Abstract
TP53 deletion (ΔTP53) in myeloma is known to be a high-risk finding associated with poorer prognosis. The prognostic impact of underlying cytogenetic heterogeneity in patients with myeloma associated with ΔTP53 is unknown. We studied 90 patients with myeloma associated with ΔTP53 identified by interphase fluorescence in situ hybridization and assessed the impact of karyotype and coexisting alterations of IGH, RB1, and CKS1B. There were 54 men and 36 women with a median age of 59 years (range 38–84); 14 patients had a normal karyotype (NK/ΔTP53), 73 had a complex karyotype (CK/ΔTP53), and 3 had a non-complex abnormal karyotype. Patients with CK/ΔTP53 showed a significantly poorer overall survival compared with patients with NK/ΔTP53 (P=0.0243). Furthermore, in the CK/ΔTP53 group, patients with IGH rearrangement other than t(11;14)(q13;q32)/CCND1-IGH, designated as adverse-IGH, had an even worse outcome (P=0.0045). In contrast, RB1 deletion, CKS1B gain, ploidy, additional chromosome 17 abnormalities, or ΔTP53 clone size did not impact prognosis. Stem cell transplant did not improve overall survival in either the NK/ΔTP53 or CK/ΔTP53 (P=0.8810 and P=0.1006) groups, but tandem stem cell transplant did improve the overall survival of patients with CK/ΔTP53 (P=0.0067). Multivariate analysis confirmed in this cohort that complex karyotype (hazard ratio 1.976, 95% CI 1.022–3.821, P=0.043), adverse-IGH (hazard ratio 3.126, 95% CI 1.192–8.196, P=0.020), and tandem stem cell transplant independently correlate with overall survival (hazard ratio 0.281, 95% CI 0.091–0.866, P=0.027). We conclude that comprehensive genetic assessment adds to TP53 status in the risk stratification of myeloma patients.
Similar content being viewed by others
Main
Multiple myeloma is a clinically and molecularly heterogeneous disease with an overall survival ranging from <1 year to over 20 years.1, 2 Reliable risk stratification is a key to predicting outcome and guiding therapy. In 2015, the International Myeloma Working Group proposed the Revised International Staging System (R-ISS) for myeloma patients.3 The system incorporates chromosomal abnormalities detected by interphase fluorescence in situ hybridization (FISH) and serum lactate dehydrogenase levels into the preexisting International Staging System (ISS), which was based on serum albumin and beta-2 microglobulin levels.4 TP53 deletion (ΔTP53) is also a known negative prognostic factor5, 6, 7 and is considered a high-risk chromosomal abnormality in the R-ISS. ΔTP53 usually results from deletion of chromosome 17p and is likely a secondary event that is often associated with disease progression.1
It is unclear if the negative prognostic impact of ΔTP53 is influenced by background chromosomal aberrations such as ploidy, karyotypic complexity, or other recurrent changes that have been reported in myeloma.8 For example, does ΔTP53 detected by FISH carry the same prognostic impact as 17p deletion detected by conventional karyotypic analysis? Does clone size of ΔTP53 as determined by FISH impact outcome? Does the presence of ΔTP53 trump other known adverse factors, such as t(4;14) or t(14;16)? Lastly, might the combination of ΔTP53 and t(4;14)/t(14;16) have a synergistic poorer effect on the prognosis of myeloma patients?
In this study, we investigated the prognostic impact of underlying cytogenetic heterogeneity in a group of patients with myeloma associated with ΔTP53, focusing on well-defined cytogenetic risk subgroups, with the goal of refining the prognostic impact of ΔTP53. Our data suggest that comprehensive assessment of genetic abnormalities in myeloma adds value to TP53 status alone in the risk stratification of patients with myeloma.
Materials and methods
Study Group
We retrospectively reviewed multiple myeloma cases assessed by conventional cytogenetics and tested for TP53 by interphase FISH at The University of Texas MD Anderson Cancer Center between 1 December 2007 and 31 December 2014. Clinical and laboratory data including hemoglobin level and serum beta-2 microglobulin and lactate dehydrogenase levels were obtained from a review of the electronic medical record. Disease stage was assessed using ISS and R-ISS. The study was approved by the Institutional Review Board.
In our laboratory, we did not routinely performed plasma cell enrichment on bone marrow aspirate specimens prior to culture until about 4 years ago. To ensure that all cases of myeloma in this study had a sufficient number of cells to avoid false-negative results, we required that all myeloma cases included in this study had 30% or more plasma cells on aspirate smears.
Cytogenetics and FISH
Conventional karyotyping was performed on cultured (unstimulated 24 and 48 h) bone marrow aspirate samples as part of the routine clinical workup and following laboratory standard procedures as have been reported.9 At least 20 metaphases were fully analyzed whenever possible for the identification of clonal cytogenetic aberrations. The karyotypic results were reported according to the International System for Human Cytogenetic Nomenclature (ISCN) 2009 and 2013. In this system, a clone is defined as chromosomal changes in two or more metaphases and a complex karyotype (CK) is defined as ≥3 chromosome abnormalities.
FISH analyses for TP53/CEN17, MYEOV/CCND1-IGH/t(11;14), RB1 (Vysis-Abbott Molecular, Downers Grove, IL, USA) and CKS1B (CytoCell, Cambridge, UK) were performed on cultured (unstimulated 24 and 48 h) bone marrow samples. Following laboratory standard protocols, all FISH cutoff values were established statistically based on the testing results on peripheral blood or bone marrow specimens from at least 20 normal healthy individuals. For TP53, we used a centromeric region of chromosome 17 as a control and the cutoff for a 200 interphase cell count was 4.7%. A clone size, that is, for the TP53 deletion was determined by the number or the percentage of positive cells in 200 interphase cells analyzed. The clone size for TP53 deletion was also further assessed by comparing it with results of RB1 and/or IGH-CCND1 tested concurrently and with the plasma cell percentage reported on the bone marrow aspirate differential count. FISH for FGFR3-IGH/t(4;14), MAF-IGH/t(14;16) were often performed (as a part of reflex testing) when FISH for MYEOV/CCND1-IGH/t(11;14)(q13;q32) was negative but with evidence of IGH rearrangement.
Patients with ΔTP53 were divided initially into two major subgroups based on karyotype: normal karyotype (NK/ΔTP53) or complex karyotype (CK/ΔTP53). The CK/ΔTP53 group, the main focus of this study, was also divided into subgroups (Figure 1) based on the presence or absence of IGH translocations. Among the patients with IGH translocations, there were three small subsets: t(4;14)/t(14;16), t(11;14), and other translocations with variable (v) loci designated here as t(v;14). Cytogenetically visible chromosome 17 aberrations, such as monosomy 17 (−17) or chromosome 17p deletion (17p−), as well as chromosome ploidy, including hypodiploidy (hypo) (chromosome count <44), hyperdiploidy (hyper) (chromosome count ≥48), and near-diploidy (chromosome count 44–47) were further analyzed in cases without IGH translocations and cases with t(11;14).
In addition, two independent groups of myeloma patients lacking ΔTP53, one with a normal karyotype (NK), and the other with a complex karyotype (CK), designated as NK/TP53nl and CK/TP53nl, respectively, were included as control groups to compare with the NK/ΔTP53 and CK/ΔTP53 study groups.
Survival Analysis
The follow-up interval and overall survival were calculated from the time of initial diagnosis until time of last follow-up or death. Statistical analyses were performed with the GraphPad Prism 6 software (GraphPad Software, La Jolla, CA, USA). Kaplan–Meyer curves for overall survival were plotted and the log-rank test was applied with a P-value <0.05 being considered as statistically significant. Multivariate analysis was performed using SPSS version 9.3 (SPSS Institute, Chicago, IL, USA). A cutoff of 50% for plasma cells and a clone size of 50% for TP53 deletion (ΔTP53) were applied to assess the impact on overall survival in the multivariate analysis.
Results
Clinicopathologic Data
From a total of 1036 myeloma patients assessed during the study interval, 90 (9%) patients had ΔTP53, including 54 men and 36 women with a median age of 59 years (range, 38–84) at the time of diagnosis. Fourteen (16%) patients had a normal karyotype, 73 (81%) had a complex karyotype, and 3 (3%) had a single chromosomal aberration (Figure 1). Demographic and clinical characteristics for patients in the NK/ΔTP53 and CK/ΔTP53 groups showed no significant differences for age, gender, or serum levels of hemoglobin, albumin, or creatinine. However, in the CK/ΔTP53 group, patients had higher serum beta-2 microglobulin (P<0.001) and lactate dehydrogenase (P=0.015) levels and higher percentage of bone marrow plasma cells (P=0.003) when compared with patients in the NK/ΔTP53 group, respectively (Table 1).
In the NK/ΔTP53 group, 29% of patients had stage I disease, 50% stage II, and 14% stage III compared with the CK/ΔTP53 group in which 15% of patients had stage I disease, 34% stage II, and 48% stage III (Table 2). When the R-ISS was applied to the study cohort there was a significant difference in patients with stage II disease: 79% of NK/ΔTP53 and 49% of the CK/ΔTP53 patients (P=0.038). No patients in either group had stage I disease and the percentage of patients with stage III disease was similar in both groups.
Of the 14 patients in the NK/ΔTP53 group, 4 (29%) were newly diagnosed and 10 (71%) had either persistent or relapsed disease. In the CK/ΔTP53 group, 11 (15%) patients were newly diagnosed and 56 (77%) had either persistent or relapsed disease; disease status in the six patients is unknown (Table 2).
Cytogenetic Heterogeneity in NK/ΔTP53 and CK/ΔTP53 Patients Groups
Among the 73 patients with myeloma associated with CK/ΔTP53, 39 had IGH translocations, including 16 with t(11;14), 16 with t(4;14)/t(14;16), and 7 with t(v;14). The other 34 patients had no IGH translocations. Combining patients with t(11;14) and patients without IGH rearrangement, that is, the group considered to be favorable (n=50), we found 19 hyperdiploid, 15 hypodiploid, and 16 near-diploid cases. The chromosome 17 findings in these 50 cases showed 24 with 17p deletion (17p−), 14 monosomy 17 (−17), and 12 with no gross abnormalities (Figure 1).
Overall, RB1 deletion in myeloma was detected in 9 of 14 (64%) NK/ΔTP53 and 55 of 73 (75%) CK/ΔTP53 cases. MYEOV/CCND1-IGH/t(11;14) was positive in 2 of 14 (14%) NK/ΔTP53 and 16 of 73 (22%) CK/ΔTP53 myeloma groups. FGFR3-IGH/t(4;14) or MAF-IGH/t(14;16) was identified in 16 cases in the CK/ΔTP53, but not in the NK/ΔTP53 myeloma groups; however, t(v;14) was detected in 2 NK/ΔTP53 and 7 CK/ΔTP53 cases. CKS1B gain was observed in 2 of 4 (50%) tested in the NK/ΔTP53 group and 26 of 32 (81%) cases tested in the CK/ΔTP53 group. There were no significant differences shown by FISH between the NK/ΔTP53 and CK/ΔTP53 groups with regard to the frequency of aberrancies (Table 3). The median TP53 deletion clone size in the NK/ΔTP53 group was 13% (range 8–40%) vs 30% (range 5–95%) in the CK/ΔTP53 group (P<0.001) (Table 1).
Cytogenetic Heterogeneity and Clinical Implications
All patients received standard clinical management with immunomodulatory drugs, proteasome inhibitors, and/or stem cell transplant (SCT). Among the patients with detailed treatment data available, all 14 (100%) patients in the NK/ΔTP53 group, 66 (90%) in the CK/ΔTP53 group, 22 (73%) in the NK/TP53nl group, and all 16 (100%) in the CK/TP53nl group (P=0.019) received proteasome inhibitors (Table 2 and Supplementary Table S1). The NK/TP53nl patient group was least often treated with proteasome inhibitors; there were no statistical differences between the CK/ΔTP53 and the CK/TP53nl patient groups. In addition, 10 (71%) patients in the NK/ΔTP53 group, 46 (63%) in the CK/ΔTP53 group, 22 (73%) in NK/TP53nl group, and 14 (88%) patients in the CK/TP53nl group received an SCT, respectively (P=0.258) (Table 2 and Supplementary Table S1). During the follow-up interval, 6 (43%) NK/ΔTP53 patients and 55 (75%) CK/ΔTP53 patients died, with the latter group showing a higher mortality rate (P=0.015) (Table 2).
The median follow-up interval for all patients in the study was 32 months (range, 3–101 months) with the NK/ΔTP53 group showing a longer follow-up interval (P=0.029) (Table 2). The overall survival was 62 months for patients in the NK/ΔTP53 group and 35 months for patients in the CK/ΔTP53 group (P=0.024) (Figure 2a).
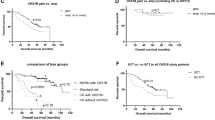
(a) Comparison of overall survival between NK/ΔTP53 (curve 1) and CK/ΔTP53 (curve 2) groups. (b) Comparison of overall survival (OS) among four subgroups: NK/TP53nl (curve 1), NK/ΔTP53 (curve 2), CK/TP53nl (curve 3), and CK/ΔTP53 (curve 4). (c) Comparison of overall survival between cases without (curve 1) and with (curve 2) adverse-IGH rearrangements in the CK/ΔTP53 group.
To further assess overall survival, we included patients without ΔTP53 and either a normal or complex karyotype designated as NK/TP53nl (n=35) and CK/TP53nl (n=21) from the same time interval to serve as the cytogenetic control groups. Median overall survival was 150 and 55 months for the NK/TP53nl and CK/TP53nl patient groups, respectively. When all four groups were compared, patients in the CK/ΔTP53 group showed the poorest overall survival whereas patients in the NK/TP53nl group had the best outcome (P<0.0001) (Figure 2b). Patients with myeloma associated with NK/ΔTP53 or CK/TP53nl had similar overall survival (62 vs 55 months), showing that a complex karyotype alone has an independent adverse impact and suggesting that a complex karyotype has an impact comparable to that of TP53 deletion.
In the CK/ΔTP53 group, the overall survival of 16 patients with t(4;14)/t(14;16) vs 7 patients with t(v;14) was similar, 23 and 33 months, respectively (P=0.3289). Combining these patients into an adverse-IGH rearrangement subgroup, the median overall survival was 23 months for those with adverse-IGH rearrangement vs 40 months for patients without adverse-IGH rearrangement (P=0.0045) (Figure 2c), suggesting an additional negative impact of adverse-IGH rearrangement within the CK/ΔTP53 subgroup.
We correlated ploidy and overall survival in 50 patients who had no adverse-IGH rearrangement, that is, patients with CCND1-IGH or no IGH rearrangement. The median overall survival was 60 vs 39 vs 32 months for hyperdiploid, near-diploid, and hypodiploid subgroups, respectively. Although the hyperdiploid subgroup showed the best overall survival, this difference did not reach statistical significance (P=0.3619) (Figure 3a). RB1 deletion or t(11;14)/CCND1-IGH had no independent impact on overall survival for patients in the CK/ΔTP53 group (Figures 3b and c).
(a) Overall survival of patients stratified according to chromosome ploidy status in the CK/ΔTP53 group (curve 1, hyperdiploidy without adverse-IGH; curve 2, near-diploidy without adverse-IGH; curve 3, hypodiploidy without adverse-IGH). (b) Comparison of overall survival between cases without (curve 1) and with (curve 2) RB1 deletion/without adverse-IGH in the CK/ΔTP53 group. (c) Comparison of overall survival between cases without IGH rearrangement (curve 1) and with t(11;14) (curve 2) in the CK/ΔTP53 group. (d) Comparison of overall survival among all non-adverse-IGH cases without chromosome 17 aberrations (curve 1), with 17p deletion (curve 2) and monosomy 17 (curve 3).
To investigate the possible prognostic impact of cytogenetically visible chromosome 17 abnormalities, we divided the 50 cases without adverse-IGH rearrangements in the CK/ΔTP53 group into three subsets: (1) cases with apparent 17p deletions (n=24); (2) cases with monosomy 17 (n=14); and (3) cases with an apparently normal 17 (n=12). The median overall survival was 42 vs 32 vs 38 months among these three subsets, respectively (P=0.9522) (Figure 3d).
Additionally, within the CK/ΔTP53 group, the median clone size of ΔTP53 was 30% (range 5–95%). Patients with a clone size below or above 30% showed no significant difference in overall survival (P=0.3759). The median plasma cell count was 78% in the CK/ΔTP53 group, and there was also no overall survival differences between patients with ≤78% and >78% plasma cells. In the CK/ΔTP53 group, we also compared the ΔTP53 clone size with the percentage of plasma cells and clone size of other altered genes including RB1, IGH, and CKS1B. We estimate that ΔTP53 likely a part of a primary (or major) clone in 53 (73%) patients whereas it was more likely a part of a secondary clone (or subclone) in 20 (27%) patients. There was no significant difference in overall survival between patients with ΔTP53 as a primary vs secondary clone.
Impact of SCT on Overall Survival
Ten (71%) patients in the NK/ΔTP53 group and 46 (63%) patients in the CK/ΔTP53 group received an SCT. In the NK/ΔTP53 group, patients who received a transplant showed no survival advantage over patients who did not receive transplant (P=0.8810) (Figure 4a). By contrast, in the CK/ΔTP53 group, although SCT did not improve the median overall survival (46 months with SCT vs 24 months without SCT; P=0.1006) (Figure 4b), patients who received a second (or tandem) SCT showed a significantly better overall survival (79 months, P=0.0067) (Figure 4c).
a) Comparison of overall survival between cases with (curve 1) and without (curve 2) stem cell transplant in the NK/ΔTP53 group. (b) Comparison of overall survival in cases with (curve 1) and without (curve 2) stem cell transplant in the CK/ΔTP53 group. (c) Comparison of overall survival among cases with tandem or more stem cell transplant (curve 1), one stem cell transplant (curve 2), and without stem cell transplant (curve 3) in the CK/ΔTP53 group.
Multivariate Analysis of Prognostic Factors
Multivariate analysis confirmed that a complex karyotype (hazard ratio 1.976, 95% CI 1.022–3.821, P=0.043) and adverse IGH rearrangement (hazard ratio 3.126, 95% CI 1.192–8.196, P=0.020) showed an independent negative impact on overall survival. In contrast, tandem SCT correlated with improved overall survival (hazard ratio 0.281, 95% CI 0.091–0.866, P=0.027) (Table 4).
Discussion
The widely disparate survival outcomes observed in patients with myeloma can be attributed to tumor genetic heterogeneity and host factors. Mounting evidence supports the idea that the genetic features of myeloma can be used to predict the outcome of these patients. To better understand the clinical implications of background cytogenetic heterogeneity in patients with myeloma associated with TP53 deletion (ΔTP53), we studied the clinicopathologic features of patients with a plasma cell count ≥30% and ΔTP53 identified by FISH in the context of a normal karyotype (NK) vs complex karyotype (CK). In patients with myeloma associated with ΔTP53, we show that patients with a normal karyotype have a significantly longer overall survival than patients with a complex karyotype. Patients with myeloma associated with a complex karyotype but without TP53 deletion also had a significantly worse overall survival than patients with NK/ΔTP53 myeloma, suggesting that a complex karyotype is an important adverse risk factor in myeloma patients.
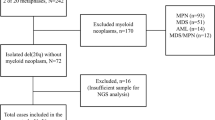
The frequency of TP53 deletion in this cohort was approximately 9%, which is consistent with what has been reported previously.1 Although others in some studies have reported a much higher frequency of TP53 deletion, ranging from 39%6 to 55%,7 we believe that these studies were likely focused on patients with advanced stage III disease, plasma cell leukemia, or relapsed disease. In contrast, this study cohort was derived from a pool of patients with myeloma at various stages of disease, including both newly diagnosed and relapsed disease, with very few plasma cell leukemia patients. Although our data were generated on non-CD138-enriched plasma cells, we only included patients with a plasma cell count 30% or higher in this study. By doing the study in this manner, we have overcome some of the known technical limitations of FISH in the analysis of non-enriched samples. In keeping with the latest recommendation by the International Myeloma Working Group, future FISH studies should be performed on plasma cell-enriched samples.
The R-ISS proposed recently by the International Myeloma Working Group for multiple myeloma considers chromosome abnormalities detected by FISH a key element in defining the biological features of myeloma. However, the role of chromosomal abnormalities detected by conventional karyotyping in the staging of myeloma patients was not clearly described. The results presented here show that knowledge of the karyotype in which ΔTP53 is detected is important: deletion detected by FISH associated with a normal karyotype does not have the same prognostic impact as does ΔTP53 associated with a complex karyotype; patients in the latter group have significantly shorter survival.
An interesting finding in this study is that patients with myeloma associated with ΔTP53 can have a normal or complex (≥3 abnormalities) karyotype, with very few patients having only one or two cytogenetic abnormalities. The reason for this apparent dichotomy is unknown, but the data suggest that ΔTP53 may occur in one of two settings: (1) an early and a likely primary event in pathogenesis; and (2) a late and/or secondary event. Most likely, a complex karyotype would be more frequent in the second group. Further studies with larger study groups may help elucidate the underlying mechanisms.9 By limiting our study cases to those with 30% of plasma cells, we aim to eliminate the possibility of a false normal karyotype result due to insufficient tumor cells.
Previous studies of myeloma patients have shown that different cytogenetic aberrations are associated with different clinical outcomes.10, 11, 12, 13, 14, 15, 16, 17, 18 Myeloma patients with t(4;14)/t(14;16) or other adverse IGH rearrangements were also found to have shorter overall survival and therefore considered to be of high-risk,19, 20 whereas myeloma patients with t(11;14) are considered low risk with a more favorable prognosis.21 In this study, we show that ΔTP53 combined with t(4;14) or t(14;16) conferred an even worse outcome, suggesting a ‘double hit’ effect. Furthermore, a complex karyotype in the context of ΔTP53 and adverse IGH rearrangement likely further contributes an even worse clinical outcome in myeloma patients. The finding is in keeping with results reported by others. A possible explanation is that the myeloma may carry multiple drug resistant clones or that the combination of multiple cytogenetic aberrations further enhances drug resistance.
Additional examination of chromosomal aberrations in myeloma cases without adverse IGH in the CK/ΔTP53 group did not show independent prognostic effects for monosomy 17 or 17p−, unlike cytogenetically visible chromosome 13q deletions in other myeloma studies.22 The median clone size in the CK/ΔTP53 group was 30% and the median plasma cell count was 78%. Although our data showed that larger clone size of ΔTP53 or higher plasma cell count did not have independent clinical impact, both findings were highly associated with a complex karyotype, which is an independent negative prognostic factor by multivariate analysis.
Myeloma can be broadly subdivided into three subsets at the chromosomal level based on ploidy: hyperdiploid, hypodiploid, and near-diploid. In general, hyperdiploid myeloma has a lower prevalence of IGH translocations compared with non-hyperdiploid myeloma.23, 24, 25, 26 Others27 have shown that hypodiploid myeloma is associated with a higher prevalence of genetic alterations and inferior outcome compared with non-hypodiploid myeloma. In the current study, although patients with hyperdiploidy in the CK/ΔTP53 group did have better overall survival, there was no statistical difference between patients in the hypodiploid or near-diploid groups, likely due to ΔTP53 and a complex karyotype overwhelming the prognostic impact of ploidy.
Although our data showed that SCTs did not improve overall survival for the overall study cohort, tandem SCT did correlate with improved overall survival, particularly in patients with CK/ΔTP53. This finding further underscores the critical role of a complex karyotype in the risk assessment and management of myeloma patients.28
In conclusion, we assessed the impact of underlying cytogenetic heterogeneity in patients with myeloma associated with ΔTP53 detected by conventional karyotype and FISH. The data presented show that conventional chromosomal analysis remains a powerful tool for risk assessment which supplements FISH data, particularly if high-resolution genomic analysis such as microarray-based testing is not available. For patients with ΔTP53 detected by FISH, those patients with a normal karyotype had a significantly longer overall survival than patients with a complex karyotype. In addition, combined TP53 deletion and adverse IGH conferred a worse clinical outcome. We therefore suggest that conventional cytogenetic analysis has a role in the work up of myeloma patients because it facilitates comprehensive assessment of molecular abnormalities in myeloma, adding value to TP53 status, in the risk stratification of patients with myeloma.
References
Brenner H, Gondos A, Pulte D . Recent major improvement in long-term survival of younger patients with multiple myeloma. Blood 2008;111:2521–2526.
McKenna RW, Kyle RA, Kuehl WM et al, Plasma cell neoplasms. In: Swerdlow SH, Campo E, Harris NL et al, (eds). WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. IARC Press: Lyon, France, 2008, pp 200–213.
Palumbo A, Avet-Loiseau H, Oliva S et al. Revised International Staging System for Multiple Myeloma: a report from International Myeloma Working Group. J Clin Oncol 2015;33:2863–2869.
Greipp PR, San Miguel J, Durie BG et al. International staging system for multiple myeloma. J Clin Oncol 2005;23:3412–3420.
Avet-Loiseau H, Li JY, Godon C et al. P53 deletion is not a frequent event in multiple myeloma. Br J Haematol 1999;106:717–719.
Drach J, Ackermann J, Fritz E et al. Presence of a p53 gene deletion in patients with multiple myeloma predicts for short survival after conventional-dose chemotherapy. Blood 1998;92:802–809.
Neri A, Baldini L, Trecca D et al. p53 gene mutations in multiple myeloma are associated with advanced forms of malignancy. Blood 1993;81:128–135.
Fonseca R, Barlogie B, Bataille R et al. Genetics and cytogenetics of multiple myeloma: a workshop report. Cancer Res 2004;64:1546–1558.
Lu G, Muddasani R, Orlowski RZ et al. Plasma cell enrichment enhances detection of high-risk cytogenomic abnormalities by fluorescence in situ hybridization and improves risk stratification of patients with plasma cell neoplasms. Arch Pathol Lab Med 2013;137:625–631.
Boyd KD, Ross FM, Chiecchio L et al. A novel prognostic model in myeloma based on co-segregating adverse FISH lesions and the ISS: analysis of patients treated in the MRC Myeloma IX trial. Leukemia 2012;26:349–355.
Boyd KD, Ross FM, Tapper WJ et al. The clinical impact and molecular biology of del(17p) in multiple myeloma treated with conventional or thalidomide-based therapy. Genes Chromosomes Cancer 2011;50:765–774.
Chang H, Qi C, Yi QL et al. p53 gene deletion detected by fluorescence in situ hybridization is an adverse prognostic factor for patients with multiple myeloma following autologous stem cell transplantation. Blood 2005;105:358–360.
Chang H, Sloan S, Li D et al. The t(4;14) is associated with poor prognosis in myeloma patients undergoing autologous stem cell transplant. Br J Haematol 2004;125:64–68.
Fonseca R, Blood E, Rue M et al. Clinical and biologic implications of recurrent genomic aberrations in myeloma. Blood 2003;101:4569–4575.
Gertz MA, Lacy MQ, Dispenzieri A et al. Clinical implications of t(11;14)(q13;q32), t(4;14)(p16.3;q32), and -17p13 in myeloma patients treated with high-dose therapy. Blood 2005;106:2837–2840.
Hanamura I, Stewart JP, Huang Y et al. Frequent gain of chromosome band 1q21 in plasma-cell dyscrasias detected by fluorescence in situ hybridization: incidence increases from MGUS to relapsed myeloma and is related to prognosis and disease progression following tandem stem-cell transplantation. Blood 2006;108:1724–1732.
Moreau P, Facon T, Leleu X et al. Recurrent 14q32 translocations determine the prognosis of multiple myeloma, especially in patients receiving intensive chemotherapy. Blood 2002;100:1579–1583.
Walker BA, Leone PE, Chiecchio L et al. A compendium of myeloma-associated chromosomal copy number abnormalities and their prognostic value. Blood 2010;116:e56–e65.
Avet-Loiseau H, Malard F, Campion L et al. Translocation t(14;16) and multiple myeloma: is it really an independent prognostic factor? Blood 2011;117:2009–2011.
Kalff A, Spencer A . The t(4;14) translocation and FGFR3 overexpression in multiple myeloma: prognostic implications and current clinical strategies. Blood Cancer J 2012;2:e89.
An G, Xu Y, Shi L et al. t(11;14) multiple myeloma: a subtype associated with distinct immunological features, immunophenotypic characteristics but divergent outcome. Leuk Res 2013;37:1251–1257.
Avet-Loiseau H, Facon T, Grosbois B et al. Oncogenesis of multiple myeloma: 14q32 and 13q chromosomal abnormalities are not randomly distributed, but correlate with natural history, immunological features, and clinical presentation. Blood 2002;99:2185–2191.
Fonseca R, Bergsagel PL, Drach J et al. International Myeloma Working Group molecular classification of multiple myeloma: spotlight review. Leukemia 2009;23:2210–2221.
Debes-Marun CS, Dewald GW, Bryant S et al. Chromosome abnormalities clustering and its implications for pathogenesis and prognosis in myeloma. Leukemia 2003;17:427–436.
Fonseca R, Debes-Marun CS, Picken EB et al. The recurrent IgH translocations are highly associated with nonhyperdiploid variant multiple myeloma. Blood 2003;102:2562–2567.
Smadja NV, Fruchart C, Isnard F et al. Chromosomal analysis in multiple myeloma: cytogenetic evidence of two different diseases. Leukemia 1998;12:960–969.
Van Wier S, Braggio E, Baker A et al. Hypodiploid multiple myeloma is characterized by more aggressive molecular markers than non-hyperdiploid multiple myeloma. Haematologica 2013;98:1586–1592.
Greslikova H, Zaoralova R, Filkova H et al. Negative prognostic significance of two or more cytogenetic abnormalities in multiple myeloma patients treated with autologous stem cell transplantation. Neoplasma 2010;57:111–117.
Acknowledgements
We thank the technologists in the Clinical Cytogenetics Laboratory at MD Anderson Cancer Center for their contributions. XL and PL designed the study; SH, PL, RLS and XL collected data; XL, SK, and LF performed statistical analyses, SH, PL, LJM, and XL analyzed data and wrote the manuscript; JS, DM, ST, EM, and RZO managed the patients and provided treatment data. All authors approved the final version of the manuscript. RZO acknowledges support from The MD Anderson Cancer Center SPORE in Multiple Myeloma (P50 CA142509), The MD Anderson Cancer Center Support Grant (P30 CA016672), the National Cancer Institute (U10 CA032102, R01 CA184464, CA194264), and the MD Anderson Cancer Center High Risk Multiple Myeloma Moonshot. We thank Wintress Dennis for technical assistance of the manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Hao, S., Lin, P., Medeiros, L. et al. Clinical implications of cytogenetic heterogeneity in multiple myeloma patients with TP53 deletion. Mod Pathol 30, 1378–1386 (2017). https://doi.org/10.1038/modpathol.2017.63
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.63