Abstract
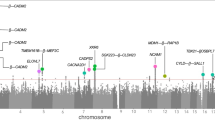
Personality can be thought of as a set of characteristics that influence people's thoughts, feelings and behavior across a variety of settings. Variation in personality is predictive of many outcomes in life, including mental health. Here we report on a meta-analysis of genome-wide association (GWA) data for personality in 10 discovery samples (17 375 adults) and five in silico replication samples (3294 adults). All participants were of European ancestry. Personality scores for Neuroticism, Extraversion, Openness to Experience, Agreeableness and Conscientiousness were based on the NEO Five-Factor Inventory. Genotype data of ∼2.4M single-nucleotide polymorphisms (SNPs; directly typed and imputed using HapMap data) were available. In the discovery samples, classical association analyses were performed under an additive model followed by meta-analysis using the weighted inverse variance method. Results showed genome-wide significance for Openness to Experience near the RASA1 gene on 5q14.3 (rs1477268 and rs2032794, P=2.8 × 10−8 and 3.1 × 10−8) and for Conscientiousness in the brain-expressed KATNAL2 gene on 18q21.1 (rs2576037, P=4.9 × 10−8). We further conducted a gene-based test that confirmed the association of KATNAL2 to Conscientiousness. In silico replication did not, however, show significant associations of the top SNPs with Openness and Conscientiousness, although the direction of effect of the KATNAL2 SNP on Conscientiousness was consistent in all replication samples. Larger scale GWA studies and alternative approaches are required for confirmation of KATNAL2 as a novel gene affecting Conscientiousness.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Costa PT, McCrae RR . Professional Manual: Revised NEO Personality Inventory (NEO-PI-R) and NEO Five-Factor-Inventory (NEO-FFI). Psychological Assessment Resources: Odessa, FL, 1992.
Samuel DB, Widiger TA . A meta-analytic review of the relationships between the five-factor model and DSM-IV-TR personality disorders: a facet level analysis. Clin Psychol Rev 2008; 28: 1326–1342.
Hettema JM, Neale MC, Myers JM, Prescott CA, Kendler KS . A population-based twin study of the relationship between neuroticism and internalizing disorders. Am J Psychiatry 2006; 163: 857–864.
Terracciano A, Lockenhoff CE, Zonderman AB, Ferrucci L, Costa PT . Personality predictors of longevity: activity, emotional stability, and conscientiousness. Psychosom Med 2008; 70: 621–627.
Dick DM, Aliev F, Wang JC, Grucza RA, Schuckit M, Kuperman S et al. Using dimensional models of externalizing psychopathology to aid in gene identification. Arch Gen Psychiatry 2008; 65: 310–318.
Kendler KS, Gatz M, Gardner CO, Pedersen NL . Personality and major depression—a Swedish longitudinal, population-based twin study. Arch Gen Psychiatry 2006; 63: 1113–1120.
Lahey BB . Public health significance of neuroticism. Am Psychol 2009; 64: 241–256.
Bienvenu OJ, Samuels JF, Costa PT, Reti IM, Eaton WW, Nestadt G . Anxiety and depressive disorders and the five-factor model of personality: a higher- and lower-order personality trait investigation in a community sample. Depress Anxiety 2004; 20: 92–97.
Fanous A, Gardner CO, Prescott CA, Cancro R, Kendler KS . Neuroticism, major depression and gender: a Population-Based Twin Study. Psychol Med 2002; 32: 719–728.
Kendler KS, Myers J . The genetic and environmental relationship between major depression and the five-factor model of personality. Psychol Med 2009; 40: 1–6.
Weiss A, Sutin AR, Duberstein PR, Friedman B, Bagby RM, Costa PT . The personality domains and styles of the five-factor model are related to incident depression in Medicare recipients aged 65 to 100. Am J Geriatr Psychiatry 2009; 17: 591–601.
Terracciano A, Costa PT . Smoking and the Five-Factor Model of personality. Addiction 2004; 99: 472–481.
Terracciano A, Lockenhoff CE, Crum RM, Bienvenu OJ, Costa PT . Five-factor model personality profiles of drug users. BMC Psychiatry 2008; 8: 22.
Saulsman LM, Page AC . The five-factor model and personality disorder empirical literature: a meta-analytic review. Clin Psychol Rev 2004; 23: 1055–1085.
Thoresen CJ, Bradley JC, Bliese PD, Thoresen JD . The big five personality traits and individual job performance growth trajectories in maintenance and transitional job stages. J Appl Psychol 2004; 89: 835–853.
De Moor MHM, Beem AL, Stubbe JH, Boomsma DI, de Geus EJC . Regular exercise, anxiety, depression and personality: a population-based study. Prev Med 2006; 42: 273–279.
Rhodes RE, Smith NEI . Personality correlates of physical activity: a review and meta-analysis. Br J Sports Med 2006; 40: 958–965.
Bouchard TJ, Loehlin JC . Genes, evolution, and personality. Behav Genet 2001; 31: 243–273.
Vernon PA, Martin RA, Schermer JA, Mackie A . A behavioral genetic investigation of humor styles and their correlations with the Big-5 personality dimensions. Pers Ind Diff 2008; 44: 1116–1125.
Distel MA, Trull TJ, Willemsen G, Vink JM, Derom CA, Lynskey MT et al. The Five Factor Model of personality and borderline personality disorder: a genetic analysis of comorbidity. Biol Psychiatry 2009; 66: 1131–1138.
Pilia G, Chen WM, Scuteri A, Orru M, Albai G, Dei M et al. Heritability of cardiovascular and personality traits in 6,148 sardinians. Plos Genet 2006; 2: 1207–1223.
Jang KL, Livesley WJ, Angleitner A, Riemann R, Vernon PA . Genetic and environmental influences on the covariance of facets defining the domains of the five-factor model of personality. Pers Ind Diff 2002; 33: 83–101.
Shifman S, Bhomra A, Smiley S, Wray NR, James MR, Martin NG et al. A whole genome association study of neuroticism using DNA pooling. Mol Psychiatry 2008; 13: 302–312.
van den Oord EJ, Kuo PH, Hartmann AM, Webb BT, Moller HJ, Hettema JM et al. Genomewide association analysis followed by a replication study implicates a novel candidate gene for neuroticism. Arch Gen Psychiatry 2008; 65: 1062–1071.
Kuo PH, Neale MC, Riley BP, Patterson DG, Walsh D, Prescott CA et al. A genome-wide linkage analysis for the personality trait neuroticism in the Irish affected sib-pair study of alcohol dependence. Am J Med Genet B 2007; 144B: 463–468.
Neale BM, Sullivan PF, Kendler KS . A genome scan of neuroticism in nicotine dependent smokers. Am J Med Genet B 2005; 132B: 65–69.
Gillespie NA, Zhu G, Evans DM, Medland SE, Wright MJ, Martin NG . A genome-wide scan for Eysenckian personality dimensions in adolescent twin sibships: psychoticism, extraversion, neuroticism, and lie. J Pers 2008; 76: 1415–1446.
Fullerton J, Cubin M, Tiwari H, Wang C, Bomhra A, Davidson S et al. Linkage analysis of extremely discordant and concordant sibling pairs identifies quantitative-trait loci that influence variation in the human personality trait neuroticism. Am J Hum Genet 2003; 72: 879–890.
Nash MW, Huezo-Diaz P, Sterne A, Purcell S, Hoda F, Cherny SS et al. Genome-wide linkage analysis of a composite index of neuroticism and mood-related scales in extreme selected sibships. Hum Mol Genet 2004; 13: 2173–2182.
Hettema JM, Van den Oord EJCG, An SS, Kendler KS, Chen XN . Follow-up association study of novel neuroticism gene MAMDC1. Psychiatric Genet 2009; 19: 213–214.
Terracciano A, Sanna S, Uda M, Deiana B, Usala G, Busonero F et al. Genome-wide association scan for five major dimensions of personality. Mol Psychiatry 2010; 15: 647–656.
Boomsma DI, de Geus EJC, Vink JM, Stubbe JH, Distel MA, Hottenga JJ et al. Netherlands Twin Register: from twins to twin families. Twin Res Hum Genet 2006; 9: 849–857.
Penninx BWJH, Beekman ATF, Smit JH, Zitman FG, Nolen WA, Spinhoven P et al. The Netherlands Study of Depression and Anxiety (NESDA): rationale, objectives and methods. Int J Methods Psychiatric Res 2008; 17: 121–140.
Manolio TA, Rodriguez LL, Brooks L, Abecasis G, Ballinger D, Daly M et al. New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nat Genet 2007; 39: 1045–1051.
Boomsma DI, Willemsen G, Sullivan PF, Heutink P, Meijer P, Sondervan D et al. Genome-wide association of major depression: description of samples for the GAIN Major Depressive Disorder Study: NTR and NESDA biobank projects. Eur J Hum Genet 2008; 16: 335–342.
Sullivan PF, de Geus EJC, Willemsen G, James MR, Smit JH, Zandbelt T et al. Genomewide association for major depressive disorder: a possible role for the presynaptic protein piccolo. Mol Psychiatry 2009; 14: 359–375.
Distel MA, Ligthart L, Willemsen G, Nyholt DR, Trull TJ, Boomsma DI . Personality, health and lifestyle in a questionnaire family study: a comparison between highly cooperative and less cooperative families. Twin Res Hum Genet 2007; 10: 348–353.
Pardo LM, MacKay I, Oostra B, van Duijn CM, Aulchenko YS . The effect of genetic drift in a young genetically isolated population. Ann Hum Genet 2005; 69: 288–295.
Bierut LJ, Agrawal A, Bucholz KK, Doheny KF, Laurie C, Pugh E et al. A genome-wide association study of alcohol dependence. Proc Natl Acad Sci USA 2010; 107: 5082–5087.
Barker DJP, Osmond C, Forsen TJ, Kajantie E, Eriksson JG . Trajectories of growth among children who have coronary events as adults. N Engl J Med 2005; 353: 1802–1809.
Eriksson JG, Osmond C, Kajantie E, Forsen TJ, Barker DJP . Patterns of growth among children who later develop type 2 diabetes or its risk factors. Diabetologia 2006; 49: 2853–2858.
Raikkonen K, Pesonen AK, Heinonen K, Lahti J, Kajantie E, Forsen T et al. Infant growth and hostility in adult life. Psychosom Med 2008; 70: 306–313.
Pergadia ML, Agrawal A, Loukola A, Montgomery GW, Broms U, Saccone SF . Genetic linkage findings for DSM-IV nicotine withdrawal in two populations. Am J Med Genet B 2009; 150B: 950–959.
Saccone SF, Pergadia ML, Loukola A, Broms U, Montgomery GW, Wang JC et al. Genetic linkage to chromosome 22q12 for a heavy-smoking quantitative trait in two independent samples. Am J Hum Genet 2007; 80: 856–866.
Aitken JF, Green A, Eldridge A, Green L, Pfitzner J, Battistutta D et al. Comparability of nevus counts between and within examiners, and comparison with computer image-analysis. Br J Cancer 1994; 69: 487–491.
Wright MJ, Martin NG . Brisbane adolescent twin study: outline of study methods and research projects. Aus J Psychol 2004; 56: 65–78.
Distel MA, Trull TJ, Derom CA, Thiery EW, Grimmer MA, Martin NG et al. Heritability of borderline personality disorder features is similar across three countries. Psychol Med 2008; 38: 1219–1229.
Deary IJ, Gow AJ, Taylor MD, Corley J, Brett C, Wilson V et al. The Lothian Birth Cohort 1936: a study to examine influences on cognitive ageing from age 11 to age 70 and beyond. BMC Geriatr 2007; 7: 28.
Deary IJ, Whiteman MC, Starr JM, Whalley LJ, Fox HC . The impact of childhood intelligence on later life: following up the Scottish Mental Surveys of 1932 and 1947. J Pers Soc Psychol 2004; 86: 130–147.
Terracciano A, McCrae RR, Brant LJ, Costa PT . Hierarchical linear modeling analyses of the NEO-PI-R scales in the Baltimore longitudinal study of aging. Psychol Aging 2005; 20: 493–506.
Terracciano A, Costa PT, McCrae RR . Personality plasticity after age 30. Pers Soc Psychol Bull 2006; 32: 999–1009.
Metspalu A . The Estonian Genome Project. Drug Develop Res 2004; 62: 97–101.
Nelis M, Esko T, Magi R, Zimprich F, Zimprich A, Toncheva D et al. Genetic structure of Europeans: a view from the North-East. PLoS ONE 2009; 4: e5472.
McCrae RR, Costa PT, Martin TA . The NEO-PI-3: a more readable revised NEO Personality Inventory. J Pers Assess 2005; 84: 261–270.
Willemsen G, de Geus EJC, Bartels M, van Beijsterveldt CEM, Brooks AI, Estourgie-van Burk GF et al. The Netherlands Twin Register Biobank: a resource for genetic epidemiological studies. Twin Res Hum Genet 2010; 13: 231–245.
Rice JP, Reich T, Bucholz KK, Neuman RJ, Fishman R, Rochberg N et al. Comparison of direct interview and family history diagnoses of alcohol dependence. Alcohol Clin Exp Res 1995; 19: 1018–1023.
First MB, Spitzer RL, Gibbon M, Williams JB . Structured Clinical Interview for DSM-IV Axis I Disorders—Patient Edition. Biometrics Research Department, New York State Psychiatric Institute: NY, 1995.
Folstein MF, Folstein SE, McHugh PR . Mini-Mental-Status-Test, German Edition. Beltz: Weinheim, 1993.
Allik J, Laidra K, Realo A, Pullmann H . Personality development from 12 to 18 years of age: changes in mean levels and structure of traits. Eur J Pers 2004; 18: 445–462.
Kallasmaa T, Allik J, Realo A, McCrae RR . The Estonian version of the NEO-PI-R: an examination of universal and culture-specific aspects of the five-factor model. Eur J Pers 2000; 14: 265–278.
Marchini J, Howie B, Myers S, McVean G, Donnelly P . A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 2007; 39: 906–913.
Chen WM, Abecasis GR . Family-based association tests for genomewide association scans. Am J Hum Genet 2007; 81: 913–926.
Abecasis G . METAL. 2009 URL: http://www.sph.umich.edu/csg/abecasis/Metal/.
Dudbridge F, Gusnanto A . Estimation of significance thresholds for genomewide association scans. Genet Epidemiol 2008; 32: 227–234.
Liu JZ, McRae AF, Nyholt DR, Medland SE, Wray NR, Brown KM et al. A versatile gene-based test for genome-wide association studies. Am J Hum Genet 2010; 87: 139–145.
Liu X, Yu XP, Zack DJ, Zhu H, Qian J . TiGER: a database for tissue-specific gene expression and regulation. BMC Bioinform 2008; 9: 271.
Karabay A, Yu WQ, Solowska JM, Baird DH, Baas PW . Axonal growth is sensitive to the levels of katanin, a protein that severs microtubules. J Neurosci 2004; 24: 5778–5788.
Lee HH, Jan LY, Jan YN . Drosophila IKK-related kinase Ik2 and Katanin p60-like 1 regulate dendrite pruning of sensory neuron during metamorphosis. Proc Natl Acad Sci USA 2009; 106: 6363–6368.
Toyo-Oka K, Sasaki S, Yano Y, Mori D, Kobayashi T, Toyoshima YY et al. Recruitment of katanin p60 by phosphorylated NDEL1, an LIS1 interacting protein, is essential for mitotic cell division and neuronal migration. Hum Mol Genet 2005; 14: 3113–3128.
Lango Allen H, Estrada K, Lettre G, Berndt SI, Weedon MN, Rivadeneira F et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010; 467: 832–838.
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet 2010; 42: 565–569.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ et al. Finding the missing heritability of complex diseases. Nature 2009; 461: 747–753.
Acknowledgements
We would like to thank the individuals who participated in the studies. Meta-analysis and statistical analyses for the NAG/IPRG, QIMR and NTR/NESDA studies were carried out on the Genetic Cluster Computer (http://www.geneticcluster.org), which is financially supported by the Netherlands Organization for Scientific Research (NWO 480-05-003). SardiNIA: We acknowledge support from the Intramural Research Program of the NIH, National Institute on Aging. Funding was provided by the National Institute on Aging, NIH Contract No. NO1-AG-1-2109 to the SardiNIA (‘ProgeNIA’) team. NTR/NESDA: We acknowledge financial support from the Netherlands Organization for Scientific Research (NWO): Grants 575-25-006, 480-04-004, 904-61-090; 904-61-193, 400-05-717 and Spinozapremie SPI 56-464-14192. MHMdeM is financially supported by ZonMW (Addiction) Grant No. 311-60008. We further acknowledge financial support from the Center for Medical Systems Biology (NWO Genomics), the Centre for Neurogenomics and Cognitive Research (CNCR-VU); EU/QLRT-2001-01254; NIMH R01 MH059160; Geestkracht program of ZonMW (10-000-1002); matching funds from universities and mental healthcare institutes involved in NESDA. Genotyping was funded by the Genetic Association Information Network (GAIN) of the Foundation for the US National Institutes of Health, and analysis was supported by grants from Genetic Association Information Network and the NIMH (MH081802). Genotype data were obtained from dbGaP (http://www.ncbi.nlm.nih.gov/dbgap, accession number phs000020.v1.p1). ERF: The genotyping for the ERF study was supported by EUROSPAN (European Special Populations Research Network) and the European Commission FP6 STRP Grant (018947; LSHG-CT-2006-01947). The ERF study was further supported by grants from the Netherlands Organization for Scientific Research, Erasmus MC, the Centre for Medical Systems Biology (CMSB) and the Netherlands Brain Foundation (HersenStichting Nederland). We are grateful to all patients and their relatives, general practitioners and neurologists for their contributions and to P Veraart for her help in genealogy, Jeannette Vergeer for the supervision of the laboratory work and P Snijders for his help in data collection. SAGE: Funding support for the Study of Addiction: Genetics and Environment (SAGE) was provided through the NIH Genes, Environment and Health Initiative (GEI) (U01 HG004422). SAGE is one of the genome-wide association studies funded as part of the Gene Environment Association Studies under GEI. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by the Gene Environment Association Studies initiative Coordinating Center (U01 HG004446). Assistance with data cleaning was provided by the National Center for Biotechnology Information. Support for collection of data sets and samples was provided by the Collaborative Study on the Genetics of Alcoholism (U10 AA008401), the Collaborative Genetic Study of Nicotine Dependence (P01 CA089392) and the Family Study of Cocaine Dependence (R01 DA013423). Funding support for genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research, was provided by the NIH GEI (U01HG004438), the National Institute on Alcohol Abuse and Alcoholism, the National Institute on Drug Abuse and the NIH contract ‘High-throughput genotyping for studying the genetic contributions to human disease’ (HHSN268200782096C). The Collaborative Study on the Genetics of Alcoholism, principal investigators: B Porjesz, V Hesselbrock, H Edenberg, LJ Bierut, includes 10 different centers: University of Connecticut (V Hesselbrock); Indiana University (HJ Edenberg, J Nurnberger Jr, T Foroud); University of Iowa (S Kuperman, J Kramer); SUNY Downstate (B Porjesz); Washington University in St Louis (LJ Bierut, A Goate, J Rice, K Bucholz); University of California at San Diego (M Schuckit); Rutgers University (J Tischfield); Southwest Foundation (L Almasy), Howard University (R Taylor) and Virginia Commonwealth University (D Dick). A Parsian and M Reilly are the NIAAA Staff Collaborators. We continue to be inspired by our memories of Henri Begleiter and Theodore Reich, founding PI and Co-PI of COGA, and also owe a debt of gratitude to other past organizers of COGA, including Ting-Kai Li, P Michael Conneally, Raymond Crowe and Wendy Reich, for their critical contributions. This national collaborative study is supported by NIH Grant U10AA008401 from the National Institute on Alcohol Abuse and Alcoholism (NIAAA) and the National Institute on Drug Abuse (NIDA). The Collaborative Genetic Study of Nicotine Dependence project is a collaborative research group and part of the NIDA Genetics Consortium. Subject collection was supported by NIH Grant CA89392 (PI—LJ Bierut) from the National Cancer Institute. Genotyping work at Perlegen Sciences was performed under NIDA Contract HHSN271200477471C. Phenotypic and genotypic data are stored in the NIDA Center for Genetic Studies (NCGS) at http://zork.wustl.edu/ under NIDA Contract HHSN271200477451C (PIs—J Tischfield and J Rice). Genotyping services were also provided by the Center for Inherited Disease Research (CIDR). CIDR is fully funded through a federal contract from the National Institutes of Health to The Johns Hopkins University, Contract No. HHSN268200782096. In memory of Theodore Reich, founding principal investigator of COGEND, we are indebted to his leadership in the establishment and nurturing of COGEND and acknowledge with great admiration his seminal scientific contributions to the field. Lead investigators directing data collection are LJ Bierut, Naomi Breslau, Dorothy Hatsukami and Eric Johnson. We thank Heidi Kromrei and Tracey Richmond for their assistance in data collection. HBCS: We acknowledge financial support from the Academy of Finland (Grant No. 120315 and 129287 to EW, 1129457 and 1216965 to KR, 120386 and 125876 to JGE), the European Science Foundation (EuroSTRESS), the Wellcome Trust (Grant No. 89061/Z/09/Z and 089062/Z/09/Z) and the Signe and Ane Gyllenberg foundation. NAG/IRPG: This study is supported by NIH Grants DA12854 (to PAFM), AA07728, AA07580, AA11998, AA13320 and AA13321 (to ACH); and grants from the Australian National Health and Medical Research Council; MLP is supported by DA019951. QIMR: We thank Marlene Grace and Ann Eldridge for sample collection; Anjali Henders, Megan Campbell, Lisa Bowdler, Steven Crooks and staff of the Molecular Epidemiology Laboratory for sample processing and preparation; Harry Beeby, David Smyth and Daniel Park for IT support. We acknowledge support from the Australian Research Council (A7960034, A79906588, A79801419, DP0212016, DP0343921), Beyond Blue and the Borderline Personality Disorder Research Foundation. Genotyping was funded by the National Health and Medical Research Council (Medical Bioinformatics Genomics Proteomics Program, 389891). Further, we gratefully acknowledge Drs Dale R Nyholt and especially Scott Gordon for their substantial efforts involving the QC and preparation of the QIMR and NAG/IRPG GWA data sets. Dr Nyholt also contributed 8% of the NAG/IRPG GWA cohort (NHMRC IDs 339462, 442981, 389938, 496739). LBC1936: We thank David Liewald and Paul Redmond for technical assistance; the study Secretary Paula Davies; Alan Gow, Michelle Taylor, Janie Corley, Caroline Brett and Caroline Cameron for data collection and data entry; nurses and staff at the Wellcome Trust Clinical Research Facility, where subjects were tested and genotyping was performed; staff at the Lothian Health Board, and the staff at the SCRE Centre, University of Glasgow. The research was supported by a program grant from Research Into Ageing. The research continues with program grants from Help the Aged/Age Concern (The Disconnected Mind). GWA funding awarded by the Biotechnology and Biological Sciences Research Council (BBSRC) to IJD and AT. ML is a Royal Society of Edinburgh/Lloyds TSB Foundation for Scotland Personal Research Fellow. The study was conducted within the University of Edinburgh Centre for Cognitive Ageing and Cognitive Epidemiology, supported by the (BBSRC), Engineering and Physical Sciences Research Council (EPSRC), Economic and Social Research Council (ESRC) and Medical Research Council (MRC), as part of the cross-council Lifelong Health and Wellbeing Initiative. This work has made use of the resources provided by the Edinburgh Compute and Data Facility (ECDF) (http://www.ecdf.ed.ac.uk/). The ECDF is partially supported by the eDIKT initiative (http://www.edikt.org.uk). Baltimore Longitudinal Study of Aging: We acknowledge support from the Intramural Research Program of the NIH, National Institute on Aging. We thank Robert McCrae. EGPUT: AM and TE received support from FP7 Grants (201413 ENGAGE, 212111 BBMRI, ECOGENE (No. 205419, EBC)) and OpenGENE. AM and TE also received targeted financing from Estonian Government SF0180142s08 and by EU via the European Regional Development Fund, in the frame of Centre of Excellence in Genomics. The genotyping of the Estonian Genome Project samples were performed in Estonian Biocentre Genotyping Core Facility, AM and TE thank Mari Nelis and Viljo Soo for their contributions. AR and JA were supported by a grant from the Estonian Ministry of Science and Education (SF0180029s08).
Author contributions
Writing group: MHMdeM, PTC, ATer., RFK, CMvanD, DIB. Analytic group: MHMdeM, J-JH, TE, ML, TT, SS, ATen, LML, NKH, SEM, NRW, EW, DLC, KR, GRA, NA. Study design and project management: DIB, EJCdeG, PSu, BWJHP, PAFM, MLP, AM, IJD, MJW, NGM, NRW, GWM, JGE, AP, LP, KR, MU, LF, DS, CMvanD, BAO, PTC, ATer. Sample and phenotype data collection: MAD, GW, EJCdeG, BWJHP, PSp, AM, AR, JA, PAFM, ACH, NGM, MLP, MJW, NGM, NRW, LJB, KR, JGE, MU, LF, DS, ACJW, PTC, ATer. Data preparation: MHMdeM, MAD, J-JH, GW, EJCdeG, CAH, TE, AR, MLP, GD, ML, ATen, LML, SEM, NKH, PL, RG, AA, JD, EW, DLC, YSA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
PTC Jr received royalties from the NEO Five-Factor Inventory. LJB is an inventor on the patent ‘Markers for Addiction’ (US 20070258898) covering the use of certain SNPs in determining the diagnosis, prognosis and treatment of addiction. Dr LJB served as a consultant for Pfizer Inc. in 2008. The other authors declare that they have no potential conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
de Moor, M., Costa, P., Terracciano, A. et al. Meta-analysis of genome-wide association studies for personality. Mol Psychiatry 17, 337–349 (2012). https://doi.org/10.1038/mp.2010.128
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2010.128
Keywords
This article is cited by
-
Functional and molecular characterization of suicidality factors using phenotypic and genome-wide data
Molecular Psychiatry (2023)
-
The molecular genetic basis of creativity: a mini review and perspectives
Psychological Research (2023)
-
Phenotypic and genetic analysis of a wellbeing factor score in the UK Biobank and the impact of childhood maltreatment and psychiatric illness
Translational Psychiatry (2022)
-
Non-genetic linkage of personality traits and the divergence of Eastern and Western cultures: association with Hofstede’s cultural dimensions
Culture and Brain (2022)
-
Uncovering the genetic architecture of broad antisocial behavior through a genome-wide association study meta-analysis
Molecular Psychiatry (2022)