Abstract
The Cancer Genome Atlas data on colorectal carcinoma have provided a comprehensive view of the tumor’s genomic alterations and their tumorigenic roles. Tumor morphology, however, has not been fully integrated into the analysis. The aim of this study was to explore relevant associations between tumor morphology and the newly characterized genomic alterations in colorectal carcinoma. Two hundred and seven colorectal carcinomas that had undergone whole-exome sequencing as part of The Cancer Genome Atlas project and had adequate virtual images in the cBioPortal for Cancer Genomics constituted our study population. Upon analysis, a tight association between ‘microsatellite instability-high histology’ and microsatellite instability-high (P<0.001) was readily detected and helped validate our image-based histology evaluation. Further, we showed, (1) among all histologies, the not otherwise specified type had the lowest overall mutation count (P<0.001 for entire cohort, P<0.03 for the microsatellite-instable group), and among the microsatellite-instable tumors, this type also correlated with fewer frameshift mutations in coding mononucleotide repeats of a defined set of relevant genes (P<0.01); (2) cytosine phosphate guanine island methylator phenotype-high colorectal cancers with or without microsatellite instability tended to have different histological patterns: the former more often mucinous and the latter more often not otherwise specified; (3) mucinous histology was associated with more frequent alterations in BRAF, PIK3CA, and the transforming growth factor-β pathway when compared with non-mucinous histologies (P<0.001, P=0.01, and P<0.001, respectively); and (4) few colorectal cancers (<9%) exhibited upregulation of immune-inhibitory genes including major immune checkpoints; these tumors were primarily microsatellite-instable (up to 43%, vs <3% in microsatellite-stable group) and had distinctly non-mucinous histologies with a solid growth. These morphology–molecular associations are interesting and propose important clinical implications. The morphological patterns associated with alterations of immune checkpoint genes bear the potential to guide patient selection for clinical trials that target immune checkpoints in colorectal cancer, and provide directions for future studies.
Similar content being viewed by others
Main
Colorectal neoplasia has traditionally served as a paradigm of tumorigenesis in which the stepwise accumulation of genetic alterations was linked to the tumor’s morphological progression from adenoma to carcinoma.1 The systematic analysis of the process and the insights gained from it exemplify the critical importance of incorporating tumor morphology into cancer research.
In recent years, the advent of next-generation sequencing technology has allowed information on tumor genetics and genomics to be uncovered at a speed and on a scale as yet unprecedented.2, 3, 4, 5 The Cancer Genome Atlas project has used next-generation sequencing and profiled the genomic changes in many different cancer types, providing fully integrated views of the genetic and molecular alterations and their significance for tumorigenesis in these cancers.4
For colorectal cancer, The Cancer Genome Atlas project not only confirmed the initial observations regarding the roles of APC, KRAS, TP53, PIK3CA, and SMAD4 as major contributors to tumorigenesis but also revealed a vast amount of new information including new gene targets and alteration patterns involving a variety of molecular pathways.3, 6 What has not been fully integrated into the analysis, however, is tumor morphology. While an association of morphology with the DNA mismatch-repair pathway in colorectal cancer is well known and this knowledge has shown significant clinical and biological importance,7, 8 it remains to be determined whether there exist additional relevant associations between tumor morphology and the newly understood genomic alterations.
Thus, the aim of this study was to characterize the morphological features of the colorectal tumors studied by The Cancer Genome Atlas project3 and correlate such features with the genomic alterations in these tumors. We hypothesized that insights gained from such correlative analysis would further our understanding of tumor morphogenesis and provide directions for future research efforts that aim at exploring the utility of tumor morphology as a predictive marker for molecular phenotype.
Materials and methods
The study cases comprised 224 primary colorectal carcinomas analyzed by The Cancer Genome Atlas project (http://www.cbioportal.org/).3, 4, 9 The analyses by The Cancer Genome Atlas project included whole-exome sequencing, DNA copy number alteration, promoter methylation, and global mRNA and microRNA expression. In addition, The Cancer Genome Atlas project also included microsatellite instability (MSI) testing by PCR.3, 10 Morphological assessment of all 224 tumors was carried out by a GI pathologist using whole slide digital images available through the cBioportal (http://www.cbioportal.org/).3, 4, 9 As the images in cBioportal are consistently scalable from the lowest power scanning view to a maximum magnification that is roughly comparable to x600, it allowed reliable histologic evaluation, including lymphocyte count.
The following morphological features were evaluated based primarily on standard definitions endorsed by the World Health Organization and the American Joint Committee on Cancer with some modifications: histologic type (conventional with gland formation (termed ‘not otherwise specified’), medullary (defined as more than 50% showing solid growth, syncytial and uniform cytology, and conspicuous lymphocytic infiltration), poorly differentiated (more than 50% showing solid growth, non-medullary), mucinous and signet ring cell (0% vs 1–50% vs >50%)); architectural complexity of the tumor cell component (arbitrarily graded 1–5: 1=simple or well differentiated; 2=some complexity with glands containing apparent secondary structure; 3=more complexity with glands containing tertiary structures; 4=complex glands with some areas of solid growth; 5=solid growth); tumor-infiltrating lymphocytes (referring to lymphocytes within tumor cells and tumor nests, exclusive of stromal lymphocytes) (0=none, 1=less than 10 per medium power field, 2=10–20 per medium power field, 3=more than 20 per medium power field); neutrophil infiltration (0=no apparent neutrophils, 1=easily discernible but low in number and scattered in the stroma, 2=conspicuous with focal abscesses within tumor glands, 3=profuse with intraepithelial neutrophils or abscesses disrupting tumor glands); and dirty necrosis (0=0, 1=low, 2=moderate, 3=high, 4=confluent).
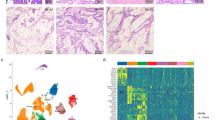
Examples of histological patterns are illustrated in Figure 1. More detailed illustrations of the different grades of glandular architectural complexity are provided in Supplementary Figure 1, and illustrations of varied amounts of tumor-infiltrating lymphocytes and neutrophils are provided in Supplementary Figure 2.
Histological patterns of colorectal carcinoma. (a) A conventional gland-forming adenocarcinoma (not otherwise specified type) with simple architecture and no increased tumor-infiltrating lymphocytes (non-MSI-H, 87 mutations). (b) A conventional gland-forming adenocarcinoma with moderate architectural complexity and elevated tumor-infiltrating lymphocytes (MSI-H, 192 mutations). (c) A mucinous adenocarcinoma with solid strips and clusters of tumor cells floating in mucin (MSI-H, 905 mutations). (d) An adenocarcinoma with high degree of architectural complexity, some medullary-like features, and elevated tumor-infiltrating lymphocytes (POLE-mutated, 6130 mutations). (e) A typical medullary carcinoma with solid and syncytial growth and elevated tumor-infiltrating lymphocytes (MSI-H, 1219 mutations). (f) A poorly differentiated adenocarcinoma with solid growth and some lymphocytes (MSI-H, 655 mutations). MSI-H, microsatellite instability-high; POLE, polymerase epsilon.
Correlative analyses were performed between morphological patterns and genetic or epigenetic alterations. Analyses were performed in three broad groups according to the presence of MSI and polymerase epsilon (POLE) mutations: MSI-H, non-MSI-H, and POLE-mutated. The POLE-mutated group comprised the ‘ultramutated’ tumors that did not show MSI-H.
Molecular alterations evaluated included overall mutation rates, rates of frameshift mutations in coding mononucleotide repeats in 20 genes that have mononucleotides repeats of six nucleotides or longer (MSH3 (A8), MSH6 (C8), MLH3 (T9), TGFβR2 (A10), PRDM2 (A9), PTEN (A6), TCF7L2 (A9), AXIN2 (C7), ACVR2A (A8), ATR (A10), AIM2 (T10), RAD50 (A9), CHEK1 (A9), BLM (A9), CASP5 (T10), GRB14 (A9), MBD4 (T10), WISP3 (A9), SEC63 (T10)6), methylation subtypes (CIMP-H, CIMP-L, cluster 3, and cluster 4), mutations, or copy number alterations of genes (Supplementary Table 1) involved in major colorectal tumorigenic pathways including WNT, mitogen-activated protein kinase (MAPK), phosphoinositol-3-kinase (PI3K), transforming growth factor-β (TGF-β), and TP53 pathways as reported by The Cancer Genome Atlas project, and major chromosomal and subchromosomal changes.3 For chromosomal changes, specific deletion peaks and regions of focal amplification, as well as translocations, were not included in this analysis owing to the small number of cases in each subset, especially when taking into consideration the difference in the sets of genes involved.
In addition to the above major alteration patterns, we also evaluated morphological associations with alterations in genes associated with tumor immunity. Specifically, we evaluated mutations, copy number alterations, and mRNA expression of the following groups of genes: (1) immune checkpoint genes (PDCD1 (encoding PD1), CD274 (encoding PD-L1), CTLA4 (encoding CTLA4), LAG3 (encoding LAG3), and TIGIT (encoding TIGIT)) and an immune regulatory gene (IDO1), all genes that are currently under investigation for targeted immunotherapies; (2) major immune-activating genes commonly associated with antitumor immunity in colorectal carcinomas including IFNG (encoding IFNγ, the canonical T-helper 1 cells (Th1)-type cytokine) and TBX21 (encoding TBET, the canonical Th1 transcription factor); and (3) immune genes encoding proinflammatory cytokines and granzymes. These gene panels were extracted from current knowledge and selected references11, 12 and are outlined in Supplementary Table 2. For determination of mRNA up- or downregulation, given that RNA-seq control samples were limited, each sample’s gene expression values (e) were normalized by calculating the mean (μ) and variance (σ) of the expression values for samples which the gene was copy number diploid. Sample level gene expression (z-score) was then calculated as (e−μ)/σ. The recommended mRNA expression z-score of ±2 was used to define up- or downregulation.4, 9
Differences in the frequency of gene and chromosomal alterations among different histologic groups were compared within and across different molecular groups (MSI-H, non-MSI-H, and POLE-mutated) using Fisher’s exact test for categorical variables and the Kruskal–Wallis test for continuous variables. All tests were descriptive; no multiplicity adjustments were considered.
Results
The Spectrum of Tumor Morphological Patterns
Of the 224 tumors analyzed, 207 had adequate virtual images for a detailed morphological assessment. The remaining 17 were insufficient because of the superficial nature or limited amount of tumor in the available digital slides. The 207 tumors included 28 MSI-H tumors (14%), 172 non-MSI-H tumors (83%), and 7 POLE-mutated tumors (3%). The morphological characteristics of all 207 tumors are detailed in Supplementary Table 3 and summarized in Table 1. We observed that those tumors that showed mucinous features often had fairly conspicuous mucin production; even those with ‘<50% mucin’ (Supplementary Table 3) almost invariably had at least 20–30% mucin. As such, we grouped together all tumors that had either <50% mucin (n=18) or ≥50% mucin (n=12) in Supplementary Table 3 as one ‘mucinous subtype’ for all subsequent analyses.
As expected, the MSI-H tumors showed the characteristic ‘MSI-H histology’ including a higher frequency of the following patterns when compared with all non-MSI-H non-POLE-mutated tumors: medullary or medullary-like morphology (21% vs 0%), mucinous features (39% vs 10%), and increased tumor-infiltrating lymphocytes (score ≥2; 57% vs 5%) (P<0.001 for all patterns) (hereafter, these features are referred to as ‘MSI-H histology’).
The seven POLE-mutated tumors showed heterogeneous morphological patterns that were intermediate between MSI-H and non-MSI-H cancers. One tumor showed a ‘medullary-like’ morphological pattern with increased tumor-infiltrating lymphocytes, resembling MSI-H tumors. Four of seven (57%) cases showed lymphocyte scores ≥2, which is the same frequency as that seen in MSI-H cancers (57%), and significantly higher than that of non-MSI-H cancers (5%). Mucinous tumors occurred in two POLE-mutated cases (29%), a frequency lower than MSI-H tumors (39%) but higher than non-MSI-H tumors (10%).
Correlation Between Tumor Morphology and Molecular Alterations
Among all histologies, the not otherwise specified type had the lowest overall mutation count within all tumors and the lowest mononucleotide frameshift mutation count within MSI-H tumors.
As expected, the overall mutation count was highest in the POLE-mutated tumors (median, 3200) followed by MSI-H (median, 664) and non-MSI (median, 67) tumors. When tumors were stratified by histological patterns, the not otherwise specified type emerged as the pattern with the lowest mutational count both in the entire cohort (median, 68 vs ≥130 for other types, P<0.001) and in the MSI-H group (median, 382 vs ≥782 for other types; P<0.03) (Figure 2). Within the entire cohort, the medullary or medullary-like morphology had the highest count at a median of 968, followed by poorly differentiated and mucinous tumors (median, 357 and 130, respectively). Within MSI-H tumors, all histologies that were not the not otherwise specified type had similar mutation counts with their median counts ranging from 782 to 792. No difference in mutation count was detected between histological types among POLE-mutated tumors alone or among non-MSI-H/non-POLE cases alone. A slight tendency of poorly differentiated tumors having more mutations than the not-otherwise-specified type was noted in non-MSI-H/non-POLE cases (median, 78 vs 67, P=0.8).
Within the MSI-H tumors, analysis of mononucleotide frameshift mutations in the 20 genes6 that contain coding mononucleotide repeats of six nucleotides or longer showed that such mutations were lowest in the not otherwise specified type as well (median 2 vs 4–8 for other types, P<0.01). Poorly differentiated tumors with a solid growth tended to have the highest number of these mutations (Table 2).
We noted a trend in which tumors with higher numbers of tumor-infiltrating lymphocytes were associated with higher overall mutation counts within the MSI-H group; however, this did not reach statistical significance. No association was detected between the number of lymphocytes and mutation counts in the non-MSI-H group and POLE-mutated group. No association was detected between the lymphocyte count and frameshift mutation rate in any group of tumors.
‘MSI-H histology’ was associated with CIMP-H when compared with other Cancer Genome Atlas methylation subtypes within the entire cohort; the most common histology in CIMP-H/MSI-H tumors was mucinous, whereas the most common histology in CIMP-H/non-MSI-H tumors was the not otherwise specified type.
The Cancer Genome Atlas project classified the tumors into four methylation subtypes: CIMP-H, CIMP-L, Cluster 3, and Cluster 4 (see Materials and Methods section); this was based on the tumor promoter DNA methylation profiles.3 As expected, the MSI-H phenotype and the ‘MSI-H histology’ were significantly associated with CIMP-H subtype (P<0.001, Supplementary Table 4a). Within MSI-H tumors, the most common histology in CIMP-H tumors was mucinous (8/20, 40%). The vast majority of MSI-H tumors clustered with CIMP-H methylation subtype, skewing the distribution of the various histological features; there were insignificant P-values between these histological features and the different methylation subtypes within these MSI-H tumors (Supplementary Table 4b).
Among non-MSI-H tumors, the most common histology in CIMP-H tumors was the not otherwise specified type (7/11, 64%). A significant difference was detected between the histology subtypes and the subtypes of methylation, with mucinous histology being more frequently represented in CIMP-L subtype (P=0.02, Supplementary Table 4b). No difference was detected with regard to tumor glandular architectural pattern, tumor-infiltrating lymphocytes, neutrophils, and necrosis.
No association between morphological features and methylation subtypes was detected within POLE-mutated tumors.
Mucinous histology was associated with frequent alterations in BRAF, PIK3CA, and the TGF-β pathway genes among MSI-H and non-MSI-H cancers.
When assessing the key genes involved in the major pathways evaluated by The Cancer Genome Atlas project (see Materials and Methods section), alterations in BRAF or PIK3CA emerged as patterns more frequently seen with mucinous histology. The trend towards mucinous histology associating with BRAF and PIK3CA alterations was observed in both MSI-H and non-MSI-H subgroups, and it was statistically significant when all tumors were assessed together regardless of the MSI status (P<0.001 for BRAF; P<0.02 for PIK3CA). When assessing the non-MSI-H group, mucinous histology was significantly associated with BRAF mutations (P<0.02), whereas the association with PIK3CA did not reach statistical significance (P=0.13).
Among the five major signaling pathways, alterations in TGF-β pathway genes were particularly frequent in MSI-H tumors. The high frequency within the MSI-H group was seen across all histologic subtypes. Within non-MSI-H cancers, mucinous tumors were significantly more frequently altered in the TGF-β pathway compared with other histologic types (59% vs 21–25% in the other types, P<0.001). The two major genes contributing to this association were SMAD2 (18% in mucinous tumors vs 5% in non-mucinous tumors, P=0.06) and SMAD4 (47% in mucinous tumors vs 10% in non-mucinous tumors, P<0.001).
Figure 3 summarizes the frequencies of alterations in BRAF, PIK3CA, SMAD2, and SMAD4 in mucinous vs non-mucinous tumors throughout the entire cohort. Altogether, 86% of mucinous tumors had alterations in at least one of these four genes, which was significantly higher than the 26% observed in non-mucinous tumors (P<0.0001). The BRAF and PIK3CA mutations observed in all tumors were predominantly recurrent hotspot mutations. BRAF V600E accounted for 100% and 73% of the BRAF mutations in mucinous and non-mucinous tumors, respectively. Recurrent hotspot mutations in PIK3CA accounted for 73% and 82% of the PIK3CA mutations in mucinous and non-mucinous tumors, respectively.
In contrast, while there was no significant difference in the alterations in the WNT pathway overall, in the full cohort, mucinous tumors trended towards having a lower rate of APC mutations (63% vs 76% in non-mucinous tumors, P=0.18). Throughout the full cohort, mucinous tumor had a significantly reduced rate of TP53 mutation (37% vs 58% in non-mucinous tumors, P<0.05).
No significant associations were detected between tumor morphology and arm level chromosomal alterations.
Fifteen of the 187 tumors with pertinent data showed arm level chromosomal alterations. As expected, all 15 tumors were non-MSI-H. A trend was noted in which tumors of the not otherwise specified type with more complex architecture or solid growth were more likely to have chromosomal alterations; however, this did not reach statistical significance. No associations were detected among any of the other histological patterns.
Colorectal carcinomas exhibiting mRNA upregulation of major immune-inhibitory genes (including checkpoints) were primarily MSI-H, frequently associated with simultaneous upregulation of antitumor immune genes, and predominantly non-mucinous with a solid growth pattern.
Of the 207 tumors with complete histological data, 181 had gene expression data. Alterations of the immune-related genes in these tumors are listed in Supplementary Table 5. Alterations (primarily in the form of mRNA upregulation) of major immune-inhibitory genes (including checkpoint genes PDCD1, CD274, CTLA4, LAG3, and TIGIT, and regulatory gene IDO1) occurred in only 3–9% of the tumors overall, and the alterations mainly occurred in MSI-H tumors (7–43% in MSI-H group vs 0–3% in non-MSI-H group). The rate of upregulation of PDCD1 and CD274 was 17% and 43% in MSI-H tumors, respectively, but 0% and only 3% in non-MSI-H tumors, respectively.
As illustrated in Figure 4, co-upregulation of more than one immune-inhibitory gene in the same patients was a frequent phenomenon (9/14, 64%). Cases that had upregulation of immune checkpoint genes also tended to exhibit upregulation of genes that are commonly regarded as immune-activating (i.e., antitumoral) including Th1-type genes (as represented by IFNG and TBX21) and proinflammatory genes such as IL10, GZMM, CXCL13, and PRF1 (Supplementary Table 6).
Correlative analysis with tumor morphological patterns revealed that upregulation of immune-inhibitory genes (as well as genes associated with antitumor immunity) occurred primarily in tumors with non-mucinous histology. The frequencies of alterations in the genes studied between mucinous and non-mucinous tumors in MSI-H and non-MSI-H groups, respectively, are outlined in Supplementary Table 6. Differentially altered immune-related genes between mucinous and non-mucinous tumors are illustrated in Figure 4. It was noted that the majority of tumors (up to 70%) that exhibited upregulation of immune checkpoint genes were tumors with a medullary or solid growth pattern.
All MSI-H tumors that had upregulation of the immune checkpoint genes had easily discernible tumor-infiltrating lymphocytes (score of ≥1). However, there was no statistically significant difference in the lymphocyte count between tumors with and without upregulation of the immune checkpoint genes. Furthermore, in the non-MSI-H group, upregulation of immune checkpoint genes occurred in both cases with and without apparent tumor-infiltrating lymphocytes.
Discussion
In this analysis, we observed several interesting associations between tumor morphology and genetic alterations in the cohort of primary colorectal carcinomas studied by The Cancer Genome Atlas project.3 First, colorectal cancers with conventional gland-forming histology (ie, the not otherwise specified type) exhibited a significantly lower mutation count than tumors with other histologies. Second, while The Cancer Genome Atlas CIMP subtypes correlated tightly with MSI-H and ‘MSI-H histology’, morphological differences were noted in CIMP-H tumors with or without MSI-H. Third, mucinous histology was associated with a specific molecular signature characterized by a higher rate of mutation or copy number alteration in BRAF, PIK3CA, SMAD2, and SMAD4, and a trend towards a lower rate of APC or TP53 mutations. Fourth and importantly, our study showed that only a small subset of colorectal carcinomas exhibited mRNA upregulation of immune checkpoint genes—targets of current checkpoint blockade therapies; these tumors were primarily MSI-H and had distinct morphological features: non-mucinous with a solid growth pattern including both medullary and non-medullary patterns.
Our study is the first to illustrate a direct association between the number of somatic mutations and the morphological patterns in tumors. Accumulating evidence has led to a firm belief that cancer is a genetic and epigenetic disease, and somatic mutations in specific genes are responsible for tumor initiation and progression. In colorectal carcinoma, it has been suggested that the development of cancer requires just three sequential mutations in ‘driver genes’.13 Within an established cancer, however, many more ‘passenger mutations’ exist. It remains to be elucidated whether and to what extent these additional seemingly ‘non-driver’ mutations contribute to the progression of the tumor.
In this context, it is intriguing to note that the mutation counts observed in colorectal carcinomas by The Cancer Genome Atlas project was significantly lower in tumors of conventional or not otherwise specified type when compared with tumors of other histologies. This was true for overall mutation counts in both the entire cohort and the subset of MSI-H tumors. This was also true with regard to mononucleotide frameshift mutations within MSI-H tumors. Several questions may be raised based on this observation. First, as the not otherwise specified type is the ‘simplest’ form of all histologies and closest in appearance to normal colonic mucosa, are the more ‘complexed’ or ‘specialized’ forms of the other histologies a result of the excess mutation burden? In MSI-H cancers, given that the degree of MSI—the number of mutations in the microsatellite repeats—is believed to be a ‘molecular clock’ of how old the tumor is,14, 15 are the not otherwise specified MSI-H cancers ‘younger’ tumors? In addition, frameshift mutations at mononucleotide repeats have been shown to yield characteristic neoantigenic peptides, and evidence exists that these peptides elicit immune responses;16, 17 are tumors of the not otherwise specified type (a type with few frameshift mutations) less immunogenic than tumors with a solid growth that tend to have more frameshift mutations? Answers to these questions await further investigation.
CIMP-H colorectal cancers have been shown to share pathologic characteristics with sporadic MSI-H tumors,18, 19 both seemingly associated with the serrated pathway and tending to have mucinous components. This is not surprising given that nearly all sporadic MSI-H tumors are molecularly based on the promoter CpG island hypermethylation-induced silencing of the MLH1 gene. In this study, while a tight association of MSI-H and CIMP-H was indeed well demonstrated, we also showed that the CIMP-H tumors that were with or without MSI-H tended to have different histological patterns: the most common histology in CIMP-H/MSI-H group was mucinous, while that in CIMP-H/non-MSI-H was the not otherwise specified type. Such findings thus broaden our view on these tumors, and suggest that these two molecular classes may actually evolve through different tumorigenesis pathways: the former may evolve through the serrated pathway (in keeping with the current belief), and the latter may potentially evolve through conventional adenomas or traditional adenomas (different from current belief). Attention should therefore be directed to such differences in our future research efforts.
In our analysis of major tumorigenic pathways, we confirmed the association between the well-known ‘MSI-H histology’ and the MSI pathway. In addition, we observed a molecular pattern that closely correlated with mucinous histology. This pattern consisted of mutations and copy number alterations in BRAF (the MAPK/ERK pathway), PIK3CA (the PI3K pathway), and SMAD2 and SMAD4 (the TGF-β pathway).
Only limited studies have evaluated the association between histological patterns and gene mutations in colorectal carcinoma, and the reported findings are largely consistent with our observations. Yaeger et al20 studied the characteristics of BRAF-mutant metastases and found that mucinous type or mucinous features were significantly more frequently encountered in BRAF-mutated tumors compared with the BRAF-wild-type counterpart (43% vs 16%, P<0.01). In evaluating mutation frequencies of 12 genes in colorectal carcinoma, Chang et al21 also found a higher frequency of mutations in BRAF (8% vs 4%), PIK3CA (25% vs 13%), and SMAD4 (8% and 4%) in their mucinous carcinomas when compared with non-mucinous tumors. Additionally, Chang et al21 also found higher mutation rates in KRAS and AKT1 in mucinous carcinomas. Furthermore, animal studies have shown that mice expressing constitutively active PI3K in the intestinal epithelium developed colonic adenocarcinomas with mucinous histology.22
Taken together, the data in the literature and our study indicate that alterations in BRAF, PIK3CA, SMAD2, and SMAD4 are more frequent in colorectal carcinomas with mucinous histology. This, in conjunction with our observation of lower rates of APC and TP53 alterations in mucinous tumors, suggests that there may exist divergent molecular pathways that underlie mucinous and non-mucinous tumorigenesis. The MAPK, PI3K, and TGF-β pathways may have a more important role in the formation of mucinous tumors, whereas WNT and p53 may be more important in non-mucinous carcinomas. Clinically, the genetic alterations associated with mucinous histology bear practical relevance: BRAF V600E is associated with a poor prognosis23 and may also predict cetuximab resistance;24 PIK3CA mutations appear to be predictive of a benefit from aspirin;25, 26 SMAD4 mutations have been shown to be an adverse prognostic marker.27 As such, the mucinous histology may serve as a readily identifiable feature that can facilitate the efficient recognition of these molecularly important tumor subsets.
Colorectal carcinoma can be an immunogenic tumor.28 It has been suggested that this tumor immunogenicity results in an antitumor immune response in the host (as represented by upregulated Th1 and cytotoxic T cell genes); simultaneously, as a defense to counter this antitumor immunity, the tumor cells and host cells also amount an immune-inhibitory response (as represented by upregulated immune checkpoints such as PDCD1, CD274, CTLA4, LAG3, and TIGIT and immune regulatory genes such as IDO1).11 It is this immune-inhibitory function of the immune checkpoints and regulatory genes that has formed the basis for the current immunotherapy trials.29 Interestingly, clinical results thus far indicate that only a small subset of colorectal cancer patients responds to immune checkpoint inhibition, and almost all responders had MSI-H tumors; a recent phase II trial30 reported immune-related objective response to treatment with the anti-PD-1 agent, pembrolizumab, in 14% of 28 colorectal cancer patients enrolled, including 40% mismatch repair-deficient cases and 0% mismatch repair-proficient ones.
In line with these observations, our assessment of mRNA expression of the immune-inhibitory genes showed that upregulation of immune-inhibitory genes was limited to only a small subset of colorectal cancer, and most of them were MSI-H. Upregulation of CD274 (the gene encoding PD-L1), for example, occurred in only 8% of the tumors and this included 43% of MSI-H cancers but only <3% of the cancers that were non-MSI-H. Furthermore, our analysis revealed frequent co-upregulation of more than one checkpoint within the same tumors, a finding that may have implications for the administration of combinatorial immunotherapies. Not surprisingly, upregulation of immune-inhibitory genes also coexisted with upregulation of genes associated with antitumor immunity in colorectal cancer, including IFNγ and TBX21.
An interesting and potentially significant finding that emerged from our analysis is that tumors with upregulated immune checkpoints tend to have specific morphological patterns. Few studies are available that have addressed such associations. A very recent publication suggested that ‘medullary’ colorectal cancers are more likely to have an immunoregulatory microenvironment.31 In our analysis, we further demonstrated that tumors with upregulated immune checkpoints were frequently non-mucinous, had a medullary or solid growth pattern, and often contained tumor-infiltrating lymphocytes (although the number of lymphocytes may not always be high, and in MSS cases, such lymphocytes may be inconspicuous).
These findings are likely to be relevant. The rate of CD274 upregulation in MSI-H colorectal cancers (43%) is strikingly similar to the reported anti-PD-1 therapy response rate (40% of MSI-H cases).30 One might hypothesize that it is the tumors that have upregulated immune checkpoints that are more likely to respond to anti-checkpoint therapy; consequently, the specific morphological patterns associated with checkpoint upregulation may have a role in patient selection for such therapy. However, it is to be emphasized that there are no data yet to connect directly immune checkpoint upregulation with clinical response to anti-checkpoint immunotherapy; inclusion of tumor morphology as well as tumor testing for immune checkpoint upregulation in clinical trial design would be desirable.
A limitation of this study is that the morphological assessment was based only on images of portions of the tumor, not the entire case. As such, features like tumor heterogeneity, tumor budding at the invasive front, and Crohn’s-type reaction could not be accurately evaluated. Nonetheless, the images represented the samples that were sequenced by The Cancer Genome Atlas project, and the essential morphological characteristics were well captured, as reflected by the almost exclusive representation of MSI-H histology within the subset of MSI-H cancers. It must also be noted that in The Cancer Genome Atlas database, clinical follow-up data were not available to examine for outcome or treatment response correlations.
In summary, our analysis revealed specific morphological patterns that are linked to clinically relevant genetic alterations. Recognizing these associations may facilitate the recognition of clinically significant tumor subsets and ultimately allow more precise patient care. As illustrated in Figure 5, mucinous histology, occurring in a minor but significant subset of colorectal cancers, is linked to relevant genetic traits such as alterations in BRAF, PIK3CA, and SMAD genes, all of which have been shown to have prognostic and/or predictive significance. Similarly, the observation that the most common histology type differs between the CIMP-H tumors that have or do not have MSI-H suggests that these two molecular classes (CIMP-H/MSI-H and CIMP-H/non-MSI-H) may evolve through different precursor lesions. Of all the histologies, the not otherwise specified type is characterized by a relatively low mutation count, and in MSI-H cancers, a lower number of mononucleotide frameshift mutations. The latter may result in a lower generation of frameshift peptides or neoantigens and thus a different degree of tumor immunogenicity. Importantly, tumors with medullary features or a solid growth pattern, with or without significantly increased tumor-infiltrating lymphocytes, appear more likely to exhibit upregulated immune checkpoints, a phenomenon that evokes the hypothesis that these tumors carry a greater potential to respond to immune checkpoint inhibitors. To this effect, efforts are ongoing at our institution to correlate tumor histopathology (along with the tumor’s MSI status and genomic alteration patterns) with treatment responses to the various immune checkpoint blockade regimens in patients with colorectal carcinoma. We anticipate that these efforts will expand the knowledge we have gained from The Cancer Genome Atlas project and facilitate its translation into practical utility in the clinical arena.
References
Fearon ER, Vogelstein B . A genetic model for colorectal tumorigenesis. Cell 1990; 61: 759–767.
Gao J, Ciriello G, Sander C,et al. Collection, integration and analysis of cancer genomic profiles: from data to insight. Curr Opin Genet Dev 2014; 24: 92–98.
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012; 487: 330–337.
Gao J, Aksoy BA, Dogrusoz U et al, Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013; 6: pl1.
Lin EI, Tseng LH, Gocke CD et al, Mutational profiling of colorectal cancers with microsatellite instability. Oncotarget 2015; 6: 42334–42344.
Donehower LA, Creighton CJ, Schultz N et al, MLH1-silenced and non-silenced subgroups of hypermutated colorectal carcinomas have distinct mutational landscapes. J Pathol 2013; 229: 99–110.
Boland CR, Goel A . Microsatellite instability in colorectal cancer. Gastroenterology 2010; 138: 2073–87 e3.
Shia J . Evolving approach and clinical significance of detecting DNA mismatch repair deficiency in colorectal carcinoma. Semin Diagn Pathol 2015; 32: 352–361.
Cerami E, Gao J, Dogrusoz U et al, The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2012; 2: 401–404.
Umar A, Boland CR, Terdiman JP et al, Revised Bethesda Guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst 2004; 96: 261–268.
Llosa NJ, Cruise M, Tam A et al, The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov 2015; 5: 43–51.
Maby P, Tougeron D, Hamieh M et al, Correlation between density of CD8+ T-cell infiltrate in microsatellite unstable colorectal cancers and frameshift mutations: a rationale for personalized immunotherapy. Cancer Res 2015; 75: 3446–3455.
Tomasetti C, Marchionni L, Nowak MA,et al. Only three driver gene mutations are required for the development of lung and colorectal cancers. Proc Natl Acad Sci USA 2015; 112: 118–123.
Blake C, Tsao JL, Wu A,et al. Stepwise deletions of polyA sequences in mismatch repair-deficient colorectal cancers. Am J Pathol 2001; 158: 1867–1870.
Calabrese P, Tsao JL, Yatabe Y et al, Colorectal pretumor progression before and after loss of DNA mismatch repair. Am J Pathol 2004; 164: 1447–1453.
Kloor M, Becker C, Benner A et al, Immunoselective pressure and human leukocyte antigen class I antigen machinery defects in microsatellite unstable colorectal cancers. Cancer Res 2005; 65: 6418–6424.
Schwitalle Y, Linnebacher M, Ripberger E,et al. Immunogenic peptides generated by frameshift mutations in DNA mismatch repair-deficient cancer cells. Cancer Immun 2004; 4: 14.
Bae JM, Kim JH, Kang GH . Epigenetic alterations in colorectal cancer: the CpG island methylator phenotype. Histol Histopathol 2013; 28: 585–595.
Kim JH, Kang GH . Molecular and prognostic heterogeneity of microsatellite-unstable colorectal cancer. World J Gastroenterol 2014; 20: 4230–4243.
Yaeger R, Cercek A, Chou JF et al, BRAF mutation predicts for poor outcomes after metastasectomy in patients with metastatic colorectal cancer. Cancer 2014; 120: 2316–2324.
Chang SC, Lin PC, Lin JK et al, Mutation spectra of common cancer-associated genes in different phenotypes of colorectal carcinoma without distant metastasis. Ann Surg Oncol 2015; 23: 849–855.
Leystra AA, Deming DA, Zahm CD et al, Mice expressing activated PI3K rapidly develop advanced colon cancer. Cancer Res 2012; 72: 2931–2936.
Tol J, Nagtegaal ID, Punt CJ . BRAF mutation in metastatic colorectal cancer. N Engl J Med 2009; 361: 98–99.
Di Nicolantonio F, Martini M, Molinari F et al, Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol 2008; 26: 5705–5712.
Domingo E, Church DN, Sieber O et al, Evaluation of PIK3CA mutation as a predictor of benefit from nonsteroidal anti-inflammatory drug therapy in colorectal cancer. J Clin Oncol 2013; 31: 4297–4305.
Liao X, Lochhead P, Nishihara R et al, Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N Engl J Med 2012; 367: 1596–1606.
Kozak MM, von Eyben R, Pai J et al, Smad4 inactivation predicts for worse prognosis and response to fluorouracil-based treatment in colorectal cancer. J Clin Pathol 2015; 68: 341–345.
Galon J, Costes A, Sanchez-Cabo F et al, Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 2006; 313: 1960–1964.
Wolchok JD . PD-1 Blockers. Cell 2015; 162: 937.
Le DT, Uram JN, Wang H et al, PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 2015; 372: 2509–2520.
Friedman K, Brodsky AS, Lu S et al, Medullary carcinoma of the colon: a distinct morphology reveals a distinctive immunoregulatory microenvironment. Mod Pathol 2016; 29: 528–541.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Shia, J., Schultz, N., Kuk, D. et al. Morphological characterization of colorectal cancers in The Cancer Genome Atlas reveals distinct morphology–molecular associations: clinical and biological implications. Mod Pathol 30, 599–609 (2017). https://doi.org/10.1038/modpathol.2016.198
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.198
This article is cited by
-
Novel biomarker SARIFA in colorectal cancer: highly prognostic, not genetically driven and histologic indicator of a distinct tumor biology
Cancer Gene Therapy (2024)
-
Deep learning model to predict Epstein–Barr virus associated gastric cancer in histology
Scientific Reports (2022)
-
Clinicopathological and molecular characteristics of RSPO fusion-positive colorectal cancer
British Journal of Cancer (2022)
-
New thiophene derivative augments the antitumor activity of γ-irradiation against colorectal cancer in mice via anti-inflammatory and pro-apoptotic pathways
Discover Oncology (2022)
-
Swarm learning for decentralized artificial intelligence in cancer histopathology
Nature Medicine (2022)