Abstract
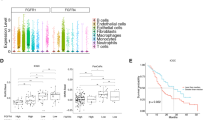
Solid-pseudopapillary neoplasm is an uncommon pancreatic tumor with distinct clinicopathologic features. Solid-pseudopapillary neoplasms are characterized by mutations in exon 3 of CTNNB1. However, little is known about the gene and microRNA expression profiles of solid-pseudopapillary neoplasms. Thus, we sought to characterize solid-pseudopapillary neoplasm-specific gene expression and identify the signaling pathways activated in these tumors. Comparisons of gene expression in solid-pseudopapillary neoplasm to pancreatic ductal carcinomas, neuroendocrine tumors, and non-neoplastic pancreatic tissues identified solid-pseudopapillary neoplasm-specific mRNA and microRNA profiles. By analyzing 1686 (1119 upregulated and 567 downregulated) genes differentially expressed in solid-pseudopapillary neoplasm, we found that the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways, as well as genes involved in epithelial mesenchymal transition, are activated in solid-pseudopapillary neoplasms. We validated these results experimentally by assessing the expression of β-catenin, WIF-1, GLI2, androgen receptor, and epithelial–mesenchymal transition-related markers with western blotting and immunohistochemistry. Our analysis also revealed 17 microRNAs, especially the miR-200 family and miR-192/215, closely associated with the upregulated genes associated with the three pathways activated in solid-pseudopapillary neoplasm and epithelial mesenchymal transition. Our results provide insight into the molecular mechanisms underlying solid-pseudopapillary neoplasm tumorigenesis and its characteristic less epithelial cell differentiation than the other common pancreatic tumors.
Similar content being viewed by others
Main
Solid-pseudopapillary neoplasm of the pancreas is rare,1, 2, 3 accounting for 1–2% of exocrine pancreatic tumors.4, 5, 6 Solid-pseudopapillary neoplasm is classified as a distinct phenotype due to its characteristic clinicopathologic features and distinct immunophenotype. The molecular genetics of solid-pseudopapillary neoplasm has recently become one of the best understood of all human tumors.1 Unlike the more common pancreatic adenocarcinoma, alterations in KRAS, TP53, CDKN2A, and SMAD4 have not been reported in solid-pseudopapillary neoplasm.7 Instead, almost all solid-pseudopapillary neoplasms exhibit somatic mutations in exon 3 of CTNNB1, which encodes β-catenin.7, 8 β-catenin is regulated by multiple signaling pathways through binding to proteins such as Tcf/Lef family members, axin, APC, and cadherins.9, 10 Under normal circumstances, β-catenin resides in the cytoplasm, where it is activated via phosphorylation and then degraded. However, this process is inhibited when exon 3 of CTNNB1 is mutated, which results in abnormal β-catenin nuclear translocation and the activation of transcription factors through formation of a β-catenin-Tcf/Lef complex.11, 12 Consistent with this, studies have demonstrated β-catenin nuclear accumulation in solid-pseudopapillary neoplasms.13
Although CTNNB1 mutation and activation of the Wnt/β-catenin pathway have been implicated in the pathogenesis of solid-pseudopapillary neoplasm, the molecular regulatory networks of solid-pseudopapillary neoplasm remain poorly understood. To date, no study has been performed to characterize microRNA expression in solid-pseudopapillary neoplasm, while only one study profiling gene expression has been reported. This investigation demonstrated the activation of the Wnt/β-catenin and Notch signaling pathways in solid-pseudopapillary neoplasm.14 However, direct comparison of the gene expression profiles of solid-pseudopapillary neoplasm to other common pancreatic carcinomas and endocrine tumors, which may provide insights into the cellular origin of solid-pseudopapillary neoplasm, has yet to be performed. Moreover, integrative analysis of the mRNA and microRNA expression profiles is necessary to identify the regulatory networks underlying the pathogenesis of solid-pseudopapillary neoplasm.
In this study, we compared the mRNA and microRNA expression profiles of solid-pseudopapillary neoplasms to those from patient-matched non-neoplastic pancreatic tissues and other ductal and endocrine pancreatic tumors. In doing so, we identified solid-pseudopapillary neoplasm-specific mRNA and microRNA expression. Moreover, we demonstrate an association between solid-pseudopapillary neoplasm and activation of the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways.
Materials and methods
Case Selection
A total of 14 solid-pseudopapillary neoplasms, six pancreatic adenocarcinomas, six neuroendocrine tumors, and five non-neoplastic pancreatic tissue samples were used in this study. The selected non-neoplastic tissues showed no evidence of chronic pancreatitis or preneoplastic lesions. The selected tissues mostly comprised pancreatic acinar cells and a few pancreatic islet cells, and therefore did not correspond exactly to the phenotypes of the three pancreatic neoplasms of our study. However, we used these non-neoplastic pancreatic tissues as a control, as they contained no neoplasms. The specimens were obtained from the archives of the Department of Pathology, Yonsei University, Seoul, Korea, and from the Liver Cancer Specimen Bank of the National Research Resource Bank Program of the Korean Science and Engineering Foundation of the Ministry of Science and Technology. Patient data were collected retrospectively from hospital records. All patients had undergone pancreatic resection between 2001 and 2011 and fresh snap-frozen samples were obtained immediately at the time of surgery. All carcinoma samples comprised ∼70% tumor cells, and none of the patients had received neo-adjuvant chemotherapy. Authorization for the use of these tissues for research purposes was obtained from the Institutional Review Board of Yonsei University of College of Medicine.
RNA Preparation
Total RNA was extracted using Trizol (Invitrogen Life Technologies, Carlsbad, CA, USA) according to the manufacturers’ protocol. After DNase digestion and other clean-up procedures, RNA samples were quantified, aliquoted, and stored at −80 °C until use. For quality control, RNA purity and integrity were evaluated by denaturing gel electrophoresis, measurement of the A260/280 ratio, and analyzed using the 2100 bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). For all samples, the RNA integrity number scores were >9.5.
Gene Expression Analysis
For DNA microarray hybridization, RNA was pooled by mixing equal amounts of total RNA. Biotin-labeled cRNA targets were synthesized starting from 1.5 μg of total RNA. Double-stranded cDNA synthesis was performed using the Illumina TotalPrep RNA Amplification Kit (Illumina, San Diego, CA, USA), while biotin-UTP-labeled antisense RNA was transcribed in vitro using Ambion’s Kit (Ambion Life Technologies, Carlsbad, CA, USA). All steps of the labeling procedure were performed according to the manufacturers’ protocol. Microarray experiments were conducted on the HumanHT-12 v4 Sentrix Expression BeadChip (Illumina). Hybridization of labeled cRNA to the BeadChip, washing, and scanning were performed according to the Illumina Bead Station 500 × manual.
MicroRNA Expression Analysis
To monitor the changes in microRNA levels associated with solid-pseudopapillary neoplasm, 100 ng of total RNA was labeled and hybridized using the Human microRNA Microarray Kit (v.16, Agilent Technologies) according to the manufacturer’s protocol (Agilent microRNA microarrays Version 1.5). Hybridization signals were detected with a DNA microarray scanner (Agilent Technologies) and the scanned images were analyzed using Agilent feature extraction software.
mRNA Gene Expression Data Preparation and Statistical Analysis
Raw data were extracted using the software provided by the manufacturer (Illumina Genome Studio v2011.1 (Gene Expression Module v1.9.0)). Expression intensities were normalized using quantile normalization techniques.15 Using the normalized intensities, differentially expressed genes between non-neoplastic pancreatic tissue and pancreatic tumors (solid-pseudopapillary neoplasm, neuroendocrine tumor, or pancreatic adenocarcinoma) were determined using the integrated statistical method previously reported.16 Briefly, (1) two independent tests were performed: Student’s t-test and log2–median-ratio test; (2) adjusted P-values from each test were computed using an empirical distribution of the null hypothesis that the means of the genes were not different, which was obtained from random permutations of the samples; (3) the P values from the two tests were combined to compute the overall P-values using Stouffer’s method17 and (4) differentially expressed genes were selected as those with P<0.01 and a fold change >1.5. Finally, functional enrichment analysis of the differentially expressed genes was performed using DAVID software18 to identify GO biological processes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways represented by the genes in individual clusters with statistical significance (Supplementary Figure 2A and B).
MicroRNA Raw Data Preparation and Statistical Analysis
Raw data were extracted using the Agilent Feature Extraction Software (v 10.7.1.1) and then processed as published previously.19 The method described above to analyze mRNA gene expression data was also used to identify differentially expressed microRNAs with P<0.01 and a fold change of >1.5.
Analysis of mRNA-micro RNA Network
We searched for possible mRNAs targeted by the 30 microRNAs specifically downregulated in solid-pseudopapillary neoplasm using miRBase,20 Targetscan,21 and Miranda.22 After the initial screening, we computed the Pearson’s correlation coefficient using expression profiles of the mRNAs and microRNAs, and then selected mRNA-microRNA pairs with correlation <−0.5 and P-value<0.01. Among these mRNA-microRNA pairs, we finally selected mRNAs specifically upregulated in solid-pseudopapillary neoplasm and associated with the three activated signaling pathways as the microRNA targets.
Quantitative RT-PCR
Primer sequences used were obtained from the Primer-Bank database (http://pga.mgh.harvard.edu/primerbank/) and are listed in Supplementary Table 5. The reaction was carried out in a final volume of 20 μl with Premix Ex Taq (Takara Bio Otsu, Japan) according to the manufacturer’s instructions.
Immunohistochemistry
Paraffin-embedded tissue blocks were cut into 4-μm sections. Immunohistochemical analysis was performed using a Ventana XT automated stainer (Ventana, Tucson, AZ, USA) with antibodies against β-catenin (diluted 1:100, BD Biosciences, San Jose, CA, USA), WIF-1 (diluted 1:50, BD Biosciences), GLI2 (diluted 1:100, Santa Cruz Biotechnology, Santa Cruz, CA, USA), androgen receptor (prediluted; Roche, Mannheim, Germany), E-cadherin for extra-cellular domains (diluted 1:100, Dako, Carpinteria, CA, USA) and intracellular domains (diluted 1:200, BD Biosciences) and N-cadherin (diluted 1:50, Abcam, Cambridge, UK).
Results
Clinicopathologic and Molecular Characteristics of Solid-pseudopapillary Neoplasms
Before performing mRNA and microRNA expression analysis, solid-pseudopapillary neoplasm was confirmed by immunohistochemistry and other molecular diagnostic tests. We performed immunostaining of β-catenin, vimentin, and CD10 for diagnosis of solid-pseudopapillary neoplasm and immunostaining of chromogranin A for diagnosis of neuroendocrine tumors (Supplementary Table 1). In doing so, we confirmed that all of the 14 solid-pseudopapillary neoplasms expressed nuclear β-catenin, vimentin, and CD10. We also demonstrated that all six neuroendocrine tumors expressed chromogranin A. Unlike the pancreatic adenocarcinomas, neuroendocrine tumors, and non-neoplastic pancreas analyzed, all 14 solid-pseudopapillary neoplasms exhibited mutation in exon 3 of CTNNB1 (Table 1 and Supplementary Table 1).
Unsupervised Clustering Analysis of mRNA and MicroRNA Expression Distinguishes Solid-pseudopapillary Neoplasm as a Distinct Type of Pancreatic Tumor
As a first step in the molecular analysis of solid-pseudopapillary neoplasms, we examined whether the mRNA and microRNA expression profiles are distinct among the different pancreatic tumors examined. mRNA and microRNA expression profiling was performed on 14 solid-pseudopapillary neoplasms, six pancreatic adenocarcinomas, six neuroendocrine tumors, and five non-neoplastic pancreatic tissues using the Human HT-12 v4 Expression Bead Chip, which contains 47 231 probes representing 31 332 annotated genes. Unsupervised hierarchical clustering analysis of 6777 differentially expressed genes in 31 microarrays grouped non-neoplastic pancreas, solid-pseudopapillary neoplasm, pancreatic adenocarcinoma, and neuroendocrine tumor into separate clusters, suggesting that these tissue types could be characterized by unique differential gene expression (Figure 1a).
Unsupervised hierarchical clustering analysis of mRNA and microRNA expression profiles of pancreatic tumors. (a) Unsupervised hierarchical clustering analysis of gene expression. Red and green indicate transcript levels above and below the sample median, respectively. Five non-neoplastic pancreatic tissues, 6 pancreatic adenocarcinomas (PCA), 14 solid-pseudopapillary neoplasm (SPN), and 6 neuroendocrine tumors (NET) were clustered based on gene expression patterns. Non-neoplastic pancreas, pancreatic adenocarcinomas, solid-pseudopapillary neoplasms, and neuroendocrine tumors are indicated by green, pink, yellow, and blue bars, respectively. (b) Unsupervised hierarchical clustering analysis of microRNA expression. Complete separation of non-neoplastic pancreas, pancreatic adenocarcinomas, SPNs, and neuroendocrine tumors were evident based on microRNA expression profiles. Non-neoplastic pancreas, pancreatic adenocarcinomas, solid-pseudopapillary neoplasms, and neuroendocrine tumors are indicated by green, pink, yellow, and blue bars, respectively. (c) Clustering analysis using the filtered 6777 differentially expressed genes in 26 pancreatic tumors composed of six pancreatic adenocarcinomas, 14 solid-pseudopapillary neoplasms, and six neuroendocrine tumors. Each mRNA expression value was divided by the median value of five non-neoplastic pancreatic tissues. Color bar gradient represents log2-fold changes in each pancreatic tumor compared to non-neoplastic pancreas. (d) Clustering expressed microRNAs in 27 samples composed of 6 pancreatic adenocarcinomas, 14 solid-pseudopapillary neoplasms, and 6 neuroendocrine tumors. Each microRNA expression value was divided by the median value of five non-neoplastic pancreatic tissues.
Clustering analysis based on the 232 differentially expressed microRNAs revealed substantial distinctions in over- and underrepresented microRNAs according to the tumor type. As evident from the resultant dendrogram, the 31 samples clustered into four groups according to the tissue type, demonstrating that microRNA expression patterns were distinct according to the type of pancreatic tumor similar to mRNA expression (Figure 1b). These gene expression and microRNA microarray data discussed in this study have been deposited in NCBI’s Gene Expression Omnibus (GEO) database (http://www.ncbi.nih.gov/geo/), and are accessible through GEO Series accession number GSE43795, GSE43796 and GSE43797.
The distinct gene and microRNA expression profiles among the pancreatic tumors were also demonstrated when unsupervised hierarchical clustering analysis of mRNAs (Figure 1c) and microRNAs (Figure 1d) were performed by using the expression data relative to non-neoplastic samples: mRNA and microRNA expression value divided by the median value of five non-neoplastic pancreatic tissues.
Solid-pseudopapillary Neoplasm-specific Gene Expression Profiles
To identify solid-pseudopapillary neoplasm-specific gene expression, we compared the gene expression profiles of solid-pseudopapillary neoplasms, pancreatic adenocarcinomas, and neuroendocrine tumors with those of non-neoplastic pancreatic tissues, namely solid-pseudopapillary neoplasm versus non-neoplastic samples, neuroendocrine tumor versus non-neoplastic samples, and pancreatic adenocarcinoma versus non-neoplastic samples. Using an integrative statistical method that we previously reported (Materials and methods), we identified 6777 differentially expressed genes (P<0.01, fold-change >1.5), of which 4134 and 2807 were up- and downregulated, respectively, in solid-pseudopapillary neoplasms, pancreatic adenocarcinomas, and neuroendocrine tumors compared with non-neoplastic samples (Figures 2a–c). While 1119 genes were exclusively upregulated in solid-pseudopapillary neoplasm, 488 were upregulated in both pancreatic adenocarcinoma and solid-pseudopapillary neoplasm. Meanwhile, 660 genes were upregulated in both neuroendocrine tumor and solid-pseudopapillary neoplasm (Figure 2a). Among the downregulated genes, 567 were exclusively downregulated in solid-pseudopapillary neoplasm. Pancreatic adenocarcinoma and solid-pseudopapillary neoplasm shared 819 downregulated genes, while 955 were downregulated in both neuroendocrine tumor and solid-pseudopapillary neoplasm (Figure 2b). The 6777 differentially expressed genes were further categorized into 25 clusters based on differential expression in the three comparisons (Supplementary Figure 1A). Clusters 5 and 21 revealed that a majority of the differentially expressed genes found in solid-pseudopapillary neoplasms (1119 of 2026 upregulated and 567 of 1757 downregulated genes) were specific to this tumor (Figure 2c), thereby defining a molecular signature for solid-pseudopapillary neoplasm tumorigenesis.
mRNAs and microRNAs are differentially expressed in non-neoplastic pancreas tissues and pancreatic tumors. Differentially expressed gene profiles between non-neoplastic pancreatic tissues and pancreatic tumors. (a–c) Venn diagram analysis of mRNAs (a) upregulated and (b) downregulated in solid-pseudopapillary neoplasm relative to non-neoplastic pancreas and other pancreatic tumors. (c) Heatmap of mRNA expression profiles of solid-pseudopapillary neoplasm-specific up- or downregulated genes. MicroRNA expression profiles differentially expressed in non-neoplastic pancreatic tissues and pancreatic tumors. (d–f) Venn diagram analysis of microRNAs (d) upregulated and (e) downregulated in solid-pseudopapillary neoplasm relative to non-neoplastic pancreas and other pancreatic tumors. (f) Heatmap of microRNA expression profiles of SPN-specific up- or downregulated genes.
Identification of Characteristic Signaling Pathways Associated with Solid-pseudopapillary Neoplasm
We then examined whether the solid-pseudopapillary neoplasm-specific up- and downregulated genes are associated with signaling pathways by performing an enrichment analysis of GO biological processes and KEGG pathways using DAVID software (Supplementary Figure 2). GO biological processes enrichment analysis demonstrated that the solid-pseudopapillary neoplasm-specific upregulated genes represent several signaling pathways, including the Wnt/β-catenin, Hedgehog, and androgen receptor signaling cascades (Supplementary Figure 2A). KEGG pathway enrichment analysis also revealed an association with the Wnt/β-catenin and Hedgehog signaling pathways in addition to identifying other pathways represented by the upregulated genes (eg, ubiquitin-mediated proteolysis) (Supplementary Figure 2B). GO biological process analysis of the downregulated genes demonstrated their association with cell motility, migration, and blood vessel development (Supplementary Figure 2A). Moreover, the KEGG pathways represented by the downregulated genes included fructose and mannose metabolism, the MAPK signaling pathway, and endocytosis (Supplementary Figure 2B). Notably, comparison of the GO biological process and KEGG pathways found in the 25 differentially expressed genes clusters (Supplementary Figure 1A) demonstrated that the genes specifically upregulated in solid-pseudopapillary neoplasms were exclusively related to the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways (cluster 5 in Supplementary Figure 2, Table 2).
Expression of Tumor Differentiation Markers
Solid-pseudopapillary neoplasms exhibit little cytologic and histologic differentiation. Therefore, solid-pseudopapillary neoplasms have been speculated to possess a stem cell-like trait.23 Also, studies have reported that ∼50% of solid-pseudopapillary neoplasms express KIT, a well-known stem cell marker.24 Interestingly, examination of stem cell marker expression in our microarray data did not reveal any significant changes in these markers in solid-pseudopapillary neoplasms (Supplementary Table 2). Instead, the expression of genes associated with epithelial–mesenchymal transition changed dramatically in solid-pseudopapillary neoplasms. Among the well-known epithelial mesenchymal transition markers,25, 26 E-cadherin (CDH1) expression was drastically decreased (−59-fold), whereas the expression of epithelial mesenchymal transition regulators, such as TWIST2 (17.09-fold) and ZEB2 (3.89-fold), was increased.
Solid-pseudopapillary Neoplasm-specific MicroRNA Expression Profiles
Using the same comparisons and statistical methods, we identified 232 differentially expressed microRNAs. We found 49 microRNAs specifically upregulated in solid-pseudopapillary neoplasms, whereas 30 were specifically downregulated (P<0.01, fold-change >1.5) (Figures 2d–f). The expression of these 232 microRNAs was categorized into 21 clusters (Supplementary Figure 1B). We focused on cluster 18, which includes 30 microRNAs specifically downregulated in solid-pseudopapillary neoplasm. Of these, nine microRNAs (miR-200c-3p, miR-192-5p, miR-215, miR-14-3p, miR141-5p, miR-127-3p, miR-379-5p, miR-338-3p, and miR-455-5p) were predicted to target >100 genes belonging to cluster 5 from our mRNA microarray analysis (ie, mRNAs specifically upregulated in solid-pseudopapillary neoplasm). Six additional microRNAs (miR-429, miR-7-5p, miR200b-3p, miR-192-3p, miR200b-5p, and miR-200a-3p) from clusters 14, 15, and 17, which consisted of microRNAs that decreased in solid-pseudopapillary neoplasm but increased in pancreatic adenocarcinomas and/or neuroendocrine tumors, were predicted to target >100 mRNAs belonging to cluster 5 of our mRNA expression analysis.
mRNA-microRNA Regulatory Network in Solid-pseudopapillary Neoplasm
Based on these results, we hypothesized that the 30 microRNAs specifically downregulated in solid-pseudopapillary neoplasm targeted genes involved in the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways. Thus, we examined the microRNAs predicted to target genes associated with these signaling pathways and that belonged to cluster 5 from our mRNA analysis. To accomplish this, the target genes of the 30 microRNAs were identified using miRBase,20 TargetScan Human,21 and Miranda.22 Then, target genes whose expression was anti-correlated with that of microRNAs (ie, up-regulated specifically in solid-pseudopapillary neoplasms; Materials and methods) and involved in the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways (Table 2) were grouped. This analysis identified 17 microRNAs that regulate genes involved in Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways (Supplementary Table 4).
Using this microRNA-mRNA target information, we constructed gene expression and a microRNA regulatory network specifically activated in solid-pseudopapillary neoplasm (Figure 3, Supplementary Figure 3). The network in Figure 3 shows that the upregulated mRNAs of the Wnt/β-catenin pathway, including WIF1, Axin2, FZD7, PRKCZ, and Wnt5A, may be regulated by the miR-200 family, miR-192/215, and miR-455 (Supplementary Figure 3, Supplementary Table 4). Figure 3b shows the regulation of Hedgehog pathway components, such as GLI2 and GLI3, by miR-141/200a, while androgen receptor could be targeted by miR-376b. According to these models, the miR-200bc/429, miR-192/215, and miR-141/200a families may contribute to upregulating Wnt/β-catenin, Hedgehog, and androgen receptor pathway gene expression (Supplementary Table 4).
Gene expression signature changes leading to activated signaling pathways in solid-pseudopapillary neoplasm. (a) Gene expression changes leading to activation of the Wnt/β-catenin signaling pathway in solid-pseudopapillary neoplasm (SPN). Alterations were defined by (1) greater than two-fold mean up- or downregulation in gene expression, and (2) >1.5-fold change in gene expression in all 14 solid-pseudopapillary neoplasms. The number in the left inside box denotes the number of altered cases/total cases. The number in the inside right box denotes the mean fold change of the gene. Red box, upregulated genes; yellow circle, genes with activating mutation; p, phosphorylation; ub, ubiquitination. (b) Gene expression changes leading to activation of the Hedgehog signaling pathway in solid-pseudopapillary neoplasm. Alterations were defined using the same criteria as in panel a.The number in the left inside box denotes the number of altered cases/total cases. The number in the inside right box denotes the mean fold change of the gene. red box, up-regulated genes; blue box, down-regulated genes.
Validation of Differentially Expressed mRNAs by Quantitative RT-PCR
Changes in mRNA levels of differentially expressed genes in solid-pseudopapillary neoplasm, pancreatic adenocarcinomas, and neuroendocrine tumors, compared with non-neoplastic pancreas, were confirmed by quantitative real-time PCR. 18S rRNA was used as a control for normalization. We selected CTNNB1 and Axin2 to assess the Wnt/β-catenin signaling pathway, GLI2 for the Hedgehog pathway, and androgen receptor for the androgen receptor signaling pathway. E-cadherin (CDH1) and N-cadherin (CDH2) were chosen as representative genes of epithelial–mesenchymal transition. Marked upregulation of CTNNB1 was found in solid-pseudopapillary neoplasm, while only a slight increase was observed in pancreatic adenocarcinoma and neuroendocrine tumor compared with non-neoplastic pancreatic tissues (Figure 4). Expression of Axin2, GLI2, and androgen receptor were markedly higher in solid-pseudopapillary neoplasm relative to pancreatic adenocarcinoma, neuroendocrine tumor, and non-neoplastic pancreatic tissues (Figure 4). Marked downregulation of E-cadherin, an epithelial marker, was found in solid-pseudopapillary neoplasm compared with normal pancreas and other pancreatic tumors. In contrast, the expression of N-cadherin, a mesenchymal marker, was significantly increased in solid-pseudopapillary neoplasm and neuroendocrine tumor relative to non-neoplastic pancreatic tissues and pancreatic adenocarcinoma (Figure 4).
Validation of the expression of genes involved in the signaling pathways activated in solid-pseudopapillary neoplasms and epithelial–mesenchymal transition. mRNA levels of CTNNB1 (β-catenin), Axin2, GLI2, androgen receptor (AR), CDH1 (E-cadherin) and CDH2 (N-cadherin) were verified by quantitative RT-PCR. The expression level of each mRNA was normalized to that of 18s rRNA and arbitrarily set to 1. *P<0.05; **P<0.01 based on the Student’s t-test.
Validation of Differentially Expressed Genes in Solid-pseudopapillary Neoplasm by Western Blotting and Immunohistochemistry
We validated β-catenin, WIF-1, GLI2, androgen receptor, E-cadherin, and N-cadherin expression by western blotting (Figure 5a) and immunohistochemistry (Figure 5b). We could not include one case of solid-pseudopapillary neoplasm in the western blotting analysis due to a lack of remaining specimen. Both western blotting and immunohistochemical analysis revealed increased levels of β-catenin, WIF-1, GLI2, androgen receptor, and N-cadherin, along with decreases in E-cadherin in solid-pseudopapillary neoplasms.
Representative western blotting and immunohistochemical analysis of the proteins involved in Wnt signaling, Hedgehog signaling, androgen receptor signaling pathway, and epithelial–mesenchymal transition in pancreatic tumors. Protein levels of components of the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways were assessed in solid-pseudopapillary neoplasm (SPN), non-neoplastic pancreatic tissues, pancreatic adenocarcinomas (PCA), and neuroendocrine tumors (NET) by western blot (a) and immunohistochemistry (b).
All 13 solid-pseudopapillary neoplasms showed increased β-catenin expression, whereas pancreatic adenocarcinomas and neuroendocrine tumors showed slightly increased β-catenin expression compared with non-neoplastic pancreas by western blot. WIF-1 protein was also highly expressed in solid-pseudopapillary neoplasm tumor tissues relative to matched normal pancreatic tissues by western blot. GLI2 was highly expressed in all solid-pseudopapillary neoplasms and three out of six pancreatic adenocarcinomas, but was not expressed in neuroendocrine tumors and non-neoplastic pancreas according to western blot. Androgen receptor was highly expressed in all of 13 solid-pseudopapillary neoplasms, but not expressed in pancreatic adenocarcinomas, neuroendocrine tumors, or non-neoplastic pancreas on western blot. Of the epithelial–mesenchymal transition-related markers, E-cadherin expression was detected in 1 out of 13 solid-pseudopapillary neoplasms, 3 out of 6 neuroendocrine tumors, and 5 out of 6 pancreatic adenocarcinomas by using E-cadherin antibody targeting for intracellular domains; N-cadherin expression was found in 8 out of 13 solid-pseudopapillary neoplasms, 4 out of 6 neuroendocrine tumors. None of the six pancreatic adenocarcinomas were assessed by western blot.
Immunohistochemical analysis revealed characteristic β-catenin nuclear accumulation exclusively in all 14 solid-pseudopapillary neoplasms, whereas no nuclear β-catenin expression was found in pancreatic adenocarcinomas or neuroendocrine tumors. Strong cytoplasmic WIF-1 expression, and strong nuclear expressions of GLI2 and androgen receptor were demonstrated in all 14 solid-pseudopapillary neoplasms (Figure 5b). E-cadherin expression was not detected in any of the 14 solid-pseudopapillary neoplasms, but was found in 3 out of 6 neuroendocrine tumors and all of 6 pancreatic adenocarcinomas by immunohistochemistry. In accordance with previous reports, we also observed nuclear E-cadherin expression in all 14 solid-pseudopapillary neoplasms and in 1 out of 6 neuroendocrine tumors by antibody targeting for intracellular domains (Supplementary Figure 4). Nuclear localization of E-cadherin in solid-pseudopapillary neoplasms has been reported, and proposed to be mechanistically related to mutations in β-catenin.27, 28 In contrast to rare membranous E-cadherin expression in solid-pseudopapillary neoplasms, N-cadherin expression was found in 12 out of 14 solid-pseudopapillary neoplasms. E-cadherin, but not N-cadherin, was expressed in the cell membranes of normal pancreas acinar cells (Figure 5b).
Discussion
In this study, we characterized the mRNA and microRNA expression profiles of solid-pseudopapillary neoplasm using a DNA microarray and microRNA array approach and compared them with other pancreatic tumors. We confirmed that solid-pseudopapillary neoplasm is a unique tumor based on mRNA and microRNA expression patterns. We also demonstrated that the Wnt/β-catenin signaling pathway is activated in solid-pseudopapillary neoplasms and revealed the involvement of the Hedgehog and androgen receptor signaling pathways. By comparing the mRNA-microRNA regulatory network of solid-pseudopapillary neoplasms, we identified several microRNAs that may participate in upregulating genes associated with these three signaling pathways.
Although solid-pseudopapillary neoplasm is rare, understanding its molecular mechanisms is still important and advantageous. Solid-pseudopapillary neoplasm is a uniquely homogeneous tumor in terms of cell morphology and genetic changes. Nearly all solid-pseudopapillary neoplasms exhibit gain of function mutations in exon 3 of CTNNB1. Thus, knowledge about the molecular ramifications of this mutation is also of great importance. Here, we provide a broad unbiased view of the molecular characteristics of solid-pseudopapillary neoplasm at the mRNA and microRNA levels by analyzing genes specifically deregulated in solid-pseudopapillary neoplasms. The gene expression profile of solid-pseudopapillary neoplasm was compared with non-neoplastic pancreas and other pancreatic tumors. Previous studies of solid-pseudopapillary neoplasm have reported the activating CTNNB1 mutations and abnormal β-catenin nuclear accumulation due to inhibition of its degradation. Thus, activation of downstream β-catenin targets was expected. CTNNB1 mutation in solid-pseudopapillary neoplasm is heterozygous and many genes encoding components of the Wnt/β-catenin pathway have been shown to be upregulated in solid-pseudopapillary neoplasm by our study and others.14 Therefore, nuclear accumulation of wild type β-catenin protein was also expected. Interestingly, both studies found that the level of CTNNB1 mRNA is approximately three to four-fold upregulated in solid-pseudopapillary neoplasm. The proportion of mutant to wild-type β-catenin protein in the nucleus and its effects on downstream targets should be studied in the future.
Besides the Wnt/β-catenin pathway, solid-pseudopapillary neoplasm also exhibits upregulation of genes involved in activation of the Hedgehog and androgen receptor signaling pathways. Based on our analysis of upregulated mRNAs in solid-pseudopapillary neoplasm, we propose that solid-pseudopapillary neoplasm is characterized by activation of the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways. At present, there has been only one report demonstrating the gene expression profiles of solid-pseudopapillary neoplasm. This study showed that the Wnt/β-catenin and Notch signaling pathways participate in solid-pseudopapillary neoplasm.14 Consistent with this, our data demonstrate that Notch signaling genes were differentially expressed in solid-pseudopapillary neoplasm compared with non-neoplastic pancreatic tissues. However, because these genes were also differentially expressed in pancreatic adenocarcinomas and/or neuroendocrine tumors, this pathway is not regarded as specifically activated pathway in solid-pseudopapillary neoplasm. Instead, our results suggest that the Hedgehog pathway is significantly activated in solid-pseudopapillary neoplasms. The Notch and Hedgehog signaling pathways are closely related to the Wnt/β-catenin pathway, with many components in common.29, 30
One novel finding of this study is activation of the androgen receptor signaling pathway in solid-pseudopapillary neoplasm. We have demonstrated increased androgen receptor expression at the transcriptional and translational levels. Studies have shown that androgen receptor binds to the armadillo repeat domain of β-catenin and enhances its nuclear localization and androgen receptor.31 Increased androgen receptor-dependent transcriptional activity was reported in prostatic tumors with CTNNB1 mutations.32 However, no direct relationship between activation of the Wnt/β-catenin and androgen receptor signaling pathways has been reported. In this study, we demonstrate for the first time a high level of nuclear androgen receptor expression in all 14 solid-pseudopapillary neoplasms. In addition to the molecular link between the Wnt/β-catenin and androgen receptor signaling pathways, tumor cells of solid-pseudopapillary neoplasm are characterized by nuclear expression of androgen receptor. These results suggest that androgen receptor may be a potential diagnostic marker for solid-pseudopapillary neoplasm.
The molecular mechanisms leading to the upregulation of these genes in solid-pseudopapillary neoplasms are mostly unknown. We evaluated the possibility that this upregulation is related to the mRNA-microRNA regulatory network. As we performed the mRNA and microRNA gene expression analysis with the same solid-pseudopapillary neoplasm samples, we could evaluate the mRNA-microRNA reciprocal relationship in our solid-pseudopapillary neoplasms. Our analysis revealed distinct microRNA expression patterns specific to solid-pseudopapillary neoplasm. In particular, 30 microRNAs were found to be specifically downregulated in solid-pseudopapillary neoplasms. Based on this data, we showed that a large number of solid-pseudopapillary neoplasm-specific upregulated genes belonging to the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways are associated with downregulated microRNAs. For instance, downregulation of the miR-200 family and miR-192/215 were related to the upregulation of components of these signaling pathways in solid-pseudopapillary neoplasm.
Finally, we found that solid-pseudopapillary neoplasm altered the expression of genes linked to epithelial–mesenchymal transition. Marked downregulation of E-cadherin (CDH1) was found in solid-pseudopapillary neoplasms, whereas upregulation of N-cadherin (CDH2) and vimentin were validated by western blotting. These findings are in accordance with the rare epithelial differentiation observed in solid-pseudopapillary neoplasm tumor cells. Interestingly, upregulation of epithelial mesenchymal transition regulator genes, TWIST2 and ZEB2, was also observed. A recent study demonstrated that the miRNA-200 family (miR-141, miR-200a, b, c, and miR-429) and miR-192/215 inhibit E-cadherin expression by targeting ZEB1 and ZEB2.33, 34, 35, 36, 37 These previous reports suggest that solid-pseudopapillary neoplasm-specific downregulation of the miR-200 family and miR-192/215 targets components of the Wnt/β-catenin, Hedgehog, androgen receptor signaling pathways as well as genes associated with epithelial–mesenchymal transition. These effects ultimately result in decreased expression of epithelial markers. Interestingly, the expression of epithelial–mesenchymal transition-related genes is linked with activation of the Wnt/β-catenin, Notch, and Hedgehog signaling pathways.30, 38, 39, 40, 41
We propose that solid-pseudopapillary neoplasm is characterized by activation of Wnt/β-catenin, Hedgehog, androgen receptor signaling pathways and epithelial–mesenchymal transition based on the signature of differentially expressed mRNAs in solid-pseudopapillary neoplasm. Analysis of microRNA expression profiles suggests that a large number of solid-pseudopapillary neoplasm-specific upregulated genes belonging to the Wnt/β-catenin, Hedgehog, and androgen receptor signaling pathways are associated with downregulated microRNAs. Especially, the downregulated miR-200 family and miR-192/215 are related to the upregulation of genes belong to epithelial–mesenchymal transition in solid-pseudopapillary neoplasm. These mRNA and microRNA expression profiles may explain reports of less epithelial cell differentiation in solid-pseudopapillary neoplasm than other common pancreatic tumors, thereby providing a molecular mechanism for solid-pseudopapillary neoplasm tumorigenesis.
Accession codes
References
Bosman FT, Carneiro F, Hruban RH et al (eds).. WHO classification of tumours of the digestive system 4th edn World Health Organization & International Agency for Research on Cancer: Lyon, 2010, pp 327–330.
Hruban RH, Pitman MB, Klimstra DS, (eds). Tumors of the Pancreas, 6th edn. American Registry of Pathology & Armed Forces Institute of Pathology: Washington, 2007, pp 231–250.
Kosmahl M, Pauser U, Anlauf M et al. [Cystic pancreas tumors and their classification: features old and new]. Pathologe 2005;26:22–30.
Buckley JA, Vaughn DD, Jabra AA et al. CT evaluation of mediastinal masses in children: spectrum of disease with pathologic correlation. Crit Rev Diagn Imaging 1998;39:365–392.
Ganne-Carrie N, Chastang C, Chapel F et al. Predictive score for the development of hepatocellular carcinoma and additional value of liver large cell dysplasia in Western patients with cirrhosis. Hepatology 1996;23:1112–1118.
Huang SC, Erdman SH . Pediatric juvenile polyposis syndromes: an update. Curr Gastroenterol Rep 2009;11:211–219.
Abraham SC, Klimstra DS, Wilentz RE et al. Solid-pseudopapillary tumors of the pancreas are genetically distinct from pancreatic ductal adenocarcinomas and almost always harbor beta-catenin mutations. Am J Pathol 2002;160:1361–1369.
Tanaka Y, Kato K, Notohara K et al. Frequent beta-catenin mutation and cytoplasmic/nuclear accumulation in pancreatic solid-pseudopapillary neoplasm. Cancer Res 2001;61:8401–8404.
Barker N, Clevers H . Catenins, Wnt signaling and cancer. Bioessays 2000;22:961–965.
Hlsken J, Behrens J . The Wnt signalling pathway. J Cell Sci 2000;113 (Pt 20):3545.
Barth AI, Nathke IS, Nelson WJ . Cadherins, catenins and APC protein: interplay between cytoskeletal complexes and signaling pathways. Curr Opin Cell Biol 1997;9:683–690.
Morin PJ, Sparks AB, Korinek V et al. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science 1997;275:1787–1790.
Aoki M, Hecht A, Kruse U et al. Nuclear endpoint of Wnt signaling: neoplastic transformation induced by transactivating lymphoid-enhancing factor 1. Proc Natl Acad Sci USA 1999;96:139–144.
Cavard C, Audebourg A, Letourneur F et al. Gene expression profiling provides insights into the pathways involved in solid pseudopapillary neoplasm of the pancreas. J Pathol 2009;218:201–209.
Bolstad BM, Irizarry RA, Astrand M et al. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003;19:185–193.
Lee HJ, Suk JE, Patrick C et al. Direct transfer of alpha-synuclein from neuron to astroglia causes inflammatory responses in synucleinopathies. J Biol Chem 2010;285:9262–9272.
Hwang D, Rust AG, Ramsey S et al. A data integration methodology for systems biology. Proc Natl Acad Sci USA 2005;102:17296–17301.
Huang da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009;4:44–57.
Lopez-Romero P, Gonzalez MA, Callejas S et al. Processing of Agilent microRNA array data. BMC Res Notes 2010;3:18.
Kozomara A, Griffiths-Jones S . miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011;39:D152–D157.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005;120:15–20.
Betel D, Koppal A, Agius P et al. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol 11:R90.
Kosmahl M, Seada LS, Janig U et al. Solid-pseudopapillary tumor of the pancreas: its origin revisited. Virchows Arch 2000;436:473–480.
Cao D, Antonescu C, Wong G et al. Positive immunohistochemical staining of KIT in solid-pseudopapillary neoplasms of the pancreas is not associated with KIT/PDGFRA mutations. Mod Pathol 2006;19:1157–1163.
Jechlinger M, Grunert S, Tamir IH et al. Expression profiling of epithelial plasticity in tumor progression. Oncogene 2003;22:7155–7169.
Lee JM, Dedhar S, Kalluri R et al. The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J Cell Biol 2006;172:973–981.
Chetty R, Serra S . Membrane loss and aberrant nuclear localization of E-cadherin are consistent features of solid pseudopapillary tumour of the pancreas. An immunohistochemical study using two antibodies recognizing different domains of the E-cadherin molecule. Histopathology 2008;52:325–330.
Tang WW, Stelter AA, French S et al. Loss of cell-adhesion molecule complexes in solid pseudopapillary tumor of pancreas. Mod Pathol 2007;20:509–513.
Jiang J, Hui CC . Hedgehog signaling in development and cancer. Dev Cell 2008;15:801–812.
Katoh Y, Katoh M . Hedgehog signaling, epithelial-to-mesenchymal transition and miRNA (review). Int J Mol Med 2008;22:271–275.
Yang F, Li X, Sharma M et al. Linking beta-catenin to androgen-signaling pathway. J Biol Chem 2002;277:11336–11344.
Truica CI, Byers S, Gelmann EP . Beta-catenin affects androgen receptor transcriptional activity and ligand specificity. Cancer Res 2000;60:4709–4713.
Gregory PA, Bracken CP, Bert AG et al. MicroRNAs as regulators of epithelial-mesenchymal transition. Cell Cycle 2008;7:3112–3118.
Korpal M, Lee ES, Hu G et al. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 2008;283:14910–14914.
Park SM, Gaur AB, Lengyel E et al. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 2008;22:894–907.
Paterson EL, Kolesnikoff N, Gregory PA et al. The microRNA-200 family regulates epithelial to mesenchymal transition. Sci World J 2008;8:901–904.
Wang B, Herman-Edelstein M, Koh P et al. E-cadherin expression is regulated by miR-192/215 by a mechanism that is independent of the profibrotic effects of transforming growth factor-beta. Diabetes 2010;59:1794–1802.
Bolos V, Grego-Bessa J, de la Pompa JL . Notch signaling in development and cancer. Endocr Rev 2007;28:339–363.
Jiang YG, Luo Y, He DL et al. Role of Wnt/beta-catenin signaling pathway in epithelial-mesenchymal transition of human prostate cancer induced by hypoxia-inducible factor-1alpha. Int J Urol 2007;14:1034–1039.
Larue L, Bellacosa A . Epithelial-mesenchymal transition in development and cancer: role of phosphatidylinositol 3' kinase/AKT pathways. Oncogene 2005;24:7443–7454.
Moustakas A, Heldin CH . Signaling networks guiding epithelial-mesenchymal transitions during embryogenesis and cancer progression. Cancer Sci 2007;98:1512–1520.
Acknowledgements
This study was supported by a faculty research grant of Yonsei University College of Medicine for 2011(6-2011-0208) and by a grant of the Korean Health 21R&D Project, Ministry of Health and Welfare, Republic of Korea (A111218- CP01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Park, M., Kim, M., Hwang, D. et al. Characterization of gene expression and activated signaling pathways in solid-pseudopapillary neoplasm of pancreas. Mod Pathol 27, 580–593 (2014). https://doi.org/10.1038/modpathol.2013.154
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.154
Keywords
This article is cited by
-
Solid Pseudopapillary Neoplasm of the Pancreas
Journal of Gastrointestinal Surgery (2021)
-
Clinicopathological and immunohistochemical study of 29 cases of solid-pseudopapillary neoplasms of the pancreas in patients under 20 years of age along with detailed review of literature
Diagnostic Pathology (2020)
-
PDAC-ANN: an artificial neural network to predict pancreatic ductal adenocarcinoma based on gene expression
BMC Cancer (2020)
-
Molecular Diagnosis of Cystic Neoplasms of the Pancreas: a Review
Journal of Gastrointestinal Surgery (2020)
-
Solid-pseudopapillary Neoplasms of the Pancreas is still an Enigma: a Clinicopathological Review
Pathology & Oncology Research (2020)