Abstract
Human immunoglobulin A (IgA) comprises two IgA subclasses, IgA1 and IgA2, whose distribution has been shown by immunohistochemistry to be different in various body compartments. In comparison with systemic immune compartments, we investigated the IgA switch profiles at the molecular level in salivary and lacrimal glands, nasal mucosa, and proximal and distal gut mucosa. Direct switching from IgM to IgA1 or IgA2 predominated in all immune compartments analyzed. Similar composition of the Sμ–Sα1 and Sμ–Sα2 junctions was observed, including microhomology usage, which suggested that there is no major difference in the actual recombination mechanism utilized during IgA subclass switching. The proportion of IgA1/IgA2 switch recombination events largely paralleled the previously published immunohistochemical representation of IgA1+ and IgA2+ plasma cells, implying that the local subclass distribution generally reflects precommitted memory/effector B cells that have undergone IgA subclass switching before extravasation at the effector site. The extremely low or undetectable levels of activation-induced cytidine deaminase (AID) and Iα–Cμ circle transcripts in intestinal lamina propria samples as compared with Peyer’s patches suggest that the cellular IgA subclass distribution outside of organized gut-associated lymphoid tissue is only to a minor extent, if at all, influenced by in situ switching.
Similar content being viewed by others
INTRODUCTION
The most abundant antibody isotype in humans is immunoglobulin A (IgA). It comprises two subclasses, IgA1 and IgA2, of which the constant (C) heavy-chain regions are encoded by separate genes (Cα1 and Cα2) on chromosome 14.1 IgA2 is relatively resistant to enzymatic degradation because of a 13-amino-acid deletion in the hinge region that removes the recognition site for IgA1-specific bacterial proteases.2, 3 Immunochemical analyses have shown that the distribution of the two IgA subclasses is strikingly different in various immune compartments. IgA1, in a monomeric form, predominates (∼90%) in serum, whereas a larger proportion of IgA2, mainly as dimers, is seen in various external secretions,4, 5, 6 which may be advantageous because of its stability.
The mucosal immune system consists of two functionally distinct tissue compartments: inductive sites, where antigens from mucosal surfaces activate naive T and B lymphocytes; and effector sites, where memory/effector B cells after extravasation undergo terminal differentiation to plasma cells. Mucosa-associated inductive sites chiefly consist of palatine tonsils, adenoids, Peyer’s patches (located mainly in the distal ileum), and isolated lymphoid follicles (ILFs)—most abundantly in the large bowel.7 Peyer’s patches, ILFs, and the appendix (with its large follicles) are collectively referred to as gut-associated lymphoid tissue (GALT). Mucosal immune effector sites comprise the subepithelial lamina propria of various mucosae, stroma of exocrine glands, and the surface epithelia.7, 8, 9 The gut mucosal system harbors by far the largest activated B-cell pool in the body, containing at least 80% of all plasmablasts and plasma cells, mostly producing IgA dimers.10 Immunohistochemical studies of these cells showed a striking disparity in local subclass distribution, with IgA2 production dominating distally in the gut.4, 5 It remains unknown whether such regional disparity is governed by inherent characteristics of the activated B cells derived by homing through lymph and peripheral blood from different inductive mucosa-associated lymphoid compartments or subsequently being tailored in situ by immunoregulatory local factors at the effector sites.9, 11, 12, 13 It was indeed suggested quite early that the topical intestinal microbiota might be involved in the distal gut preference for IgA2.14
Isotype switching, or class switch recombination (CSR), is initiated by activation-induced cytidine deaminase (AID) in activated B cells15 and involves specific DNA regions, referred to as switch (S) regions located upstream of the genes encoding the Ig heavy-chain C regions.16, 17 DNA double-strand breaks are introduced in the donor and acceptor S regions and these breaks are processed by a number of DNA repair pathways, eventually leading to recombination of the two S regions.18 This allows previously rearranged Ig heavy-chain V domains to be expressed in association with a C region different from Cμ, thus promoting B-cell switching to IgA, IgG, or IgE expression. CSR is also accompanied by deletion of circular DNA from the Ig heavy-chain locus. These so-called “switch circles” contain the DNA segments between the Sμ and a target S region, including the I (intervening) region promoter. Under the influence of the I promoter, the switch circle produces an I–Cμ transcript, referred to as circle transcript (Figure 1a), which, together with AID transcripts, have been suggested to serve as hallmarks of ongoing or recent CSR.19
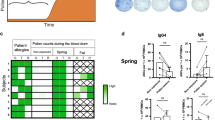
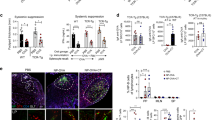
Nested PCR amplification of Sμ–Sα fragments generated by class switch recombination (CSR) in systemic and mucosal immune compartments. (a) Schematic map of the human Ig heavy-chain constant (C) locus. Horizontal small arrows indicate the approximate positions of the primers used for amplification of Iα1–Cα1 germline transcripts, Sμ–Sα fragments, and Iα–Cμ circle transcripts. (b) Typical gel electrophoresis of PCR products as determined with genomic DNA in one representative tissue sample of spleen, ileum, and colon. For each specimen, the numbers of Sμ–Sα switch fragments were based on 10 PCR reactions run in parallel (lanes 1–10). M, molecular weight marker.
The mechanisms of CSR are partly mediated by cytokines and B-cell activators that regulate transcription of unrearranged CH genes, yielding the so-called germline (GL) transcripts (Figure 1a). These transcripts are a prerequisite for CSR and their regulation is a decisive mechanism for isotype selection.20 Thus, transforming growth factor-β1 is an important cytokine for directing the IgA switching through regulation of Iα GL transcripts, both in human and mouse B cells.21, 22 However, as the transforming growth factor-β1-responsive elements in the human Iα1 and Iα2 promoters are identical, this cytokine probably does not provide subclass-specific targeting.17 More recently, APRIL (A Proliferation-Inducing Ligand) has been proposed as the most important cytokine directing IgA2 switching, whereas BAFF (B-cell-Activating Factor belonging to the TNF Family), in addition to its ability to induce IgA1 switching, can enhance APRIL-stimulated IgA2 expression, apparently by promoting the survival of IgA2+ B cells.23
In this study, the differential induction of IgA1 and IgA2 revealed in various human tissues was investigated by molecular methods. We compared the IgA subclass switch patterns detected at mucosal effector and inductive sites, as well as in systemic immune compartments, by analyzing the composition of Sμ–Sα recombination junctions. Furthermore, in order to explore the mechanisms underlying the differential regulation of IgA subclass switching, we analyzed the level of GL Iα1, GL Iα2, APRIL, and BAFF transcripts in specimens obtained from the proximal and distal gut mucosa as well as in Peyer’s patches. In addition, the expression of Iα–Cμ circle transcripts and AID was investigated in these samples to investigate possible in situ CSR activities outside the Peyer’s patches.
RESULTS
Profiles of Sμ–Sα1/Sμ–Sα2 B-cell switching in systemic and mucosal immune compartments
Genomic DNA purified from peripheral blood lymphocytes (PBLs), spleen, and various mucosal tissues was subjected to nested PCR to amplify the Sμ–Sα switch fragments that contain the recombination junctions. The approximate positions of the PCR primers are shown in Figure 1a. The number of Sμ–Sα fragments was determined from 10 reactions with DNA from each sample and represents random amplification of clones that had switched in vivo. As shown in Figure 1b, the number of Sμ–Sα switch fragments generated from one representative spleen sample ranged from 10 to 15, each with a unique length. The number and intensity of amplified Sμ–Sα fragments in mucosal tissues (represented by one ileal and one colonic sample in Figure 1b) were comparable to those in spleen, confirming that there was a sizable IgA-committed B-cell pool in the different mucosal compartments, regardless of anatomical site.
To further characterize the Sμ–Sα junctions obtained from systemic and mucosal tissue samples, we subsequently cloned and sequenced the amplified Sμ–Sα fragments. Altogether, 322 unique Sμ–Sα sequences were obtained from various tissues (Table 1) and (Supplementary Figure S1 online). In addition, 159 Sμ–Sα fragments from PBLs of adults and 141 fragments from PBLs of children, which have previously been partially described,24, 25, 26, 27 were also included for comparison.
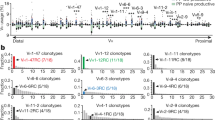
All sequences could be reliably matched with the GL Sμ and Sα1 or Sα2 regions, which enabled us to assessed the proportion of Sμ–Sα1/Sμ–Sα2 recombination junctions in IgA+ memory/effector B cells from systemic and mucosal tissues. The spleen contained mainly IgA1-switched B cells, representing by a predominance of Sμ–Sα1 switch fragments (93%, Figure 2). The proportion of IgA1-switched cells was significantly reduced in PBLs from adults as compared with the spleen (66%, χ2 test; P<0.01), but not in PBLs from children (80%).
Proportions of Sμ–Sα1/Sμ–Sα2 recombination junctions in switched B cells present in samples of spleen, unstimulated peripheral blood lymphocytes (PBLs), and various mucosal immune compartments. Black and white columns indicate IgA1 and IgA2 switching, respectively. Statistical analysis was performed with the χ2 test and statistically significant differences between spleen and PBLs or mucosal specimens are marked with asterisks. *P <0.05; **P <0.01; ***P <0.001.
Figure 2 illustrates the IgA subclass switch profiles of B cells in mucosal tissues. Palatine tonsils and adenoids, which make up most of Waldeyer’s ring and are referred to as nasopharynx-associated lymphoid tissue (NALT),7 showed a predominance of Sμ–Sα1 recombinational junctions (78% and 80%, respectively). However, salivary glands, lacrimal glands, and nasal mucosa, where effector/memory B cells are believed to be recruited mainly from NALT,28 contained a lower proportion of Sμ–Sα1 junctions (55%, 54%, and 60%, respectively; significantly lower as compared with spleen; χ2 test; P<0.05 or P<0.01). In the proximal small intestine (duodenum), the percentage of IgA1-switched cells (80%) was not significantly different from that in the spleen, whereas in the distal small intestine (ileum and Peyer’s patches), the frequency of IgA2-switched cells was markedly increased, reaching a nearly equal ratio of Sμ–Sα1/Sμ–Sα2 switching (χ2 test; P<0.05 between duodenum and ileum or Peyer’s patches). In the appendix and colonic mucosa, switching to IgA2 was much more frequent than to IgA1, with a dominance of Sμ–Sα2 junctions (≥70%) identified at both sites.
Composite Sμ–Sα junctions in switched B cells from systemic and mucosal immune compartments
We next determined the nucleotide sequences surrounding the Sμ–Sα breakpoints (Supplementary Figure S1). In most instances, a clear Sμ–Sα boundary could be identified, whereas in some sequences, a small stretch of nucleotides surrounding the putative breakpoint displayed perfectly matched homology with both Sμ and Sα, a phenomenon referred to as microhomology usage. Occasionally, a piece of the Sγ region was inserted between Sμ and Sα sequences, representing a sequential Sμ–Sγ–Sα CSR event.
The average length of microhomology at the Sμ–Sα junctions from various mucosal tissues was comparable to that in the spleen (Table 1). However, among the various mucosal tissues, significantly shorter stretches of microhomology were observed in both lacrimal gland and appendix than in the nasal mucosa (Student’s two-sided t-test; P<0.001). Point mutations on one or both sides of the Sμ–Sα junctions, reflecting the error-prone end-joining mechanism operating during the CSR process, were identified in some of the sequences. Although there appeared to be a lower frequency of mutations in the switch junctions derived from the lacrimal gland and duodenum, no statistically significant differences were found as compared with the spleen and PBLs (χ2 test, P>0.05). Furthermore, when Sμ–Sα1 and Sμ–Sα2 sequences were directly compared, no significant difference was observed in the composition of the recombination junctions. Taken together, as the nature of the recombination junctions, i.e., the microhomology usage as well as the frequency and pattern of mutations around the recombination breakpoints, reflects the DNA repair pathway used,18, 29 we conclude that the actual repair/recombination mechanisms involved during the IgA1 and IgA2 switching processes in systemic and mucosal immune compartments are largely similar.
It is, however, notable that the length of the S fragments was shorter in mucosal tissues than in spleen and PBLs, particularly so in the gut (Table 1; Student’s two-sided t-test; P<0.001). As the Sμ–Sα2 fragments were generally shorter than the Sμ–Sα1 fragments (average length 320 vs. 374 bp, based on all the Sμ–Sα fragments included in this study), this difference might simply be explained by a higher proportion of IgA2-switched cells in the gut. However, this subclass disparity could not fully account for the fragment length difference. Thus, although the proportion of Sμ–Sα1 in the duodenum was similar to that in the spleen and PBLs (Figure 2), the length of S fragments was still significantly shorter. Indeed, when the two types of S fragments, Sμ–Sα1 and Sμ–Sα2, were calculated separately, they were both significantly shorter in the duodenum, ileum, Peyer’s patches, appendix, and colon than in PBLs (data not shown). An alternative explanation could be that PCR preferentially amplifies shorter products whose overrepresentation in the gut is because of more abundant IgA-switched clones. However, this possibility was also excluded, as we did not observe any difference in the length of switch fragments when genomic DNA from a representative appendix sample was further diluted (from 30 ng to 3 ng, 1 ng, and 0.3 ng, respectively, per PCR reaction). Yet another possible explanation is that IgA switching in the gut is directed more toward the 5' of the Sμ and 3' end of the Sα regions, possibly through multiple rounds of recombination, and that the observed length of S fragments actually reflected the position of the breakpoints in the Sμ and Sα sequences (a plot of Sμ–Sα2 breakpoints derived from duodenum, appendix, colon, spleen, and PBL samples is shown in Supplementary Figure S2).
Direct vs. sequential IgA switching in different immune compartments
We subsequently investigated whether sequential switching contributes to the increased IgA2 production in the distal gut. To that end, we quantified the percentage of direct and sequential IgA switching in mucosal tissues and compared that with systemic lymphoid tissues. The data demonstrated that >90% of IgA switching had progressed directly from IgM to IgA in both compartments (Table 2). A small proportion of sequential switching to IgA2 tended to occur in the appendix and colon, but no significant difference was found between the distal gut and other tissue sites (Table 2).
Interestingly, some variations in the type of sequential IgA switching were observed between the systemic and mucosal immune compartments. Thus, in the spleen only Sμ–Sγ1–Sα1 switching was observed, whereas in the salivary glands and distal gut mucosa S regions from other IgG subclasses were also involved (Sγ2 and Sγ3; Table 2). Sequential switching in nasal mucosa involved two Sα regions, namely Sμ–Sα1–Sα2 and Sμ–Sα2–Sα1.
Regulation of IgA1/IgA2 switching in proximal and distal gut mucosa
To further understand the mechanisms underlying differential regulation of IgA subclass switching at different mucosal effector sites, we designed a series of reverse transcriptase-PCR (RT-PCR) assays to measure the level of GL and circle transcripts (Figure 1a) as well as the expression of BAFF and APRIL in tissue specimens from both the small bowl (duodenum and ileum) and the large bowl (colon). Samples from Peyer’s patches, representing the major intestinal inductive site, and PBLs were also included for comparison.
As shown in Figure 3a, the Iα1–Cα1 GL transcripts were detectable in some of the duodenal/ileal (lanes 2, 4, 5, and 10) and colonic (lane 6) tissue samples. However, as compared with the samples of Peyer’s patches (lanes 11–14, with 2% of complementary DNA (cDNA) input) and IL-10+CD40 ligand-activated PBLs (lane 16, with 5% of cDNA input), the expression level at these mucosal effector sites was extremely low. The Iα2–Cα2 GL transcripts were undetectable in half of the lamina propria samples and barely detectable in the remaining ones derived from duodenum/ileum/colon (lanes 2, 3, 4, 8, and 10), but were easily seen in the Peyer’s patch samples (lanes 11–14, with 2% of cDNA input).
Expression of Iα1–Cα and Iα2–Cα2 germline (GL) transcripts, Iα–Cμ circle transcripts, and AID, CD19, APRIL, and BAFF in proximal and distal gut mucosa. Mucosal specimens from the duodenum/ileum and colon are labeled with blue and red numbers, respectively, whereas specimens of Peyer’s patches are indicated by green numbers. Samples 1 and 3 were taken from colonic and samples 2, 4, and 5 were taken from duodenal lamina propria of different individuals, whereas samples 6–10 were obtained from the same individual at different sites (6, ascending colon; 7, ascending colon with inflammation; 8, descending colon; 9, duodenum; 10, ileum). Samples 11–14 represented Peyer’s patches from four different individuals. Samples 15 and 16 were derived from unstimulated and IL-10+CD40L-stimulated peripheral blood lymphocytes (PBLs) respectively. (a) Gel electrophoresis of GL transcripts amplified by reverse transcriptase-PCR (RT-PCR). Arrows indicate the desired size of the amplicons. Because of the large difference in the expression levels of GL transcripts among different samples, only 1/50th or 1/20th of complementary DNA (cDNA), respectively, was added in the PCR reactions for Peyer’s patches or PBLs, but equal amounts of cDNA were added throughout for the amplification of Iα–Cμ circle transcripts and the housekeeping gene β-actin. Three independent experiments were performed and representative pictures are shown. M, molecular marker. (b) Measurement of AID, CD19, APRIL, and BAFF expression by real-time quantitative RT-PCR. Equal amounts of cDNA were added in all the amplifications. The expression level of each gene in every sample was first normalized in relation to the β-actin housekeeping gene. A relative expression level was subsequently determined by comparing the normalized value of each sample with sample 15, the unstimulated PBLs. An average of three independent experiments was plotted and the dispersion bar indicates the s.e. value.
We subsequently studied the expression of AID and Iα–Cμ circle transcripts, two hallmarks for ongoing CSR activities. Compared with the Peyer’s patch samples and the IL-10+CD40-ligand activated PBLs, AID was expressed at an extremely low or even undetectable level (not reliably determined by real-time PCR, with the cycle threshold (CT) value of >34) in all the duodenal, ileal, and colonic lamina propria specimens (Figure 3b). This is in line with the extremely low number of B lymphocytes in the lamina propria samples as measured by the expression level of CD19 (Figure 3b), and as previously demonstrated by immunohistochemistry outside of GALT structures.30 Because of the extensive homology between the Iα1 and Iα2 sequences, we designed an assay to amplify the Iα1 or Iα2 circle transcripts together (Iα–Cμ). These transcripts were reliably detected in each experiment (and confirmed by sequencing) in all the Peyer’s patch samples and in activated PBLs, but could only be amplified occasionally in a few ileal and colonic lamina propria samples after multiple attempts (Figure 3a). Furthermore, the weak bands detected in lamina propria samples were often nonspecific and could only be confirmed by sequencing as a real circle transcript in one ileal and one colonic sample respectively. Thus, although ongoing IgA switching could occasionally be detected in samples derived from proximal as well as distal intestinal effector sites, this is an extremely rare event as compared with Peyer’s patch samples.
We next determined the mRNA expression of BAFF and APRIL. BAFF, a cytokine that has been suggested to induce IgA1 switching as well as to promote the survival of IgA2+ B cells, was clearly present at a higher level in the duodenum (IgA1 dominant; lanes 2, 4, 5, and 9 in Figure 3b) than in the colon (IgA2 dominant; lanes 1, 3, 6, 7, and 8 Figure 3b). However, APRIL, the cytokine that has been suggested to promote IgA2 switching specifically, was expressed at more comparable levels in these sites (Figure 3b). In Peyer’s patches, where nearly equal proportions of Sμ–Sα1/Sμ–Sα2 switch fragments were observed, the expression level of APRIL was much lower than in both the duodenal (IgA1 dominant) and colonic (IgA2 dominant) effector sites (Figure 3b). Similar results were also obtained in the nasal mucosal samples (lanes 17–19 in Supplementary Figure S3). Taken together, there was no clear relationship of expression of BAFF and APRIL, or the ratio of APRIL/BAFF, with the proportion of switched IgA1 and IgA2 cells in the mucosal samples (Supplementary Figure S4).
DISCUSSION
We studied for the first time at the molecular level the subclass profiles of IgA expression among systemic and mucosal immune compartments. Although our PCR-based strategy could not make a distinction between productive and nonproductive alleles, our molecular results generally paralleled the distribution of IgA1- and IgA2-producing plasma cells as previously determined by immunohistochemistry,4 with some minor deviations.
The spleen collects and responds to circulating antigens and thus plays an important role in maintaining systemic immunity. On average, 93% of the IgA+ splenic plasma cells are of the IgA1 phenotype,4, 5 which is similar to our molecular data (94%). This organ is a likely major source of the predominating monomeric IgA1 in serum.
The human NALT structures (palatine tonsils and adenoids) also contained a high proportion of Sμ–Sα1 junctions (78% and 80%, respectively), which is reasonable because these organs act as a “cross-road” between mucosal and systemic immunity.7 B cells activated in human NALT express several homing molecules shared between the systemic immune system and mucosal effector sites of the upper aero-digestive tract.28 These lymphoid structures also exert local effector functions of both systemic and secretory immunity.31
Salivary glands, lacrimal glands, and nasal mucosa, where the IgA-committed B cells apparently are mainly recruited from NALT, showed a higher level of IgA2 switching than NALT. Several possibilities could explain this discrepancy. First, as secretory IgA antibodies against microbial polysaccharides are predominantly of the IgA2 subclass,32, 33 increased exposure to such antigens at these secretory effector sites might influence the retention and local differentiation of B cells that have already switched to IgA2. Second, some of the IgA2-producing cells in this region are most likely derived from Peyer’s patches.8, 9 Third, a relatively high level of switched IgA2 cells in nasal mucosa might be contributed by local CSR, which reportedly may take place at this site,34 and where scattered ILFs may occur.7
It follows that the percentage of Sμ–Sα1-switched cells (60%) did not match the high proportion of IgA1+ plasma cells (90%) previously detected by immunohistochemistry in nasal mucosa.4 One possible interpretation is that coexistence of B cells not differentiated to plasma cells could have diluted out the switch junctions of surface IgA+ cells of the other subclass. Alternatively, there are individual variations among the tissue samples; and we examined nasal mucosa from three subjects, of which one biopsy showed a dominance of IgA1 switching. It is also interesting to note that in nasal mucosa, IgA1+ B cells could serve as precursors of IgA2+ B cells through sequential switching (Sμ–Sα1–Sα2), and vice versa, involving both sequential switching and potentially trans-switching (Sμ–Sα2–Sα1, where the two Sα regions are not from the same allele). These switch activities between the two IgA subclasses might reflect the alteration of antigens encountered in the nasal mucosa of the same individual and could contribute to the varying proportion of IgA1+ or IgA2+ plasma cells observed in different individuals.
The intestinal IgA system is the best-understood contributor to mucosal immunity.7, 9, 35 It is generally accepted that Peyer’s patches are the main inductive sites for gut immunity.36, 37 Although a nearly equal ratio of Sμ–Sα1/Sμ–Sα2 switch junctions was found in Peyer’s patches, there was a striking difference between the duodenal (Sμ–Sα1, 80%) and ileal (Sμ–Sα1, 52%) lamina propria effector sites. The reason for this regional disparity is unclear but could reflect an impact of the local luminal antigens or microbiota.14, 38 The proximal small intestine has a low bacterial density but is abundantly exposed to food proteins, whereas the distant small intestine has a massive microbial load. The shift from IgA1- to IgA2-producing cells probably represents an adaptive response to protect the mucosa from the increased exposure to lipopolysaccharide-containing bacteria through mechanisms discussed above.14 Consistent with this, the colonic lamina propria, adjacent to the gut lumen where lipopolysaccharide-containing bacteria reach the highest density (1010–1012 organisms per g), showed an overwhelming predominance of Sμ–Sα2 switch junctions (70%), contributing to an abundance of IgA2-producing plasma cells. APRIL, produced by intestinal epithelial cells or dendritic cells, has been suggested as the key factor driving the T-cell-independent IgA2 CSR in colonic mucosa.23 Our data could to some extent be taken to support this idea, as we detected a relatively high ratio of APRIL/BAFF as well as signs of ongoing CSR in the colonic samples. However, the expression level of APRIL/BAFF, or the ratio of APRIL/BAFF in duodenal and ileal lamina propria and Peyer’s patch samples, as well as in nasal mucosa, did not show a clear relationship to the distribution of IgA1/IgA2-switched cells (Figure 3b and (Supplementary Figure S4). The mRNA expression levels of APRIL/BAFF may not correlate with the amounts of cytokines secreted, but it is also possible that these cytokines do not have a decisive effect on the IgA subclass switching/distribution in vivo. Furthermore, it is noteworthy that in the studies by Fagarasan et al.39 and He et al.,23 where the concept of in situ IgA switching in mouse and human gut lamina propria was first proposed, resection material from the gut was used and the possibility that ILFs might have contaminated the analyzed cells was not considered. However, in more recent studies by Barone et al.,40 Bergqvist et al.,41 and Shikina et al.,42 gut tissues containing organized lymphoid follicles were carefully excluded, and a different conclusion was reached, i.e., GALT is the major site for IgA CSR and no evidence was obtained to suggest that IgA CSR occurs in the intestinal lamina propria of humans and mice. Small CD19+ B lymphocytes are rare outside the immediate vicinity of GALT structures in the human gut and they usually express surface IgA.30 Some of these cells may show sign of proliferation (nuclear marker Ki-67+), which is a prerequisite for CSR; but as they also express the postgreminal center marker CD27, they were deemed to represent cells recently migrated from GALT structures.43 Thus, although we did not microdissect the included lamina propria samples to fully exclude potentially contaminating organized lymphoid tissues such as ILFs, our results suggested that if in situ switching occurs outside of GALT, it is a very rare event compared with the IgA switch activity in organized lymphoid structures, especially the Peyer’s patches. This was clearly indicated by the extremely low or undetectable level of AID and IgA circle transcripts in lamina propria samples.
Activated B cells from GALT and NALT migrate through the blood and cannot be easily distinguished from each other in the circulation.13, 28 The proportion of Sμ–Sα2 junctions in PBLs from children (20%) was lower than that in the PBLs from adults (34%), which could reflect an increasing contribution of circulating IgA precursor cells from GALT with age. Immunohistochemical studies of salivary glands have indeed suggested that the proportion of IgA2-producing plasma cells increases with age,44 perhaps reflecting distant homing from GALT.
Both direct and sequential IgA switching activities have been suggested to participate in the development of IgA+ plasma cells.45, 46 APRIL can reportedly induce sequential CSR to IgA2 in vitro, either in IgA1+ or in IgG1+ B cells, and intestinal lamina propria IgA2+ B cells have been shown to contain Sμ–Sα1–Sα2 or Sμ–Sγ1–Sα2 junctions.23 However, to what extent sequential CSR contributes to IgA subclass production in vivo remains unclear. Our study demonstrated that switching to IgA1 and IgA2 in systemic and mucosal immune compartments, including at the intestinal mucosa, mainly progresses directly from Cμ to Cα, whereas only a minor proportion proceeds from Cμ to Cα via Cγ or the other Cα gene. This is in agreement with the recent study on human memory B-cell subsets where, in contrast to IgG+ cells, the CD27+IgA+ and CD27−IgA+ (presumably resembling the IgA+ cells from the gut lamina propria) cells mainly showed direct Sμ–Sα switching (>95%).11 Similarly, in mouse B cells, sequential CSR from IgM through IgG to IgA contributed very little to IgA production in vitro and in vivo, in contrast to the situation for IgE.47 It is also interesting to note that in colonic mucosa, the rare sequential events that we observed were Sμ–Sγ2–Sα2. IgG2+ cells rather than IgG1+ or IgG3+ cells could thus act as precursors to add to the IgA2 repertoire in colonic mucosa. Indeed, IgG2+ plasma cells are more abundant than the IgG3 counterparts at this effector site, whereas the reverse is true in nasal mucosa.48
Taken together, our data demonstrate that there is a striking disparity of IgA subclass switch profile at different mucosal sites. The remarkable regionalization of activated B cells that have undergone IgA CSR is closely related to the distribution of plasma cells producing the two IgA subclasses. The subclass commitment seems to be largely determined by CSR at regional inductive sites before the IgA effector cells are distributed by their homing molecules. Thus, our study suggested that at most a minor molecular tailoring of the local subclass profiles might take place by in situ switching at mucosal effector sites—that is, outside of GALT and NALT structures. Such compartmentalized differentiation of IgA responses has to be taken into account in the development of mucosal vaccines and in their application strategy.
METHODS
Tissue specimens The following mucosal tissues were sampled with ethical approval from immunologically normal subjects (aged 10–77 years)28 who had endoscopic biopsy or surgical resection performed for diagnostic or therapeutic purposes: duodenum (n=7), ileum (n=4), Peyer’s patches (n=7), colon (n=6), adenoids (n=3), palatine tonsils (n=5), salivary glands (n=4), lacrimal glands (n=3), nasal mucosa (n=6), appendix (n=2), and spleen (n=3). The tissue specimens were placed directly in ice-chilled tubes and brought to the laboratory within 1 h, or they were snap-frozen bed-side in liquid nitrogen. It is noteworthy that the samples excised from the surgical resection preparations were of a small size comparable to the targeted biopsies. Control sections (from the excised samples) stained with hematoxylin and eosin were examined to exclude as well as feasible that organized lymphoid tissue was present in the lamina propria samples. One biopsy (sample 7 in Figure 3) was obtained from an inflamed mucosal area; the remaining specimens were all from sites without macroscopical signs of inflammation and were deemed to be normal by hematoxylin and eosin staining. PBLs were obtained from 17 healthy adults and 14 healthy children (1–6 years old), as previously reported.24, 25
PCR amplification of Sμ–Sα fragments. Genomic DNA was purified from various tissue specimens and PBLs by previously described methods.26, 28, 49 The amplification of Sμ–Sα fragments from in vivo switched cells was also performed as described before.49 Briefly, two pairs of Sμ– and Sα-specific primers were used in a nested PCR assay. The Sα-specific primers were located in a region 3′ of the repetitive sequences and perfectly matched with both Sα1 and Sα2 sequences; together with the Sμ-specific primers, they were therefore expected to amplify the Sμ–Sα1 and Sμ–Sα2 fragments with the same efficiency. In the first run, 30 ng of DNA was added in a 25 μl reaction mix and amplified for a total of 30 cycles consisting of 1 min at 94 °C, 1 min at 60 °C and 1 min 30 s at 72 °C. Then, 1 μl of PCR products from the first run was used in the second run as templates and under the same conditions, except that the annealing temperature was changed to 65 °C and the extension time at 72 °C was 1 min. The number of Sμ–Sα fragments was determined from 10 PCR reactions run in parallel with DNA from one individual, representing random amplification of in vivo IgA-switched clones. The PCR error rate (0.9/1,000 nucleotides) was estimated previously.49
Analysis of Sμ–Sa junctions. The PCR-amplified Sμ–Sα fragments were subsequently purified, cloned, and sequenced. The Sμ–Sα breakpoints were determined by aligning the S fragment sequences with the Sμ (X54713), Sα1 (L19121), and Sα2 (AF030305) sequences.49 S fragment sequences that did not completely match with the Sμ sequences were subsequently aligned with the Sγ1 (U39737), Sγ2 (U39934), Sγ3 (U39935), or Sγ4 (Y1254-52) sequences.50 Direct IgA switching was defined by progressing from Sμ to Sα, whereas sequential IgA switching was defined by proceeding from Sμ to Sα via Sγ (Sμ–Sγ–Sα) or from one IgA subclass to another (Sμ–Sα1–Sα2 or Sμ–Sα2–Sα1).
Sμ–Sα fragments, rather than Sμ–Sγ–Sα fragments, were included for microhomology and mutation analyses. Microhomology was defined as successive nucleotides that were shared by both the Sμ and Sα regions at the switch junction (without mismatches). Analysis of microhomology usage and mutation pattern around the switch junctions was performed as previously described25 and followed the recently suggested guideline.29
Amplification of Iα–Cα GL and Iα–Cμ circle transcripts. Total RNA was isolated from 20 to 30 mg snap-frozen mucosal tissue by the RNeasy mini kit (Qiagen, Hilden, Germany). Dnase I (Qiagen) was used to remove trace amounts of genomic DNA. First-strand cDNA synthesis was performed with a NotI-d(T)18 primer using a cDNA synthesis kit (Pharmacia, Uppsala, Sweden).
Iα-consensus (5'-CCCCACAGCAGCCCTCTTG-3′), Cα1-specific (5'-GCGGTTA GCGGGGTCTTGG-3′), and Cα2-specific (5'-GGCGGTTAGTGGGGTCTTCA-3′) primers were used to amplify the Iα1–Cα1 and Iα2–Cα2 GL transcripts. Iα-consensus and Cμ-antisense (5'-AATCTGCCGGGGACTGAAAC-3′) primers were used to amplify the Iα–Cμ circle transcripts. Amplification was performed in 40 cycles, at 94 °C for 50 s, at 68 °C (66 °C for Iα–Cμ) for 50 s, and at 72 °C for 1.5 min (1 min for Iα–Cμ), respectively. Amplification of β-actin was used to monitor the integrity of RNA and cDNA synthesis. The desired amplicons were checked for correctness by sequencing.
Expression of AID, BAFF, APRIL , and CD19 analyzed by real-time RT-PCR. Real-time RT-PCR assays were performed to quantitatively measure the expression level of AID, BAFF, and APRIL transcripts. A qPCR Core kit for SYBR Green I (Eurogentec, Seraing, Belgium) was used together with the previously described primers for amplification of AID transcripts and the housekeeping gene β-actin,51 as well as with the following primers for amplification of BAFF, APRIL, and CD19 transcripts: BAFF-forward (5'-TGCCTGAAACACTACCCAATAA-3′) and BAFF-reverse (5'-AGCAGTTTCAATGCACCAAA-3′); APRIL-forward (5'-GGGTATCCCTGGCAGA GTC-3′) and APRIL-reverse (5'-CTCCTTTTCCGGGATCTCTC-3′); CD19-forward (5'-AACCTGACCATGTCATTCCACC-3′) and CD19-reverse (5'-GCCAAA GTCACAGCTGAGACCT-3′). Amplification was performed on a StepOne Real-Time PCR System (Applied Biosystems, Foster City, CA) with the following conditions: at 95 °C for 10 min, followed by 40 cycles, each cycle at 95 °C for 10 s and at 60 °C for 1 min. Melting curve analysis was performed immediately after the above run to ensure that the amplification was specific. Relative expression of AID, CD19, BAFF, and APRIL was calculated as previously described.52
Statistical analysis. Categorical variables, such as mutation frequency and proportion of Sμ–Sα subclass, were represented as numbers (%), and compared by the χ2 test. Continuous variables, such as microhomology usage and length of the S fragments, were expressed as mean±s.d. and compared by Student’s t-test. All statistical assessments were two sided and evaluated at the 0.05 level of significance. The analyses were performed with the SPSS 13.0 software (IBM, NY).
References
Flanagan, J.G. & Rabbitts, T.H. Arrangement of human immunoglobulin heavy chain constant region genes implies evolutionary duplication of a segment containing gamma, epsilon and alpha genes. Nature 300, 709–713 (1982).
Schroeder, H.W. Jr & Cavacini, L. Structure and function of immunoglobulins. J. Allergy Clin. Immunol. 125 (2 Suppl 2), S41–S52 (2010).
Kilian, M., Reinholdt, J., Lomholt, H., Poulsen, K. & Frandsen, E.V. Biological significance of IgA1 proteases in bacterial colonization and pathogenesis: critical evaluation of experimental evidence. APMIS 104, 321–338 (1996).
Kett, K., Brandtzaeg, P., Radl, J. & Haaijman, J.J. Different subclass distribution of IgA-producing cells in human lymphoid organs and various secretory tissues. J. Immunol. 136, 3631–3635 (1986).
Crago, S.S. et al. Distribution of IgA1-, IgA2-, and J chain-containing cells in human tissues. J. Immunol. 132, 16–18 (1984).
Delacroix, D.L., Dive, C., Rambaud, J.C. & Vaerman, J.P. IgA subclasses in various secretions and in serum. Immunology 47, 383–385 (1982).
Brandtzaeg, P. Function of mucosa-associated lymphoid tissue in antibody formation. Immunol. Invest. 39, 303–355 (2010).
Brandtzaeg, P. & Johansen, F.E. Mucosal B cells: phenotypic characteristics, transcriptional regulation, and homing properties. Immunol. Rev. 206, 32–63 (2005).
Brandtzaeg, P. Mucosal immunity: induction, dissemination, and effector functions. Scand. J. Immunol. 70, 505–515 (2009).
Brandtzaeg, P. et al. Immunobiology and immunopathology of human gut mucosa: humoral immunity and intraepithelial lymphocytes. Gastroenterology 97, 1562–1584 (1989).
Berkowska, M.A. et al. Human memory B cells originate from three distinct germinal center-dependent and -independent maturation pathways. Blood 118, 2150–2158 (2011).
Fernandes, J.R. & Snider, D.P. Polymeric IgA-secreting and mucosal homing pre-plasma cells in normal human peripheral blood. Int. Immunol. 22, 527–540 (2010).
Pakkanen, S.H. et al. Expression of homing receptors on IgA1 and IgA2 plasmablasts in blood reflects differential distribution of IgA1 and IgA2 in various body fluids. Clin. Vaccine Immunol. 17, 393–401 (2010).
Kett, K., Baklien, K., Bakken, A., Kral, J.G., Fausa, O. & Brandtzaeg, P. Intestinal B-cell isotype response in relation to local bacterial load: evidence for immunoglobulin A subclass adaptation. Gastroenterology 109, 819–825 (1995).
Muramatsu, M., Kinoshita, K., Fagarasan, S., Yamada, S., Shinkai, Y. & Honjo, T. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Stavnezer, J., Guikema, J.E. & Schrader, C.E. Mechanism and regulation of class switch recombination. Annu. Rev. Immunol. 26, 261–292 (2008).
Pan-Hammarstrom, Q., Zhao, Y. & Hammarstrom, L. Class switch recombination: a comparison between mouse and human. Adv. Immunol. 93, 1–61 (2007).
Kotnis, A., Du, L., Liu, C., Popov, S.W. & Pan-Hammarstrom, Q. Non-homologous end joining in class switch recombination: the beginning of the end. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 364, 653–665 (2009).
Kinoshita, K., Harigai, M., Fagarasan, S., Muramatsu, M. & Honjo, T. A hallmark of active class switch recombination: transcripts directed by I promoters on looped-out circular DNAs. Proc. Natl. Acad. Sci. USA 98, 12620–12623 (2001).
Stavnezer, J. Antibody class switching. Adv. Immunol. 61, 79–146 (1996).
Islam, K.B., Nilsson, L., Sideras, P., Hammarstrom, L. & Smith, C.I. TGF-beta 1 induces germ-line transcripts of both IgA subclasses in human B lymphocytes. Int. Immunol. 3, 1099–1106 (1991).
Shockett, P. & Stavnezer, J. Effect of cytokines on switching to IgA and alpha germline transcripts in the B lymphoma I.29 mu. Transforming growth factor-beta activates transcription of the unrearranged C alpha gene. J. Immunol. 147, 4374–4383 (1991).
He, B. et al. Intestinal bacteria trigger T cell-independent immunoglobulin A(2) class switching by inducing epithelial-cell secretion of the cytokine APRIL. Immunity 26, 812–826 (2007).
Du, L. et al. Involvement of Artemis in nonhomologous end-joining during immunoglobulin class switch recombination. J. Exp. Med. 205, 3031–3040 (2008).
Pan-Hammarstrom, Q. et al. Impact of DNA ligase IV on nonhomologous end joining pathways during class switch recombination in human cells. J. Exp. Med. 201, 189–194 (2005).
Pan, Q., Petit-Frere, C., Lahdesmaki, A., Gregorek, H., Chrzanowska, K.H. & Hammarstrom, L. Alternative end joining during switch recombination in patients with ataxia-telangiectasia. Eur. J. Immunol. 32, 1300–1308 (2002).
Pan-Hammarstrom, Q. et al. Disparate roles of ATR and ATM in immunoglobulin class switch recombination and somatic hypermutation. J. Exp. Med. 203, 99–110 (2006).
Johansen, F.E., Baekkevold, E.S., Carlsen, H.S., Farstad, I.N., Soler, D. & Brandtzaeg, P. Regional induction of adhesion molecules and chemokine receptors explains disparate homing of human B cells to systemic and mucosal effector sites: dispersion from tonsils. Blood 106, 593–600 (2005).
Stavnezer, J., Bjorkman, A., Du, L., Cagigi, A. & Pan-Hammarstrom, Q. Mapping of switch recombination junctions, a tool for studying DNA repair pathways during immunoglobulin class switching. Adv. Immunol. 108, 45–109 (2010).
Farstad, I.N., Carlsen, H., Morton, H.C. & Brandtzaeg, P. Immunoglobulin A cell distribution in the human small intestine: phenotypic and functional characteristics. Immunology 101, 354–363 (2000).
Brandtzaeg, P. Potential of nasopharynx-associated lymphoid tissue for vaccine responses in the airways. Am. J. Respir. Crit. Care. Med. 183, 1595–1604 (2011).
Tarkowski, A., Lue, C., Moldoveanu, Z., Kiyono, H., McGhee, J.R. & Mestecky, J. Immunization of humans with polysaccharide vaccines induces systemic, predominantly polymeric IgA2-subclass antibody responses. J. Immunol. 144, 3770–3778 (1990).
Ladjeva, I., Peterman, J.H. & Mestecky, J. IgA subclasses of human colostral antibodies specific for microbial and food antigens. Clin. Exp. Immunol. 78, 85–90 (1989).
Takhar, P. et al. Allergen drives class switching to IgE in the nasal mucosa in allergic rhinitis. J. Immunol. 174, 5024–5032 (2005).
Fagarasan, S. & Honjo, T. Intestinal IgA synthesis: regulation of front-line body defences. Nat. Rev. Immunol. 3, 63–72 (2003).
Macpherson, A.J., McCoy, K.D., Johansen, F.E. & Brandtzaeg, P. The immune geography of IgA induction and function. Mucosal Immunol. 1, 11–22 (2008).
Mora, J.R. et al. Generation of gut-homing IgA-secreting B cells by intestinal dendritic cells. Science 314, 1157–1160 (2006).
Macpherson, A.J. IgA adaptation to the presence of commensal bacteria in the intestine. Curr. Top. Microbiol. Immunol. 308, 117–136 (2006).
Fagarasan, S., Kinoshita, K., Muramatsu, M., Ikuta, K. & Honjo, T. In situ class switching and differentiation to IgA-producing cells in the gut lamina propria. Nature 413, 639–643 (2001).
Barone, F., Patel, P., Sanderson, J.D. & Spencer, J. Gut-associated lymphoid tissue contains the molecular machinery to support T-cell-dependent and T-cell-independent class switch recombination. Mucosal Immunol. 2, 495–503 (2009).
Bergqvist, P., Stensson, A., Lycke, N.Y. & Bemark, M. T cell-independent IgA class switch recombination is restricted to the GALT and occurs prior to manifest germinal center formation. J. Immunol. 184, 3545–3553 (2010).
Shikina, T. et al. IgA class switch occurs in the organized nasopharynx- and gut-associated lymphoid tissue, but not in the diffuse lamina propria of airways and gut. J. Immunol. 172, 6259–6264 (2004).
Halstensen, T.S. & Brandtzaeg, P. Activated T lymphocytes in the celiac lesion: non-proliferative activation (CD25) of CD4+ alpha/beta cells in the lamina propria but proliferation (Ki-67) of alpha/beta and gamma/delta cells in the epithelium. Eur. J. Immunol. 23, 505–510 (1993).
Thrane, P.S., Rognum, T.O. & Brandtzaeg, P. Ontogenesis of the secretory immune system and innate defence factors in human parotid glands. Clin. Exp. Immunol. 86, 342–348 (1991).
Hammarstrom, L., Mellstedt, H., Persson, M.A., Smith, C.I. & Ahre, A. IgA subclass distribution in paraproteinemias: suggestion of an IgG-IgA subclass switch pattern. Acta. Pathol. Microbiol. Immunol. Scand. C 92, 207–211 (1984).
Conley, M.E. & Bartelt, M.S. In vitro regulation of IgA subclass synthesis. II. The source of IgA2 plasma cells. J. Immunol. 133, 2312–2316 (1984).
Kaminski, D.A. & Stavnezer, J. Stimuli that enhance IgA class switching increase histone 3 acetylation at S alpha, but poorly stimulate sequential switching from IgG2b. Eur. J. Immunol. 37, 240–251 (2007).
Bjerke, K. & Brandtzaeg, P. Terminally differentiated human intestinal B cells. J chain expression of IgA and IgG subclass-producing immunocytes in the distal ileum compared with mesenteric and peripheral lymph nodes. Clin. Exp. Immunol. 82, 411–415 (1990).
Pan, Q. et al. Regulation of switching and production of IgA in human B cells in donors with duplicated alpha1 genes. Eur. J. Immunol. 31, 3622–3630 (2001).
Pan, Q., Rabbani, H. & Hammarstrom, L. Characterization of human gamma 4 switch region polymorphisms suggests a meiotic recombinational hot spot within the Ig locus: influence of S region length on IgG4 production. J. Immunol. 161, 3520–3526 (1998).
Cagigi, A. et al. CD27(-) B-cells produce class switched and somatically hyper-mutated antibodies during chronic HIV-1 infection. PLoS One 4, e5427 (2009).
Wen, S., Felley, C.P., Bouzourene, H., Reimers, M., Michetti, P. & Pan-Hammarstrom, Q. Inflammatory gene profiles in gastric mucosa during Helicobacter pylori infection in humans. J. Immunol. 172, 2595–2606 (2004).
Acknowledgements
We thank S. Dai and W. Zhou for help with the switch junction analysis; N. Wang for help with the real-time PCR assay; R. Peng and H. Eliassen for manuscript preparation; and H.S. Carlsen and G.R. Melum for providing some of the mucosal tissue samples. This work was supported by the Swedish Research Council, the Swedish Cancer Society, the European Research Council (242551-ImmunoSwitch), the Research Council of Norway, and Oslo University Hospital.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
Author contributions
P.B. and Q.P.-H. designed and supervised the study; M.L., L.D., and Q.P.-H. performed the experiments; M.L., P.B., and Q.P.-H. wrote the manuscript.
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Rights and permissions
About this article
Cite this article
Lin, M., Du, L., Brandtzaeg, P. et al. IgA subclass switch recombination in human mucosal and systemic immune compartments. Mucosal Immunol 7, 511–520 (2014). https://doi.org/10.1038/mi.2013.68
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2013.68
This article is cited by
-
NOD2 deficiency increases retrograde transport of secretory IgA complexes in Crohn’s disease
Nature Communications (2021)
-
Immunoprofiles associated with controlled human malaria infection and naturally acquired immunity identify a shared IgA pre-erythrocytic immunoproteome
npj Vaccines (2021)
-
Local eosinophils are associated with increased IgA subclass levels in the sinonasal mucosa of chronic rhinosinusitis with polyp patients
Allergy, Asthma & Clinical Immunology (2020)
-
FcαRI co-stimulation converts human intestinal CD103+ dendritic cells into pro-inflammatory cells through glycolytic reprogramming
Nature Communications (2018)
-
The human intestinal B-cell response
Mucosal Immunology (2016)