 77: 493–503.
Hancock DB, Reginsson GW, Gaddis NC, Chen X, Saccone NL, Lutz SM et al. Genome-wide meta-analysis reveals common splice site acceptor variant in CHRNA4 associated with nicotine dependence. Transl Psychiatry 2015; 5: e651.
Wain LV, Shrine N, Miller S, Jackson VE, Ntalla I, Artigas MS et al. Novel insights into the genetics of smoking behaviour, lung function, and chronic obstructive pulmonary disease (UK BiLEVE): a genetic association study in UK Biobank. Lancet Respir Med 2015; 3: 769–781.
Baker TB, Breslau N, Covey L, Shiffman S. DSM criteria for tobacco use disorder and tobacco withdrawal: a critique and proposed revisions for DSM-5. Addiction 2012; 107: 263–275.
Heatherton TF, Kozlowski LT, Frecker RC, Fagerstrom KO. The fagerstrom test for nicotine dependence: a revision of the fagerstrom tolerance questionnaire. Br J Addict 1991; 86: 1119–1127.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010; 26: 2190–2191.
Pe'er I, Yelensky R, Altshuler D, Daly MJ. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet Epidemiol 2008; 32: 381–385.
Fadista J, Manning AK, Florez JC, Groop L. The (in)famous GWAS P-value threshold revisited and updated for low-frequency variants. Eur J Hum Genet 2016; 24: 1202–1205.
Kanai M, Tanaka T, Okada Y. Empirical estimation of genome-wide significance thresholds based on the 1000 Genomes Project data set. J Hum Genet 2016; 61: 861–866.
de Vries PS, Sabater-Lleal M, Chasman DI, Trompet S, Ahluwalia TS, Teumer A et al. Comparison of HapMap and 1000 genomes reference panels in a large-scale genome-wide association study. PLoS One 2017; 12: e0167742.
Sveinbjornsson G, Albrechtsen A, Zink F, Gudjonsson SA, Oddson A, Masson G et al. Weighting sequence variants based on their annotation increases power of whole-genome association studies. Nat Genet 2016; 48: 314–317.
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 2010; 26: 2336–2337.
Boyles AL, Harris SF, Rooney AA, Thayer KA. Forest Plot Viewer: a new graphing tool. Epidemiology 2011; 22: 746–747.
Huedo-Medina TB, Sanchez-Meca J, Marin-Martinez F, Botella J. Assessing heterogeneity in meta-analysis: Q statistic or I2 index? Psychol Methods 2006; 11: 193–206.
Hannon E, Spiers H, Viana J, Pidsley R, Burrage J, Murphy TM et al. Methylation QTLs in the developing brain and their enrichment in schizophrenia risk loci. Nat Neurosci 2016; 19: 48–54.
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 2015; 348: 648–660.
Ramasamy A, Trabzuni D, Guelfi S, Varghese V, Smith C, Walker R et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat Neurosci 2014; 17: 1418–1428.
McKay JD, Hung RJ, Han Y, Zong X, Carreras-Torres R, Christiani DC et al. Large scale genetic analysis identifies novel loci and histological variability in susceptibility to lung cancer. Nat Genet 2017; 49: 1126–1132.
Fagerstrom KO. Measuring degree of physical dependence to tobacco smoking with reference to individualization of treatment. Addict Behav 1978; 3: 235–241.
Gibbs JR, van der Brug MP, Hernandez DG, Traynor BJ, Nalls MA, Lai SL et al. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet 2010; 6: e1000952.
Hancock DB, Wang JC, Gaddis NC, Levy JL, Saccone NL, Stitzel JA et al. A multiancestry study identifies novel genetic associations with CHRNA5 methylation in human brain and risk of nicotine dependence. Hum Mol Genet 2015; 24: 5940–5944.
Wang JC, Spiegel N, Bertelsen S, Le N, McKenna N, Budde JP et al. Cis-regulatory variants affect CHRNA5 mRNA expression in populations of African and European ancestry. PLoS One 2013; 8: e80204.
Wang JC, Cruchaga C, Saccone NL, Bertelsen S, Liu P, Budde JP et al. Risk for nicotine dependence and lung cancer is conferred by mRNA expression levels and amino acid change in CHRNA5. Hum Mol Genet 2009; 18: 3125–3135.
Goldstein RZ, Volkow ND. Dysfunction of the prefrontal cortex in addiction: neuroimaging findings and clinical implications. Nat Rev Neurosci 2011; 12: 652–669.
Miquel M, Vazquez-Sanroman D, Carbo-Gas M, Gil-Miravet I, Sanchis-Segura C, Carulli D et al. Have we been ignoring the elephant in the room? Seven arguments for considering the cerebellum as part of addiction circuitry. Neurosci Biobehav Rev 2016; 60: 1–11.
Moulton EA, Elman I, Becerra LR, Goldstein RZ, Borsook D. The cerebellum and addiction: insights gained from neuroimaging research. Addict Biol 2014; 19: 317–331.
Strick PL, Dum RP, Fiez JA. Cerebellum and nonmotor function. Annu Rev Neurosci 2009; 32: 413–434.
Weissman J, Naidu S, Bjornsson HT. Abnormalities of the DNA methylation mark and its machinery: an emerging cause of neurologic dysfunction. Semin Neurol 2014; 34: 249–257.
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999; 99: 247–257.
Kaneda M, Okano M, Hata K, Sado T, Tsujimoto N, Li E et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 2004; 429: 900–903.
Jin B, Tao Q, Peng J, Soo HM, Wu W, Ying J et al. DNA methyltransferase 3B (DNMT3B) mutations in ICF syndrome lead to altered epigenetic modifications and aberrant expression of genes regulating development, neurogenesis and immune function. Hum Mol Genet 2008; 17: 690–709.
Teneng I, Tellez CS, Picchi MA, Klinge DM, Yingling CM, Snider AM et al. Global identification of genes targeted by DNMT3b for epigenetic silencing in lung cancer. Oncogene 2015; 34: 621–630.
Oka M, Meacham AM, Hamazaki T, Rodic N, Chang LJ, Terada N. De novo DNA methyltransferases Dnmt3a and Dnmt3b primarily mediate the cytotoxic effect of 5-aza-2'-deoxycytidine. Oncogene 2005; 24: 3091–3099.
Gnyszka A, Jastrzebski Z, Flis S. DNA methyltransferase inhibitors and their emerging role in epigenetic therapy of cancer. Anticancer Res 2013; 33: 2989–2996.
Ward LD, Kellis M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 2012; 40: D930–D934.
Domino EF, Minoshima S, Guthrie S, Ohl L, Ni L, Koeppe RA et al. Nicotine effects on regional cerebral blood flow in awake, resting tobacco smokers. Synapse 2000; 38: 313–321.
Zubieta JK, Heitzeg MM, Xu Y, Koeppe RA, Ni L, Guthrie S et al. Regional cerebral blood flow responses to smoking in tobacco smokers after overnight abstinence. Am J Psychiatry 2005; 162: 567–577.
Brody AL, Mandelkern MA, Jarvik ME, Lee GS, Smith EC, Huang JC et al. Differences between smokers and nonsmokers in regional gray matter volumes and densities. Biol Psychiatry 2004; 55: 77–84.
Gallinat J, Meisenzahl E, Jacobsen LK, Kalus P, Bierbrauer J, Kienast T et al. Smoking and structural brain deficits: a volumetric MR investigation. Eur J Neurosci 2006; 24: 1744–1750.
Kuhn S, Romanowski A, Schilling C, Mobascher A, Warbrick T, Winterer G et al. Brain grey matter deficits in smokers: focus on the cerebellum. Brain Struct Funct 2012; 217: 517–522.
Sutherland MT, Riedel MC, Flannery JS, Yanes JA, Fox PT, Stein EA et al. Chronic cigarette smoking is linked with structural alterations in brain regions showing acute nicotinic drug-induced functional modulations. Behav Brain Funct 2016; 12: 16.
Wood AR, Perry JR, Tanaka T, Hernandez DG, Zheng HF, Melzer D et al. Imputation of variants from the 1000 Genomes Project modestly improves known associations and can identify low-frequency variant-phenotype associations undetected by HapMap based imputation. PLoS One 2013; 8: e64343.
Huang J, Ellinghaus D, Franke A, Howie B, Li Y. 1000 Genomes-based imputation identifies novel and refined associations for the Wellcome Trust Case Control Consortium phase 1 Data. Eur J Hum Genet 2012; 20: 801–805.
Jostins L, Ripke S, Weersma RK, Duerr RH, McGovern DP, Hui KY et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 2012; 491: 119–124.
Ananthakrishnan AN. Environmental risk factors for inflammatory bowel disease. Gastroenterol Hepatol (NY) 2013; 9: 367–374.
Thorgeirsson TE, Steinberg S, Reginsson GW, Bjornsdottir G, Rafnar T, Jonsdottir I et al. A rare missense mutation in CHRNA4 associates with smoking behavior and its consequences. Mol Psychiatry 2016; 21: 594–600.
Xie P, Kranzler HR, Krauthammer M, Cosgrove KP, Oslin D, Anton RF et al. Rare nonsynonymous variants in alpha-4 nicotinic acetylcholine receptor gene protect against nicotine dependence. Biol Psychiatry 2011; 70: 528–536.
Siedlinski M, Cho MH, Bakke P, Gulsvik A, Lomas DA, Anderson W et al. Genome-wide association study of smoking behaviours in patients with COPD. Thorax 2011; 66: 894–902.
Hirvonen K, Korhonen T, Salomaa V, Männistö S, Kaprio J. Effect of the DBH polymorphism rs3025343 on smoking cessation in a large population-based sample. Nic Tob Research 2017; 19: 1112–1115.
Yang J, Wang S, Yang Z, Hodgkinson CA, Iarikova P, Ma JZ et al. The contribution of rare and common variants in 30 genes to risk nicotine dependence. Mol Psychiatry 2015; 20: 1467–1478.
Amos CI, Wu X, Broderick P, Gorlov IP, Gu J, Eisen T et al. Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nat Genet 2008; 40: 616–622.
Barrie ES, Weinshenker D, Verma A, Pendergrass SA, Lange LA, Ritchie MD et al. Regulatory polymorphisms in human DBH affect peripheral gene expression and sympathetic activity. Circ Res 2014; 115: 1017–1025.
Lee PN, Forey BA, Coombs KJ. Systematic review with meta-analysis of the epidemiological evidence in the 1900s relating smoking to lung cancer. BMC Cancer 2012; 12: 385.
Lee PN, Forey BA. Indirectly estimated absolute lung cancer mortality rates by smoking status and histological type based on a systematic review. BMC Cancer 2013; 13: 189.
Acknowledgments
We thank the many study participants. We also thank Michael E. Hall for reviewing the manuscript. This work was supported by the National Institute on Drug Abuse grant numbers R01 DA035825, R01 DA036583 and R01 DA042090. Acknowledgments for the nicotine dependence studies are included in the Supplementary Information. Funding for lung cancer studies was provided by the National Cancer Institute grant number U19 CA148127.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Dr Bierut and the spouse of Dr Saccone are listed as inventors on U.S. Patent 8,080,371, ‘Markers for Addiction’ covering the use of certain SNPs in determining the diagnosis, prognosis and treatment of addiction. Authors listed with the affiliation deCODE Genetics/AMGEN are employees of deCODE genetics/AMGEN. Although unrelated to this research, Dr Kranzler has been a consultant or advisory board member for Lundbeck and Indivior and is a member of the American Society of Clinical Psychopharmacology’s Alcohol Clinical Trials Initiative, which was supported in the last 3 years by AbbVie, Alkermes, Ethypharm, Indivior, Lilly, Lundbeck, Otsuka, Pfizer, Arbor and Amygdala Neurosciences. Dr Kaprio has consulted for Pfizer in 2012–2014 on nicotine dependence. The remaining authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hancock, D.B., Guo, Y., Reginsson, G.W. et al. Genome-wide association study across European and African American ancestries identifies a SNP in DNMT3B contributing to nicotine dependence. Mol Psychiatry 23, 1911–1919 (2018). https://doi.org/10.1038/mp.2017.193
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2017.193
This article is cited by
-
Deep sequencing of candidate genes identified 14 variants associated with smoking abstinence in an ethnically diverse sample
Scientific Reports (2024)
-
The Comprehensive Effect of Socioeconomic Deprivation on Smoking Behavior: an Observational and Genome-Wide by Environment Interaction Analyses in UK Biobank
International Journal of Mental Health and Addiction (2024)
-
Multivariate genome-wide association meta-analysis of over 1 million subjects identifies loci underlying multiple substance use disorders
Nature Mental Health (2023)
-
Genetic insights into smoking behaviours in 10,558 men of African ancestry from continental Africa and the UK
Scientific Reports (2022)
-
The addiction risk factor: A unitary genetic vulnerability characterizes substance use disorders and their associations with common correlates
Neuropsychopharmacology (2022)