Abstract
MicroRNAs (miRNAs) are a class of small (∼22 nucleotides), non-coding, highly conserved single-stranded RNAs with posttranscriptional regulatory features, including the regulation of cell proliferation, differentiation, survival, and apoptosis. They are deregulated in a broad variety of tumors showing characteristic expression patterns and can, thus, be used as a diagnostic tool. In contrast to non-small cell carcinoma of the lung neuroendocrine lung tumors, encompassing typical and atypical carcinoids, small cell lung cancer and large cell neuroendocrine lung cancer, no data about deregulation of tumor entity-specific miRNAs are available to date. miRNA expression differences might give useful information about the biological characteristics of these tumors, as well as serve as helpful markers.
In 12 pulmonary neuroendocrine tumors classified as either typical carcinoid, atypical, large cell neuroendocrine or small cell lung cancer, screening for 763 miRNAs known to be involved in pulmonary cancerogenesis was conducted by performing 384-well TaqMan low-density array real-time qPCR. In the entire cohort, 44 miRNAs were identified, which showed a significantly different miRNA expression. For 12 miRNAs, the difference was highly significant (P<0.01). Eight miRNAs showed a negative (miR-22, miR-29a, miR-29b, miR-29c, miR-367*; miR-504, miR-513C, miR-1200) and four miRNAs a positive (miR-18a, miR-15b*, miR-335*, miR-1201) correlation to the grade of tumor biology. The miRNAs let-7d; miR-19; miR-576-5p; miR-340*; miR-1286 are significantly associated with survival. Members of the miR-29 family seem to be extremely important in this group of tumors. We found a number of miRNAs, which showed a highly significant deregulation in pulmonary neuroendocrine tumors. Moreover, some of these deregulated miRNAs seem to allow discrimination of the various subtypes of pulmonary neuroendocrine tumors. Thus, the analysis of specific sets of miRNAs can be proposed as diagnostic and/or predictive markers in this group of neoplasias.
Similar content being viewed by others
Main
Lung cancer is the leading cause of cancer death worldwide.1, 2 Pulmonary neuroendocrine tumors form a distinct group of neoplasms that share characteristic morphological, immunohistochmical, ultrastructural and molecular features.3 These tumors considerably differ in their biological behaviors, thus, they are classified as either low-grade typical carcinoids, intermediate-grade atypical carcinoids, high-grade large cell neuroendocrine lung cancer, or small cell lung cancer.3, 4 Small cell lung cancers account for the largest group of these tumors (∼15% of all lung malignancies) and are associated with an extremely poor prognosis,5, 6, 7, 8 whereas large cell euroendocrine cancers and carcinoids are rare. Currently, the 2004 World Health Organization classification of pulmonary neuroendocrine tumors is based on combined architectural patterns considering the two most relevant morphological parameters, that is, the mitotic index and the presence of necrosis, diagnosed by haematoxylin and eosin morphology.3, 8, 9 However, these tumors represent a broad spectrum of phenotypically distinct entities, and the assignment of a given neuroendocrine lung tumor can occasionally be cumbersome even for experienced pathologists.3, 9, 10 Thus, reproducible and objective (molecular) diagnostic criteria with clinical and prognostic value need to be established to achieve a more accurate assignment of the various types of pulmonary neuroendocrine tumors.3
Micro-RNA (miRNA) molecules are evolutionarily conserved, small RNA-molecules with a size of 19–24 nucleotides and, unlike mRNA, do not encode amino-acid sequences.3, 11, 12 miRNAs function as negative endogenous gene-expression regulators by binding complementary sequences in target mRNAs, resulting in their selective degradation or selective steric inhibition of translation.2, 3, 12, 13 Therefore, miRNAs are involved in a wide range of biological functions, including cellular proliferation, differentiation, angiogenesis, apoptosis, and metastatic potential.2, 12, 13, 14, 15, 16, 17 Aberrant miRNA expression has been reported to be involved in the pathogenesis of Alzheimer’s disease,18 cardiovascular disease,19 spinal motor neuron anomalies,20 and numerous others.2 Furthermore, deregulated expression of miRNAs has been identified in a variety of human malignancies,12, 13, 21, 22, 23, 24, 25, 26, 27, 28, 29 suggesting that miRNAs function as potential oncogenic factors or tumor suppressors, depending on the cell type or tissue investigated and their target genes.3 For example, the miR-34 family members are direct transcriptional targets of p53 and their expression induces cell cycle arrest in cancer cell lines.5, 30, 31 miRNA-29 family members act as tumor suppressors through the restoration of a normal DNA methylation pattern.5, 31 Among other tumor entities, both of these miRNAs are known to be deregulated in non-small cell lung cancer.31, 32, 33, 34, 35, 36 The role of miRNAs in cancer biology, prognosis prediction, and as a diagnostic tool in non-small cell lung cancer without neuroendocrine features has been intensively studied.3, 5, 37 Besides miR-34 and miR-29 family members, let-7a overexpression, possibly through suppression of the RAS oncogene, was shown to be related to increased overall survival in non-small cell lung cancer patients.2, 5, 38, 39, 40 In contrast, overexpression of the ‘oncomirs’ miR-21 and miR-155 was shown to be related to a decreased overall survival in non-small cell lung cancer patients.2, 5, 39, 41 It has been suggested that miR-31 may act as an oncogenic miRNA by inhibiting the tumor suppressors LATS2 and PPP2R2A,42 and the expression of miR-205 has been suggested to distinguish squamous from non-squamous lung cancer.2, 43 miR-126 may promote lung cancer cells irradiation-induced apoptosis through the PI3K-AKT pathway.2, 44 However, data on the role of miRNA expression in neuroendocrine tumors of the lung are limited, and the function of several miRNAs, which are well-documented cancer-related miRNAs in other cancer types, has not been investigated in neuroendocrine lung tumors so far.3, 5 Furthermore, we hypothesized that specific ‘biomarker miRNAs’ might exist that could distinguish more accurately and reliably between subtypes of pulmonary neuroendocrine tumors. Against this background, 763 different miRNAs in all four different subtypes of neuroendocrine human pulmonary tumors were evaluated.
Materials and methods
Patient Cohort and Study Design
Twelve tumor specimens including three of each typical and atypical carcinoid tumors, large cell neuroendocrine lung cancer, and small cell lung cancer were investigated. The specimens were retrieved from the archives of the Institute of Pathology and Neuropathology of the University Hospital Essen (Germany). All of these tumor specimens were classified as either atypical or typical carcinoids of the lung, large cell lung cancer or small cell lung cancer, respectively. To verify the initial diagnosis, all specimens were reevaluated by two experienced pathologists (JWO, THA). The patients had undergone surgical resection for their lung tumors between 2005 and 2012. In addition, clinical follow-up was available, as well as information about a broad range of other investigations (immunohistochemistry, sequencing-data, and mRNA expression profiles). All the investigated samples contained a sufficient amount of tumor cells for RNA isolation and a low percentage of contaminating benign or desmoplastic stroma cells. The investigations conform to the principles declared in the declaration of Helsinki. This retrospective study was approved by the Ethics Committee of the Medical Faculty of the University Duisburg-Essen (13-5382-BO).
TaqMan qPCR miRNA Assay Cards
Formalin-fixed, paraffin-embedded tissue specimens were used for RNA isolation. Sections of tumor tissue prepared for RNA isolation were stored at −20 °C until used. Total RNA isolation (including isolation of miRNA) was performed with the ‘miRNeasy FFPE’ kit from Qiagen (Hilden, Germany) according to the manufacturers’ protocol, using adapted times and volumes (eg, overnight digestion). RNA not immediately used was stored at –20 °C until use.
For cDNA synthesis the Megaplex RT Human Primer Pools A and B (Applied Biosystems, Foster City, CA, USA) in combination with the TaqMan miRNA Reverse Transcription Kit (Applied Biosystems) was engaged. cDNA not immediately used was stored at −20 °C. Preamplification was also conducted with Megaplex Preamplification Primer Pools A and B (Applied Biosystems) in combination with the TaqMan miRNA Preamplification Kit (Applied Biosystems). Preamplified samples were stored at −20 °C until used for expression analysis. For 384-well TaqMan low-density array real-time qPCR, preamplification products were pressed through microchannels into wells fixed in the card, preloaded with immobilized, dehydrated target-specific primers-probe pairs. PCR analysis was run on the ABI PRISM 7900 System. Analysis was done with TaqMan Array Human MicroRNA Card A and TaqMan Array Human MicroRNA Card B v3.0 (both Applied Biosystems), respectively. PCR was performed using the ready-to-use TaqMan Universal PCR Master Mix (Applied Biosystems). Of note, qPCR analysis was performed in concordance to the MIQE-guidelines.45
Statistical Analysis
Correlation analysis was performed with a custom-programmed algorithm for the R software. The exact Wilcoxon–Mann–Whitney Rank Sum test was used to test associations between miRNA expression and dichotomous variables (eg, gender). To rule out a possible association between miRNA and clinical variables (age, age of blocks, etc), a Spearman’s rank correlation test was performed. The Spearman’s rank correlation test was done to test associations between tumor type and expression levels of different miRNAs.
Overall survival was compared by performing Kaplan–Meier curves. Survival analysis for overall survival was calculated by Cox-regression (COXPH-model), statistical significance was determined using Likelihood ratio test and Score (logrank) test.
Because of the better distribution of our expression levels in logarithmic scale, all further analysis were additionally performed using log(miR)-values. A Bonferroni correction for multiple testing was performed to all calculated P-values to overcome the problem of false-positive results in this high-throughput statistics. The level of statistical significance was defined as P<0.05.
Bioinformatics
miRNA target-sites were descriptively determined using the miRanda-database (miRanda), inhibition scores were calculated by the mirSVR algorithm. The 10 most strongly regulated miRNA-targets of every miRNA being differently regulated in the various neuroendocrine tumors were selected for further analysis. In addition, all targets with a score over 2.00 were selected for the identification of possible deregulated pathways. Associated pathways, which were affected by differences in the miRNA expression levels, were discovered and, if possible, correlated to gene- and protein expression levels already available. In addition, a gene ontology (GO)-term analysis to classify differentially regulated biological processes was performed.
Databases used for pathway identification are uniprot.org and the NCBI website gene-search and BioSystems-search, respectively.
Results
qPCR analysis was accomplished successfully in all cases. The endogenous control always showed up between Ct 12.5 and 13.9 on both plates.
In the entire cohort of patients including typical and atypical carcinoids, large cell neuroendocrine cancers and small cell lung cancers, 44 significantly differently expressed miRNAs could be identified. For 11 of these differently expressed miRNAs, the level of statistical significance was <0.01. Seven of these miRNAs (miR-22, miR-29a, miR-29b, miR-29c, miR-367*; miR-504, miR-513C, miR-1200) were negatively correlated to biological aggressiveness (determined by tumor grade), for the remaining four miRNAs a positive correlation with tumor biology (miR-18a; miR-15b*, miR-335*, miR-1201) was observed (data are summarized in Table 1).
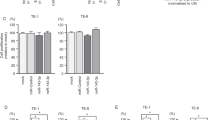
miR-18a was significantly correlated with the tumor grading, that is, being lowest in typical and increasing in atypical carcinoids to large cell neuroendocrine lung cancer, and being highest in small cell lung cancer, showing a 20.25-fold higher expression in high-grade neuroendocrine pulmonary tumors compared with carcinoid tumors. Similar observations were made for miR-15b* and miR-335*, 5.25-fold and 4.62-fold, respectively). miR-22 expression decreased >12 times from typical to atypical carcinoids and large cell carcinoma (both showed identical expression levels) and again >12 times to small cell lung cancer (overall decrease from carcinoids to carcinomas: 23.9-fold). Moreover, miR-29a, miR-29b, and miR-29c expression levels decline with increasing aggressiveness of the tumor. They demonstrated almost identical expression levels in carcinoids and the expression levels significantly decreased in large cell carcinoma and further to small cell lung cancer. miR-504 expression was rather stable in carcinoids and large cell lung cancer but was completely abolished in small cell cancer. miR-367* showed a similar pattern of expression compared with miR-22, but the inverse correlation with aggressiveness was much stronger (typical carcinoid/typical carcinoid: ∼21.85-fold; large cell cancer/small cell cancer: ∼28.22-fold). Expression of miR-513C was only detected in typical carcinoids, but was significantly decreased to a minimum in atypical carcinoids and high-grade neuroendocrine lung cancers. miR-1201 was stably expressed in both carcinoids, but its expression increased in high-grade neuroendocrine lung cancers (Figure 1).
(a) miRNA-19 family members are significantly different expressed in the four analyzed entities if pulmonary NELCs. miRNA-19 family members mainly regulate DNMTs and, therefore, gene silencing mechanisms. Also TP53 expression is strongly regulated by miRNA-19 family members. (b) Overview of different highly significant (P<0.001) expressed miRNAs between the different types of pulmonary neuroendocrine tumors. miR-513C has a strong diagnostic impact to discriminate between typical and atypical carcinoids; miR-504 can be a supporting diagnostic tool for discrimination of large cell neuroendocrine lung cancer and small cell lung cancer. miR-18a and miR-1201 are much highly expressed in carcinomas than in carcinoids, also miR-15b* and miR-335* show higher expression in HG than in LG tumors. In contrast, miR-22 and miR-367* are less expressed in HG than in LG NET of the lung.
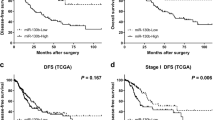
Forty-five miRNAs were identified showing a significant impact on survival time. Highly significant associations were noted for nine miRNAs using the Likelihood ratio test and the Score (logrank) test (let-7d; miR-139-5p; miR-197; miR-301; miR-576-5p; miR-582-5p; miR-448; miR-340*; miR-1286). Five miRNAs were still significantly associated with survival if an additional Wald test was performed (let-7d; miR-19; miR-576-5p; miR-340*; miR-1286). Data are summarized in Table 2.
In this cohort it was observed that the increasing malignant potential of the tumors was correlated with the differential expression of miRNAs leading to pathway upregulation. The pathway affected the most was the extracellular signal-regulated kinase (ERK)/MAPK-signaling pathway. Also in the phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K) pathway, which is involved in different cellular processes, significant differences in miRNAs expression between the four tumor types of pulmonary neuroendocrine tumors were noted. Furthermore, the TGF-beta pathway becomes activated with increasing potential for the development of metastasis, indicating a causative relation between augmented angiogenesis and more frequent development of metastatic disease in these tumors. In addition, the Wnt-signaling pathway, NFkB pathway, JNK-MAPK-signaling network as well as the mTOR-complex associated signaling was differentially regulated by miRNA expression (Figure 2).
The GO analysis revealed 256 hits at regulation of gene expression and transcriptional regulation. The next major differentially regulated field of cellular processes was cell cycle regulation (139 hits) followed by the development and differentiation of the nervous system, more precisely the part of neuronal development (134 hits). In addition, proliferation (125 hits), immune response (105 hits), apoptosis (96 hits), and DNA repair (59) were strongly deregulated in this collection of neuroendocrine lung cancer specimens. Furthermore, growth factors as well as growth-factor receptors and their pathways were clearly affected (53 hits). Less, but also differentially regulated were processes such as ion uptake/carrier-depended uptake of compounds and angiogenesis (48 and 42 hits, respectively). Moreover, there was a strongly aberrant regulation of the expression of structural cell compounds (42 hits), especially of collagens (19 hits) (Figure 3).
Discussion
Despite the accuracy of diagnostic methods used in modern pathology has dramatically improved, the discrimination between the different subtypes of neuroendocrine lung tumors remains challenging. Especially the differential diagnosis between atypical carcinoids and small cell lung cancers as well as the differentiation between small cell lung cancer and large cell neuroendocrine lung cancer can need supporting molecular-biological facts. To improve this discrimination, the establishment of reliable markers for the differentiation between different subtypes of neuroendocrine pulmonary tumors in routine pathology is desirable. This study was designed to test the miRNA profiles of these lung cancer types in order to establish feasible and reliable markers that facilitate finding correct diagnosis in routine diagnostic surgical pathology. At present, the diagnosis of the different types of neuroendocrine lung tumors is usually based on morphological and histological findings. The identification of a clear-cut correlation between the expression of a defined set of miRNAs and a given neuroendocrine tumor type would be of benefit for both the diagnostic process and individualization of therapeutic concepts. Also when isolated from formalin-fixed, paraffin-embedded tissue, RNA can be a useful and robust marker rendering reproducible results.46 In addition, miRNAs are highly stable also in formalin-fixed tissue and, therefore, can be used for normal pathological routine diagnostic purposes.47 In the past, some miRNAs were found to distinguish between different types of lung tumors, having a crucial role in the tumorgenesis of lung cancers and/or having a prognostic or predictive impact on these tumors. However, the majority of these studies focused on non-small cell lung cancer patients, whereas data concerning pulmonary neuroendocrine tumors are still lacking.
Expression of the miR-29 family was inversely correlated to an increasing degree of malignant behavior in our cohort of patients. Decreased expression was associated with an increasing aggressive biological behavior. In large cell neuroendocrine lung cancers, the expression of miR-29 family members is known to be strongly correlated with the DNA methylation pattern of tumor cells, because the miR-29 family members target the DNA-methyltransferases 3A and 3B (DNMT3A/DNMT3B). Their function is required for the de novo methylation and for the establishment of DNA methylation patterns during development. Therefore, a high methylation pattern is present, but an induced expression of miR-29 restores normal pattern and reexpression of silenced tumor suppressor genes like fragile histidine triad in cell lines.31 Finally, this leads to a reprogramming of tumor cells to the physiological state, which indicates that, the miR-29 family (especially miR-29a, miR-29b, and miR-29c) act as tumor-suppressor-miRNAs.31 Furthermore, this mechanism might explain why downregulation of miR-29 expression is accountable for hypermethylation and frequent silencing of tumor suppressor genes. Although the expression levels in carcinoids were almost identical, there was a huge difference between low and high-grade neuroendocrine lung tumors as well as between large cell neuroendocrine lung cancer and small cell lung cancer (Figure 2). Furthermore, other targets of the miR-29 family are p85α and CDC42.48 CDC42 and p85α are negative regulators of p53, which has apoptotic activity. Lower miR-29 expression decreases p53 levels resulting in escape from apoptosis and is associated with biologically malignant behavior. miR-34 family members are suspected to be associated with the p53 tumor suppressor network to inhibit inappropriate cell proliferation.30 Nevertheless, no significant difference in the miR-34 expression levels between the four types of neuroendocrine tumors could be outlined in this study. Our findings are consistent with a report from Lee et al5 demonstrating that none of the miR-34 family members could be used as a prognostic marker in neuroendocrine lung cancers, including small cell lung cancer. Members of the miRNA family let-7 are also thought to be deregulated in human lung cancers, including cancers with neuroendocrine differentiation.3, 12 However, in this study, these miRNAs were not able to discriminate between the different subtypes of neuroendocrine lung tumors. Thus, we were unable to confirm the association observed in other retrospective studies.2, 5, 38, 39, 40 The let-7d expression was significantly associated with overall survival, suggesting that let-7d could have an important role in metastasis and tumor progression. The miRNAs miR-21 and miR-155, both reported to be overexpressed in neuroendocrine lung cancers,1, 3 did not show a significant association with overall survival in our cohort. Similar results were revealed for miR-210, a miRNA overexpressed in cancer tissue vs normal lung parenchyma.1 Especially miR-21 turns out to be an interesting target, because it is a key regulator of the mTOR pathway and is reported to be differentially regulated in neuroendocrine tumors.49 In addition, miR-96 expression co-induces the methylation pattern of normal cells and is upregulated in non-small cell lung cancer.2 Nevertheless, we were unable to confirm that miR-96 is a potential diagnostic marker whose expression differs between the neuroendocrine tumor types. In this context, the differential expression of the miR-143 in the different tumor types was reported. miR-143 targets mainly the hexokinase 2 gene, which is the key regulator of glycolysis activated in multiple human cancers.50 If the malignant potential increases, the expression of miR-143 decreases leading to an augmented glycolytic activity of the tumors (Warburg effect). High-grade neuroendocrine cancers with an increased proliferative activity require a stronger metabolic activity and depend on the glycolytic utilization of glucose.
However, the expression of miR-375 did not differ significantly between the various tumor types. miR-375 is a key downstream effector of achaete-scute homolog 1 (ASH1)-mediated induction of neuroendocrine features. The transcriptional coactivator of ASH1, YAP1, was determined to be a direct target of miR-375.37 ASH1 is a proneural basic helix-loop-helix transcription factor, leading to an exhibition of neuroendocrine features.51 In addition, a functional knockdown of ASH1 elicits prominent apoptosis in lung cancer cell lines with neuroendocrine features.52
Considering the results of the pathway analyses, in which the investigated miRNAs are involved, the MAPK pathway was identified to be the most affected signal transduction pathway. MAPK pathway is one of the most important evolutionary conserved intracellular signaling cascades.53, 54 MAPKs are intracellular serine/threonine kinases55 and the three most intensively studied kinases are the ERK, c-jun NH2-terminal kinase, and p38.53, 56, 57 Kit, proteinkinase B (PKB)/Akt and MAPK are expressed in a high percentage of small cell lung cancers.55 Furthermore, miRNAs targeting the PI3K/Akt/mTOR signaling pathway were differentially expressed in the four types of pulmonary neuroendocrine tumors investigated in this study.
It has to be kept in mind that the number of patients included is small and the results have to be regarded as a pilot. Owing to the excellent results, however, the proof of our findings in a larger cohort seems highly desirable.
In conclusion, the different subtypes of neuroendocrine lung tumors (typical carcinoids, atypical carcinoids, large cell neuroendocrine lung cancer, and small cell lung cancer) demonstrate significant differences in their miRNA expression profile. Moreover, significant insight into the molecular biology of pulmonary neuroendocrine tumors can be derived from these data. These might potentially be able to be used for diagnostic purposes and the assessment of prognosis. In particular, the expression of miR-29 family members is decreased paralleling an increasing grade of malignancy of the tumor. miRNA targeting protein expression of the MAPK and PI3K signaling pathways are differentially expressed in the various pulmonary neuroendocrine tumor subtypes. These data might prompt further scientific effort to explore potential novel therapeutic approaches for the treatment of these tumors. Nevertheless, a validation of these results on a bigger collective is urgently needed before these markers can be de facto integrated in a routine diagnostical setting.
References
Guan P, Yin Z, Li X et al. Meta-analysis of human lung cancer microRNA expression profiling studies comparing cancer tissues with normal tissues. J Exp Clin Cancer Res 2012;31:54.
Ma L, Huang Y, Zhu W et al. An integrated analysis of miRNA and mRNA expressions in non-small cell lung cancers. PloS one 2011;6:e26502.
Lee HW, Lee EH, Ha SY et al. Altered expression of microRNA miR-21, miR-155, and let-7a and their roles in pulmonary neuroendocrine tumors. Pathol Int. 2012;62:583–591.
Travis WD . Lung tumours with neuroendocrine differentiation. Eur J Cancer 2009;45 (Suppl 1):251–266.
Lee JH, Voortman J, Dingemans AM et al. MicroRNA expression and clinical outcome of small cell lung cancer. PloS one 2011;6:e21300.
Jackman DM, Johnson BE . Small-cell lung cancer. Lancet 2005;366:1385–1396.
Govindan R, Page N, Morgensztern D et al. Changing epidemiology of small-cell lung cancer in the United States over the last 30 years: analysis of the surveillance, epidemiologic, and end results database. J Clin Oncol 2006;24:4539–4544.
Travis WD et al. Tumours of the Lung In: World Health Organization International Histological Classification of Tumours. Pathology and Genetics of Tumours of the Lung, Pleura, Thymus and Heart 1st edn IARC Press: Lyon, 2004, pp 9–124.
den Bakker MA, Willemsen S, Grunberg K et al. Small cell carcinoma of the lung and large cell neuroendocrine carcinoma interobserver variability. Histopathology 2010;56:356–363.
Travis WD, Gal AA, Colby TV et al. Reproducibility of neuroendocrine lung tumor classification. Hum Pathol 1998;29:272–279.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281–297.
Caldas C, Brenton JD . Sizing up miRNAs as cancer genes. Nat Med 2005;11:712–714.
Lagos-Quintana M, Rauhut R, Lendeckel W et al. Identification of novel genes coding for small expressed RNAs. Science 2001;294:853–858.
Hutvagner G, Zamore PD . A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002;297:2056–2060.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006;6:857–866.
Iorio MV, Croce CM . MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol 2009;27:5848–5856.
Cho WC . MicroRNAs in cancer - from research to therapy. Biochim Biophys Acta 2010;1805:209–217.
Nunez-Iglesias J, Liu CC, Morgan TE et al. Joint genome-wide profiling of miRNA and mRNA expression in Alzheimer’s disease cortex reveals altered miRNA regulation. PloS one 2010;5:e8898.
Clop A . Insights into the importance of miRNA-related polymorphisms to heart disease. Hum Mutat 2009;30 doi:10.1002/humu.21087.
Haramati S, Chapnik E, Sztainberg Y et al. miRNA malfunction causes spinal motor neuron disease. Proc Natl Acad Sci USA 2010;107:13111–13116.
Ruvkun G . Clarifications on miRNA and cancer. Science 2006;311:36–37.
Bandres E, Agirre X, Ramirez N et al. MicroRNAs as cancer players: potential clinical and biological effects. DNA Cell Biol 2007;26:273–282.
Sassen S, Miska EA, Caldas C . MicroRNA: implications for cancer. Virchows Arch 2008;452:1–10.
Esquela-Kerscher A, Slack FJ . Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 2006;6:259–269.
Schmitz KJ, Helwig J, Bertram S et al. Differential expression of microRNA-675, microRNA-139-3p and microRNA-335 in benign and malignant adrenocortical tumours. J Clin Pathol 2011;64:529–535.
Sheu SY, Vogel E, Worm K et al. Hyalinizing trabecular tumour of the thyroid-differential expression of distinct miRNAs compared with papillary thyroid carcinoma. Histopathology 2010;56:632–640.
Sheu SY, Grabellus F, Schwertheim S et al. Differential miRNA expression profiles in variants of papillary thyroid carcinoma and encapsulated follicular thyroid tumours. Br J Cancer 2010;102:376–382.
Schwertheim S, Sheu SY, Worm K et al. Analysis of deregulated miRNAs is helpful to distinguish poorly differentiated thyroid carcinoma from papillary thyroid carcinoma. Horm Metab Res 2009;41:475–481.
Sheu SY, Grabellus F, Schwertheim S et al. Lack of correlation between BRAF V600E mutational status and the expression profile of a distinct set of miRNAs in papillary thyroid carcinoma. Horm Metab Res 2009;41:482–487.
He L, He X, Lim LP et al. A microRNA component of the p53 tumour suppressor network. Nature 2007;447:1130–1134.
Fabbri M, Garzon R, Cimmino A et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 2007;104:15805–15810.
Plaisier CL, Pan M, Baliga NS . A miRNA-regulatory network explains how dysregulated miRNAs perturb oncogenic processes across diverse cancers. Genome Res 2012;22:2302–2314.
Gebeshuber CA, Zatloukal K, Martinez J . miR-29a suppresses tristetraprolin, which is a regulator of epithelial polarity and metastasis. EMBO rep 2009;10:400–405.
Balca-Silva J, Sousa Neves S, Goncalves AC et al. Effect of miR-34b overexpression on the radiosensitivity of non-small cell lung cancer cell lines. Anticancer Res 2012;32:1603–1609.
Wang LG, Ni Y, Su BH et al. MicroRNA-34b functions as a tumor suppressor and acts as a nodal point in the feedback loop with Met. Int J Oncol 2013;42:957–962.
Wang Z, Chen Z, Gao Y et al. DNA hypermethylation of microRNA-34b/c has prognostic value for stage non-small cell lung cancer. Cancer Biol Ther 2011;11:490–496.
Nishikawa E, Osada H, Okazaki Y et al. miR-375 is activated by ASH1 and inhibits YAP1 in a lineage-dependent manner in lung cancer. Cancer Res 2011;71:6165–6173.
Johnson SM, Grosshans H, Shingara J et al. RAS is regulated by the let-7 microRNA family. Cell 2005;120:635–647.
Yanaihara N, Caplen N, Bowman E et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006;9:189–198.
Yu SL, Chen HY, Chang GC et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell 2008;13:48–57.
Markou A, Tsaroucha EG, Kaklamanis L et al. Prognostic value of mature microRNA-21 and microRNA-205 overexpression in non-small cell lung cancer by quantitative real-time RT-PCR. Clin Chem 2008;54:1696–1704.
Liu X, Sempere LF, Ouyang H et al. MicroRNA-31 functions as an oncogenic microRNA in mouse and human lung cancer cells by repressing specific tumor suppressors. J Clin Invest 2010;120:1298–1309.
Lebanony D, Benjamin H, Gilad S et al. Diagnostic assay based on hsa-miR-205 expression distinguishes squamous from nonsquamous non-small-cell lung carcinoma. J Clin Oncol 2009;27:2030–2037.
Wang XC, Du LQ, Tian LL et al. Expression and function of miRNA in postoperative radiotherapy sensitive and resistant patients of non-small cell lung cancer. Lung Cancer. 2011;72:92–99.
Bustin SA, Benes V, Garson JA et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009;55:611–622.
Walter RF, Mairinger FD, Wohlschlaeger J et al. FFPE tissue as a feasible source for gene expression analysis - a comparison of three reference genes and one tumor marker. Pathol Res Pract 2013;209:784–789.
Peiro-Chova L, Pena-Chilet M, Lopez-Guerrero JA et al. High stability of microRNAs in tissue samples of compromised quality. Virchows Arch 2013;463:765–774.
Park SY, Lee JH, Ha M et al. miR-29 miRNAs activate p53 by targeting p85 alpha and CDC42. Nat Struct Mol Biol 2009;16:23–29.
Cingarlini S, Bonomi M, Corbo V et al. Profiling mTOR pathway in neuroendocrine tumors. Target Oncol 2012;7:183–188.
Fang R, Xiao T, Fang Z et al. MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem 2012;287:23227–23235.
Osada H, Tomida S, Yatabe Y et al. Roles of achaete-scute homologue 1 in DKK1 and E-cadherin repression and neuroendocrine differentiation in lung cancer. Cancer Res 2008;68:1647–1655.
Osada H, Tatematsu Y, Yatabe Y et al. ASH1 gene is a specific therapeutic target for lung cancers with neuroendocrine features. Cancer Res 2005;65:10680–10685.
Vicent S, Garayoa M, Lopez-Picazo JM et al. Mitogen-activated protein kinase phosphatase-1 is overexpressed in non-small cell lung cancer and is an independent predictor of outcome in patients. Clin Cancer Res 2004;10:3639–3649.
Widmann C, Gibson S, Jarpe MB et al. Mitogen-activated protein kinase: conservation of a three-kinase module from yeast to human. Physiol Rev 1999;79:143–180.
Blackhall FH, Pintilie M, Michael M et al. Expression and prognostic significance of kit, protein kinase B, and mitogen-activated protein kinase in patients with small cell lung cancer. Clin Cancer Res 2003;9:2241–2247.
Su B, Karin M . Mitogen-activated protein kinase cascades and regulation of gene expression. Curr Opin Immunol 1996;8:402–411.
Cano E, Mahadevan LC . Parallel signal processing among mammalian MAPKs. Trends Biochem Sci 1995;20:117–122.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mairinger, F., Ting, S., Werner, R. et al. Different micro-RNA expression profiles distinguish subtypes of neuroendocrine tumors of the lung: results of a profiling study. Mod Pathol 27, 1632–1640 (2014). https://doi.org/10.1038/modpathol.2014.74
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.74
Keywords
This article is cited by
-
Zinc Gluconate Induces Potentially Cancer Chemopreventive Activity in Barrett’s Esophagus: A Phase 1 Pilot Study
Digestive Diseases and Sciences (2021)
-
Exosomal microRNA-144 from bone marrow-derived mesenchymal stem cells inhibits the progression of non-small cell lung cancer by targeting CCNE1 and CCNE2
Stem Cell Research & Therapy (2020)
-
Identifying the miRNA signature associated with survival time in patients with lung adenocarcinoma using miRNA expression profiles
Scientific Reports (2017)
-
Aberrant regulation of miR-15b in human malignant tumors and its effects on the hallmarks of cancer
Tumor Biology (2016)
-
A molecular signature for the prediction of recurrence in colorectal cancer
Molecular Cancer (2015)