Abstract
Most familial breast cancers arise in patients who tested negative for germline mutations in BRCA1 and BRCA2 genes (also referred to as BRCAX cases). Several studies aimed to define histopathological and molecular profiles characteristic of BRCA1, BRCA2 and BRCAX tumors have been performed. Major pathological and immunohistochemical differences have been reported in BRCA1 cancers compared to the other two groups, whereas less difference has been observed between BRCA2 and BRCAX cases. The aim of this study was to investigate the ability of selected tumor markers to discriminate BRCAX breast cancers from cancers arising in carriers of mutations in BRCA genes, and their usefulness in selecting familial cases in whom testing for such mutations is more likely to result uninformative. We carried out a morphological and immunohistochemical analysis on 22 BRCA1, 16 BRCA2 and 33 BRCAX familial breast cancers. Age at first diagnosis, histological type and grade, and immunostaining for estrogen receptor (ER), progesterone receptor (PR), p53, HER2/Neu, E-cadherin and cyclin D1 were investigated. The occurrence of somatic mutations of the TP53 gene was also verified. BRCA1 tumors resulted clearly distinguishable from BRCAX cases, occurring at a younger age, being more frequently of higher grade, negative for ER, PR and cyclin D1 expression and positive for p53 alterations. The predictive value of age at diagnosis, histological grade and PR expression was confirmed in a multivariable analysis. When comparing BRCA2 with BRCAX tumors, the only parameter that differed was cyclin D1, which was significantly overexpressed in BRCA2 cases both in the univariable and the multivariable analyses. If confirmed by further studies, our observations indicate that the investigation of cyclin D1 expression in familial breast cancer cases could be used, in conjunction with the analysis of other tumor markers preferentially associated with BRCA1 or BRCA2 tumors, to prioritize hereditary cases for mutation testing in BRCA genes.
Similar content being viewed by others
Introduction
Breast cancer is the most common malignancy among women in developed countries. It is estimated that approximately 5–10% of all breast cancers are hereditary and occur in women with a positive family history for the disease. The tumor suppressor BRCA11 and BRCA22 are the genes most commonly implicated at germline level in hereditary breast cancers.
Testing for BRCA1 and BRCA2 alterations within high-risk families is a time consuming and expensive procedure because of the large size of both genes and the heterogeneity of mutation spectra. Moreover, mutations in BRCA genes account for no more than 25% of all familial aggregations of breast cancer.3 To provide a tool for accurate selection of individuals to address for genetic testing, several statistical models for the prediction of mutations in BRCA genes have been developed. However, these methods did not prove to be sufficiently efficient to be used in a clinical setting.4 On the other hand, different studies have shown that the phenotypic and molecular features of breast cancers arising in BRCA1 and BRCA2 mutation carriers could be used to identify, among familial cases, those who are more likely to be associated with these genes.
BRCA1-related tumors tend to be more frequently of ductal type with an excess of medullary and atypical medullary cases, and of higher histological grade than BRCA2 and sporadic cancers.5, 6, 7 In addition, breast carcinomas arising in BRCA1 mutation carriers are more frequently estrogen receptor (ER) and progesterone receptor (PR) negative and p53 positive compared to sporadic age-matched controls.6, 8, 9, 10, 11 Moreover, they show positivity for a few markers, including cytokeratin 5/6, cytokeratin 8/18, EGFR, and vimentin, which define the so-called ‘basal-like’ phenotype.11, 12, 13
Identifying a phenotype that characterizes breast cancers associated with BRCA2 mutations is more elusive. In general, BRCA2-related tumors do not show a specific phenotype. They tend to be of higher grade than sporadic cancers, although to a lesser extent compared to BRCA1-related cases, but are not significantly different from sporadic controls with respect to ER and PR positivity and p53 alterations.6, 8, 9, 10, 11 In addition, both BRCA1- and BRCA2-associated breast cancers are prevalently negative for HER2/Neu expression.6, 8, 9, 11, 14
Only a few studies have been performed on the morphological and immunophenotypic profiles of familial breast tumors in patients negative for mutations in BRCA genes, also termed BRCAX, which appear to be a rather heterogeneous group.15 They have been shown to be of lower grade and more frequently of lobular type than breast cancers arising in BRCA1 mutation carriers,14, 16 although the latter finding has been disputed by another study.17 The BRCAX tumors are also of significantly lower grade in comparison to BRCA2 tumors and with control breast cancers unselected for family history of the disease, even if these differences are less pronounced in comparison to BRCA1.14, 16 In addition, they are more frequently ER and PR positive and p53 negative than BRCA1 tumors, but no statistically significant difference is generally found in the expression of HER2/Neu and E-cadherin.14, 17, 18, 19 One study reported that when compared to BRCA2, BRCAX tumors show a less frequently preserved expression of E-cadherin and a lower frequency of HER2/Neu positivity.14 However, the latter finding was not confirmed by subsequent reports.17, 19, 20 A higher frequency of breast cancers overexpressing cyclin D1 was recently reported in BRCAX patients compared to both BRCA1 and BRCA2 cases.17 This finding is in keeping with the results of a previous study that reported a lower frequency of cyclin D1-positive tumors among BRCA gene mutation carriers aged <40 years compared to age-matched mutation-negative patients.8 However, no such difference was observed in another study comparing hereditary breast cancer associated with germline BRCA2 mutation and control tumors unlinked to BRCA1 and BRCA2 ascertained through a breast cancer family registry.20
Overall, the major histopathological and molecular differences are observed in cancers arising in BRCA1 mutation carriers compared to all other groups, whereas few differences are apparent among BRCA2, BRCAX and sporadic breast cancers.
The aims of this study were (a) to investigate the ability of selected tumor markers to discriminate, when used in combination, BRCAX from both BRCA1 and BRCA2 breast cancers, and (b) to verify their predictivity in selecting families in whom testing for BRCA gene mutations is more likely to result uninformative and, therefore, of no benefit.
To this end, we carried out a morphological and immunohistochemical analysis on 71 hereditary breast cancers (22 BRCA1, 16 BRCA2 and 33 BRCAX) that were tested for a set of different markers, including ER, PR, p53, HER2/Neu, E-cadherin, and cyclin D1. The occurrence of somatic mutations of the TP53 gene was also investigated.
Materials and methods
Patients
A total of 71 breast cancer tissue samples from an equal number of female patients were included in the study. Information regarding age at diagnosis, laterality and personal cancer history was acquired from clinical reports. Enrolled patients belonged to families with a history of breast and/or ovarian cancer complying with the intake criteria to BRCA testing in use at the Istituto Nazionale Tumori (INT) of Milan, as previously reported.21 All subjects received genetic counseling and provided a written consent to BRCA mutation testing for the use of their biological material for research purposes, approved by the local ethical committee. Thirty-eight patients carried frankly deleterious alterations (including nonsense and frameshift mutations or exon deletion) in either BRCA1 (22 subjects form 20 families) or BRCA2 (16 subjects from 13 families). The remaining 33 patients (all from different families) tested negative for mutations in both genes (BRCAX) after screening of all coding exons and splice sites by both direct sequencing and multiplex ligation-dependant probe amplification. Overall, 57 primary tumors, 6 metachronous contralateral tumors and 8 ipsilateral tumors (with a time interval form first event ranging from 4 to 13 years) were collected from the Department of Pathology of INT in the form of Bouin's fluid- or formalin-fixed, paraffin-embedded blocks. All tumors were invasive carcinomas and were classified for type and grading by reviewing hematoxylin–eosin slides by the pathologist, blinded to BRCA1 and BRCA2 germline mutational status.
Immunohistochemistry analyses
Immunostaining for ER, PR, p53, HER2/Neu, E-cadherin and cyclin D1 was performed on 2-μm cut tumor sections using the streptavidin–biotin immunoperoxidase method (1:300 in PBS; Dako, Carpinteria, CA, USA) in accordance with the manufacturer's instructions. Staining was developed using 3,3′-diaminobenzidine (Sigma, St Louis, MO, USA). Antibodies, dilutions and suppliers are listed in Table 1. Antigen retrieval was performed in 10 mM citrate buffer (pH 6.0) at 95°C for 6 min except for cyclin D1, for witch a treatment with 1 mM EDTA buffer (pH 8.0) at 95°C for 15 min was employed. For ER and PR, a sample was classified as positive when the percentage of stained nuclei was ≥10% while for cyclin D1 and p53 cutoff values of 20 and 50%, respectively, were considered. For HER2/Neu and E-cadherin, membranous staining was considered.
DNA extraction
Areas of tumor tissue were chosen from formalin-fixed paraffin blocks using the corresponding hematoxylin- and eosin-stained slides as a guide. For each sample, six consecutive 8-μm-thick sections were cut and used as such, if only tumor tissue was present, or counterstained with methylene blue and microdissected under light microscope. The sections were deparaffinated in xylene and rehydrated through graded alcohols. For samples fixed in Bouin's fluid, after xylene treatment the sections were washed in a solution containing 60% ethanol and 40 mM NaOH. The tumor tissues were then digested with proteinase K and DNA was purified using QIAamp® DNA Mini kit, according to the manufacturer's instructions (Qiagen, Hilden, Germany), and resuspended in 200 μl elution buffer.
TP53 mutation analysis
All tumor DNAs were screened in exons 4–9 of the TP53 gene. In samples that tested mutation negative, the analysis was extended to the remaining coding exons. Each time a mutation was identified in the tumor DNA, its presence at the constitutional level was verified in the corresponding blood DNA. Polymerase chain reaction (PCR) analysis was carried out using 3 μl DNA solution and 1 U of AmpliTaq Gold DNA polymerase (Applied Biosystems) in a finale volume of 20 μl. For each sample, two sequential rounds of amplification, consisting of 35 cycles each, were performed, as described elsewhere, with minor modifications.22 Primer sequences are available upon request. PCR products were analyzed on 3% agarose gel stained with ethidium bromide and purified directly from reaction solutions using the QIAquick PCR Purification kit (Qiagen) or after excision from gel if nonspecific amplified DNA was present, using the QIAquick Gel Extraction kit (Qiagen, QIAamp DNA Mini kit). Purified DNA was sequenced in both directions with nested primers using the ABI PRISM® BigDyeTM Terminator Cycle Sequencing kit (Applied Biosystems) and examined on an ABI PRISM 377 DNA Sequencer (Applied Biosystems) using the Sequencing Analysis software (Applied Biosystems). Mutations were verified by performing second independent PCR and sequence analysis to exclude possible artifacts.
Statistical methods
Tumors in BRCA1 and BRCA2 mutation carriers were separately compared with tumors in BRCAX cases. Contingency tables were created to describe the distribution of clinical, pathological and molecular features of breast cancer lesions among BRCA1, BRCA2 and BRCAX cases, and univariable associations were tested with the Fisher exact test. To determine which factors were independently predictive of genetic status, we also performed multivariable analyses by means of logistic regression models, which summarize the effect of each factor in terms of odds ratios (ORs). Results from logistic models are shown by reporting the estimated ORs, together with corresponding standard errors and P-values. An OR greater (lower) than 1 indicates an increased (decreased) probability of BRCA1 or BRCA2 mutation in a given category compared to the reference category of the predictive factor. As a measure of model predictive capability, the Harrell's c-statistic was calculated. A c-statistic of 1 indicates perfect predictive capability, a value of 0.5 indicates the lack of predictive capability and values in between indicate from poor to strong predictive capability. Two-sided P-values below 0.05 were considered significant. Computations were made by use of the SAS™ software.
Results
Clinical and morphological features
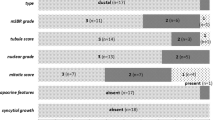
The distributions of ages at disease onset (first event) and of pathological characteristics of the cases included in the study are reported in Table 2.
BRCA1 but not BRCA2 patients were diagnosed with breast cancer at a significantly younger age than BRCAX cases (P=0.0110).
As for the histological types, a significant difference (P=0.0214) was observed between BRCAX and BRCA1 cases, which showed an excess of medullary carcinomas, but not between BRCAX and BRCA2 cases.
BRCA1 tumors were of higher histological grade than BRCAX cases (P<0.0001), whereas BRCA2 and BRCAX cases showed a similar grade distribution.
Immunohistochemistry markers
Immunostaining for p53 was successfully performed on all 71 specimens considered. For the other markers, a few cases were excluded because of the low quality of the staining or because the available tumor material was not sufficient (Table 3).
When compared to BRCAX cases, BRCA1 tumors were observed to be more frequently ER negative (100 vs 30%; P<0.0001), PR negative (90 vs 36%; P<0.0001), p53 positive (45 vs 6%; P=0.0008) and cyclin D1 negative (95 vs 64%; P=0.0186). No meaningful difference was observed for the above markers between BRCA2 and BRCAX cases, with the exception of cyclin D1, which showed a significantly higher frequency of staining in BRCA2 tumors (73 vs 36%; P=0.0285).
A strong correlation was observed between the expression of cyclin D1 and that of ER in the BRCA2 group, in which 10 of the 11 cyclin D1-positive tumors (91%) were also ER positive. Conversely, in the BRCAX group only 8 of 12 cyclin D1-positive tumors (67%) tested positive also for ER.
The expression of HER2/Neu and E-cadherin did not significantly differ in the three groups analyzed, being HER2/Neu negative and E-cadherin positive in the large majority (≥90%) of cases examined.
TP53 mutations
Positive nuclear staining for p53 protein in tumor cells is considered a diagnostic of altered gene function. However, not all TP53 mutations are detectable at the immunohistochemistry level.23, 24 To obtain a more accurate estimate of the frequency of TP53 alterations in our group of hereditary breast cancers, a DNA sequencing analysis was performed in a subset of cases (n=56, including 21 BRCA1, 13 BRCA2 and 22 BRCAX tumors). For the remaining 15 tumors, the quality of obtained DNA was not adequate to mutation analyses.
TP53 was mutated in 52% (11/21) of BRCA1 tumors vs 15% (2/13) of BRCA2 and 9% (2/22) of BRCAX tumors. Identified mutations are reported in Table 4.
In the BRCA1 group, nine mutations were missense and two were frameshift deletions (c.599delA and c.631delA), both introducing a premature truncation site at residue 246. In the BRCA2 group, the same missense mutation at codon 220 (p.Tyr>Cys) was detected in two unrelated tumors. Finally, the two alterations detected in the BRCAX group were a missense mutation at codon 248 (p.Arg>Trp) and a nonsense mutation at codon 342. In addition, silent mutations were detected at codon 88 in a BRCA1 case and at codons 290 and 299 in a BRCA2 case. None of the identified mutations were detected at the germline level.
According to the TP53 website (http://p53.free.fr/Database/p53_database.html; 2007 version), all but three of the alterations detected in this analysis have been already reported in breast cancer. The exceptions, all in BRCA1 tumors, were the frameshift deletion at codon 200 and the missense mutation at codon 92, which were previously observed only in tumors other than breast carcinoma, and the frameshift deletion at codon 211, which was not present in the database. In addition, the majority of the other TP53 mutations found in BRCA1 tumors were rarely represented (<1% of all entries) in the database.
When compared with immunohistochemistry analyses, discrepant results were observed in 12 tumors (Table 5). In particular, five cases (B2, B15, B37, B56 and B75) showed p53 stabilization in the absence of detectable gene mutations, strongly suggesting the presence of an alternative mechanism of deregulation of the protein, while seven tumors with mutated TP53 tested negative at immunostaining. These included three cases (B11, B16 and B80) with mutations producing truncated, and possibly unstable, proteins and four cases (B1, B3, B22 and B34) with missense mutations (Table 4). Noticeably, three of the latter tumors carried the same alteration (p.Y220C).
Taken together, the molecular and immunohistochemistry analyses showed that 14/22 (64%) BRCA1 and 3/16 (19%) BRCA2 breast cancers carried p53 alterations compared to 3/33 (9%) BRCAX cases (P<0.0001 and P=0.3773, respectively) (Table 5).
These figures might underestimate the actual proportions of tumors with altered p53 in each group, since, as reported above, the mutational screening could not be performed in 15 cases, all immunohistochemistry negative. However, considering that TP53 mutations were detected in only 7 of 43 immunohistochemistry-negative tumors tested at the DNA level (16%), we assumed that the introduced biases were unlikely to affect the results of the statistical analyses.
Multivariable analyses
A multivariable analysis was carried out on the factors that appeared to be predictive of BRCA1 or BRCA2 status in the univariable analyses, including age, histological grade, PR, p53 and cyclin D1 (Table 6). The effect of ER was not estimable, being all BRCA1 carriers negative for this parameter.
In regards to BRCA1, only histological grade and PR were statistically significant in the multivariable analysis, after adjusting for the remaining factors, whereas early age of disease onset (≤35 years) was associated with a borderline significance (P=0.0675). The model c-statistic for all considered variables was 0.928, quite a high figure denoting a strong model capability to predict BRCA1 status.
In regards to BRCA2, the only significant factor that emerged from the analysis was cyclin D1. In accordance, the overall c-statistic (0.748) was only slightly superior to that achievable considering cyclin D1 only (0.685). This means that cyclin D1 was the most important predictor, and that the remaining factors included in the multivariable analysis added little information in terms of predictive power.
A complementary quantification of the capability of cyclin D1 to predict BRCA2 status was achieved by calculating the positive and negative predictive values, namely the probability of a BRCA2 mutation when cyclin D1 is positive or negative, respectively. For this calculation, we considered relative prevalence figures of BRCA1, BRCA2 and BRCAX cases of 14, 9 and 77%, respectively, observed among 700 familial breast cancer patients fulfilling the same intake criteria for genetic testing that were used to select the patients enrolled in the present study (unpublished data). The results obtained indicated a positive predictive value of 19% and a negative predictive value of 95%.
Discussion
The results of our combined morphological, immunohistochemical and molecular analyses on a panel of 71 hereditary breast cancers were largely in keeping with previously reported findings.25
In particular, we confirmed that overall BRCA1-associated tumors are clearly distinguishable from BRCAX cases, occurring at a younger age, being more frequently of higher grade, ER and PR negative and p53 positive by immunohistochemistry. Moreover, consistently with a previous study,18 we observed a low rate of TP53 gene mutations among BRCAX cases. Such a rate was comparable to that detected in BRCA2 tumors and significantly lower than in BRCA1 tumors. In the latter group, we also found that most identified mutations had never or only rarely been reported in human cancers, in keeping with the notion that TP53 mutation spectrum in BRCA1-associated tumors is highly heterogeneous,26 possibly due to a reduced efficiency of DNA repair activity in BRCA1-deficient cells. The predictive value of age at diagnosis, histological grade and PR expression was confirmed by the multivariable analysis, in which a c-statistic value of 0.928 was obtained, quite a high figure denoting a strong model capability to predict BRCA1 status.
In our study, no difference was observed among the three groups considered for the expression of the HER2/Neu and the E-cadherin proteins. On the contrary, the BRCAX tumors were found to differ form both BRCA1 and BRCA2 cases for the expression of cyclin D1, a protein involved in the regulation of the G1–S phase transition, which plays an important role in mammary carcinogenesis.27, 28 In particular, BRCAX cases exhibit a significantly higher frequency of positivity compared to BRCA1-associated tumors, the majority of which tested negative for this marker, in agreement with previous reports.8, 11, 17 On the other hand, BRCAX tumors were significantly less positive than BRCA2 cases. Indeed, in our study, cyclin D1 was the only parameter that differentiated these two groups of familial breast cancers. Such difference was not observed in previous studies in which, however, the frequency of cyclin D1-positive cases among BRCA2 tumors ranged from 12 to 56%8, 17, 20 and, therefore, was lower than that we detected in the same group (75%).27, 28 Apart from sampling biases, such discrepancy could be due to technical reasons related to the immunostaining procedures, in particular to antigen retrieval, and/or to differences in cutoff values adopted to score a sample as positive.
The preferential association of cyclin D1 positivity with BRCA2 tumors was confirmed by multivariable analysis. In addition, the computed c-statistic was 0.748, a figure denoting a model predictivity weaker than that achievable for BRCA1 (0.928), but that was almost entirely attributable to the contribution of cyclin D1. Moreover, cyclin D1 was associated with a remarkably high negative predictive value (95%) for BRCA2 mutations.
Another interesting finding of our study was the observation that BRCAX tumors did not exhibit the strong correlation in the expression of ER and cyclin D1 detected in sporadic breast cancers, where the great majority of cyclin D1-positive cases (>90%) are also positive for ER.29, 30, 31, 32 Such correlation is in keeping with the notion that the expression of cyclin D1 is induced by estrogens and is, therefore, dependent on the presence of ER. On the other hand, cyclin D1 has been demonstrated to trigger the transactivation properties of ER even in the absence of its ligand,33, 34 a property that might account for its oncogenic activity in ER-positive breast cancers.28 Consistently, of the 20 BRCA1 tumors, all ER negative, that were analyzed for cyclin D1 expression, 19 (95%) tested negative, whereas 10 of 11 (91%) cyclin D1-positive BRCA2 tumors were also ER positive. However, among BRCAX tumors, of the 12 cases scored positive for cyclin D1, only 8 (67%) expressed ER. This suggests that at least a fraction of hereditary breast cancer unlinked to BRCA1 and BRCA2 might develop through mechanisms that are specific or preferentially associated with this group of malignancies and distinguish them from other breast cancers.
Other than providing clues on the specific molecular mechanisms involved in the development of familial breast cancers differentially associated with BRCA genes, our observations suggest that the analysis of cyclin D1 expression might be useful in discriminating among the various groups of familial breast cancers and, in particular, between BRCA2 and BRCAX cases. Owing to the paucity of tumor markers reported to date as preferentially associated with BRCA2-linked breast cancers, these findings are of particular interest in the management of familial cases, since they may be of help for the identification of patients more likely to carry mutations in this gene. Under this respect, cyclin D1 might be used in combination with other markers that have been recently reported as equally able to differentiate BRCA2-associated breast cancers, including the DNA repair proteins CHEK2 and RAD1.35
References
Miki Y, Swensen J, Shattuck-Eidens D, et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994;266:66–71.
Wooster R, Bignell G, Lancaster J, et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 1995;378:789–792.
Antoniou AC, Easton DF . Models of genetic susceptibility to breast cancer. Oncogene 2006;25:5898–5905.
Kang HH, Williams R, Leary J, et al. Evaluation of models to predict BRCA germline mutations. Br J Cancer 2006;95:914–920.
Armes JE, Egan AJ, Southey MC, et al. The histologic phenotypes of breast carcinoma occurring before age 40 years in women with and without BRCA1 or BRCA2 germline mutations: a population-based study. Cancer 1998;83:2335–2345.
Chappuis PO, Nethercot V, Foulkes WD . Clinico-pathological characteristics of BRCA1- and BRCA2-related breast cancer. Semin Surg Oncol 2000;18:287–295.
Breast Cancer Linkage Consortium. Pathology of familial breast cancer: differences between breast cancers in carriers of BRCA1 or BRCA2 mutations and sporadic cases. Lancet 1997;349:1505–1510.
Armes JE, Trute L, White D, et al. Distinct molecular pathogeneses of early-onset breast cancers in BRCA1 and BRCA2 mutation carriers: a population-based study. Cancer Res 1999;59:2011–2017.
Lakhani SR, Van De Vijver MJ, Jacquemier J, et al. The pathology of familial breast cancer: predictive value of immunohistochemical markers estrogen receptor, progesterone receptor, HER-2, and p53 in patients with mutations in BRCA1 and BRCA2. J Clin Oncol 2002;20:2310–2318.
Foulkes WD, Metcalfe K, Sun P, et al. Estrogen receptor status in BRCA1- and BRCA2-related breast cancer: the influence of age, grade, and histological type. Clin Cancer Res 2004;10:2029–2034.
Palacios J, Honrado E, Osorio A, et al. Phenotypic characterization of BRCA1 and BRCA2 tumors based in a tissue microarray study with 37 immunohistochemical markers. Breast Cancer Res Treat 2005;90:5–14.
Foulkes WD, Stefansson IM, Chappuis PO, et al. Germline BRCA1 mutations and a basal epithelial phenotype in breast cancer. J Natl Cancer Inst 2003;95:1482–1485.
Lakhani SR, Reis-Filho JS, Fulford L, et al. Prediction of BRCA1 status in patients with breast cancer using estrogen receptor and basal phenotype. Clin Cancer Res 2005;11:5175–5180.
Palacios J, Honrado E, Osorio A, et al. Immunohistochemical characteristics defined by tissue microarray of hereditary breast cancer not attributable to BRCA1 or BRCA2 mutations: differences from breast carcinomas arising in BRCA1 and BRCA2 mutation carriers. Clin Cancer Res 2003;9:3606–3614.
Honrado E, Osorio A, Milne RL, et al. Immunohistochemical classification of non-BRCA1/2 tumors identifies different groups that demonstrate the heterogeneity of BRCAX families. Mod Pathol 2007;20:1298–1306.
Lakhani SR, Gusterson BA, Jacquemier J, et al. The pathology of familial breast cancer: histological features of cancers in families not attributable to mutations in BRCA1 or BRCA2. Clin Cancer Res 2000;6:782–789.
Oldenburg RA, Kroeze-Jansema K, Meijers-Heijboer H, et al. Characterization of familial non-BRCA1/2 breast tumors by loss of heterozygosity and immunophenotyping. Clin Cancer Res 2006;12:1693–1700.
Sensi E, Tancredi M, Aretini P, et al. p53 inactivation is a rare event in familial breast tumors negative for BRCA1 and BRCA2 mutations. Breast Cancer Res Treat 2003;82:1–9.
Eerola H, Heikkila P, Tamminen A, et al. Relationship of patients' age to histopathological features of breast tumours in BRCA1 and BRCA2 and mutation-negative breast cancer families. Breast Cancer Res 2005;7:R465–R469.
Bane AL, Beck JC, Bleiweiss I, et al. BRCA2 mutation-associated breast cancers exhibit a distinguishing phenotype based on morphology and molecular profiles from tissue microarrays. Am J Surg Pathol 2007;31:121–128.
Manoukian S, Peissel B, Pensotti V, et al. Germline mutations of TP53 and BRCA2 genes in breast cancer/sarcoma families. Eur J Cancer 2007;43:601–606.
Lehman TA, Bennett WP, Metcalf RA, et al. p53 mutations, ras mutations, and p53-heat shock 70 protein complexes in human lung carcinoma cell lines. Cancer Res 1991;51:4090–4096.
Wynford-Thomas D . P53 in tumour pathology: can we trust immunocytochemistry? J Pathol 1992;166:329–330.
Casey G, Lopez ME, Ramos JC, et al. DNA sequence analysis of exons 2 through 11 and immunohistochemical staining are required to detect all known p53 alterations in human malignancies. Oncogene 1996;13:1971–1981.
Honrado E, Benitez J, Palacios J . Histopathology of BRCA1- and BRCA2-associated breast cancer. Crit Rev Oncol Hematol 2006;59:27–39.
Greenblatt MS, Chappuis PO, Bond JP, et al. TP53 mutations in breast cancer associated with BRCA1 or BRCA2 germ-line mutations: distinctive spectrum and structural distribution. Cancer Res 2001;61:4092–4097.
Arnold A, Papanikolaou A . Cyclin D1 in breast cancer pathogenesis. J Clin Oncol 2005;23:4215–4224.
Roy PG, Thompson AM . Cyclin D1 and breast cancer. Breast 2006;15:718–727.
Michalides R, Hageman P, van Tinteren H, et al. A clinicopathological study on overexpression of cyclin D1 and of p53 in a series of 248 patients with operable breast cancer. Br J Cancer 1996;73:728–734.
Jares P, Rey MJ, Fernandez PL, et al. Cyclin D1 and retinoblastoma gene expression in human breast carcinoma: correlation with tumour proliferation and oestrogen receptor status. J Pathol 1997;182:160–166.
Zukerberg LR, Yang WI, Gadd M, et al. Cyclin D1 (PRAD1) protein expression in breast cancer: approximately one-third of infiltrating mammary carcinomas show overexpression of the cyclin D1 oncogene. Mod Pathol 1995;8:560–567.
Barbareschi M, Pelosio P, Caffo O, et al. Cyclin-D1-gene amplification and expression in breast carcinoma: relation with clinicopathologic characteristics and with retinoblastoma gene product, p53 and p21WAF1 immunohistochemical expression. Int J Cancer 1997;74:171–174.
Neuman E, Ladha MH, Lin N, et al. Cyclin D1 stimulation of estrogen receptor transcriptional activity independent of cdk4. Mol Cell Biol 1997;17:5338–5347.
Zwijsen RM, Wientjens E, Klompmaker R, et al. CDK-independent activation of estrogen receptor by cyclin D1. Cell 1997;88:405–415.
Honrado E, Osorio A, Palacios J, et al. Immunohistochemical expression of DNA repair proteins in familial breast cancer differentiate BRCA2-associated tumors. J Clin Oncol 2005;23:7503–7511.
Acknowledgements
This work was partially supported by grants from the Italian Foundation for Cancer Research (FIRC, Special Project ‘Hereditary Tumors’).
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Colombo, M., Giarola, M., Mariani, L. et al. Cyclin D1 expression analysis in familial breast cancers may discriminate BRCAX from BRCA2-linked cases. Mod Pathol 21, 1262–1270 (2008). https://doi.org/10.1038/modpathol.2008.43
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.43
Keywords
This article is cited by
-
First description of an acinic cell carcinoma of the breast in a BRCA1 mutation carrier: a case report
BMC Cancer (2013)
-
Characterisation of amplification patterns and target genes at chromosome 11q13 in CCND1-amplified sporadic and familial breast tumours
Breast Cancer Research and Treatment (2012)
-
Subtypes of familial breast tumours revealed by expression and copy number profiling
Breast Cancer Research and Treatment (2010)