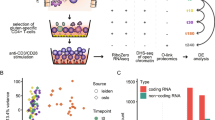
Abstract
We have analyzed the production of the effector cytokines interleukin (IL)-17, IL-21, and IL-22 in gluten-reactive CD4+ T cells of celiac disease patients, either cultured from small intestinal biopsies or isolated from peripheral blood after an oral gluten challenge. Combining intracellular cytokine staining with DQ2-α-II gliadin peptide tetramer staining of intestinal polyclonal T-cell lines, we found that gluten-specific T cells produced interferon-γ (IFN-γ) and IL-21, but not IL-17 or IL-22, even if other T cells of the same lines produced these cytokines. Similarly, in DQ2-α-II-specific T cells in peripheral blood of gluten-challenged patients, very few stained for intracellular IL-17, whereas many cells stained for IFN-γ. We conclude that gluten-reactive T cells produce IL-21 and IFN-γ, but not IL-17. Their production of IL-21 suggests a role for this cytokine in the pathogenesis of celiac disease.
Similar content being viewed by others
Introduction
Celiac disease is a chronic inflammatory disease of the small intestine precipitated by ingestion of gluten proteins (consisting of gliadin and glutenin subcomponents) in genetically susceptible individuals. The disease lesion is characterized by lymphocytic infiltration of the epithelium and the lamina propria, crypt cell hyperplasia, and loss of villus structures.1 Gluten-reactive T cells can be cultured from intestinal biopsies of celiac disease patients, but not from control individuals.2 The gluten-reactive T cells recognize a variety of different epitopes,3, 4 which generally are much better recognized by the T cells after deamidation by the enzyme transglutaminase 2 (TG2).5, 6 The strong HLA association of celiac disease with HLA-DQ2 and HLA-DQ87 and selective recognition of gluten peptides in the context of the very same HLA molecules by CD4+ T cells8 indicate that these T cells control the immune reaction that leads to the lesion formation.
CD4+ T-helper (Th) cells can be divided into subsets based on their transcription factors, main cytokines produced, and their function. The subsets Th1, Th2, and T regulatory cells TR are well established. Recently, Th17 and follicular T-helper cells were defined.9 Th17 cells are characterized by expression of the transcription factor RORC (RAR-related orphan receptor C; RORγt in mouse) and the production of the proinflammatory cytokines interleukin (IL)-17A, IL-17F, and often IL-21 and IL-22. The IL-21 cytokine is also produced by follicular T-helper cells that help B cells in making antibodies during T cell-dependent responses.9 The cytokine production by the gluten-reactive CD4+ T cells of the intestinal lesion is dominated by interferon-γ (IFN-γ),10 although several other cytokines are also produced by such cells, including IL-410 and occasionally IL-10.11 Two recent studies showed an increase in IL-17 mRNA in small intestinal biopsies of untreated celiac disease patients.12, 13 Both T cells and non-T cells were found to produce increased IL-17 in celiac disease, and gliadin challenge of intestinal biopsies from treated celiac disease patients identified CD4+ as well as CD4+CD8+ T cells to be the source of IL-17. Furthermore, blockage of IL-21 in the challenge experiments with neutralizing antibodies led to diminished IL-17 production. These observations suggest that IL-17 and IL-21 may be implicated in celiac disease, but it is unclear whether gluten-reactive CD4+ T cells are the source of these cytokines. In this study we have addressed whether gluten-reactive T cell produce IL-17, IL-21, and IL-22 by analyzing gluten-reactive CD4+ T cells of celiac disease patients, either cultured from small intestinal biopsies or isolated from peripheral blood after an oral gluten challenge. The results, for which tetramer staining of polyclonal lines was particularly informative, show that gluten-reactive CD4+ T cells of the celiac lesion produce IL-21, but not IL-17 or IL-22.
Results
Gluten-specific T-cell clones do not produce IL-17 but produce IL-21
We first examined whether gluten-reactive T-cell clones generated from duodenal biopsies of celiac disease patients produce IL-17. The T-cell clones chosen for analysis were well characterized; all were DQ2.5 (i.e., DQA1*0501/DQB1*0201) restricted, recognized several different gliadin epitopes (DQ2-α-I, DQ2-α-II, DQ2-γ-I, or DQ2-γ-II), and were generated from different patients. The clones were stimulated with either TG2-treated and chymotrypsin-digested gluten (CT-gluten), peptide antigen, or phorbol 12-myristate 13-acetate (PMA)/ionomycin, and the culture supernatants were analyzed for the presence of IFN-γ, IL-17, IL-21, and IL-22 by Luminex assay or enzyme-linked immunosorbent assay. Several of the gluten-reactive T-cell clones produced both IL-21 and IFN-γ, whereas some produced IL-22 (Figure 1). However, none of the gluten-reactive T-cell clones produced IL-17. Intracellular cytokine staining and flow cytometry on these T-cell clones confirmed the findings of the culture supernatants. Indeed, many of the T-cell clones were double producers of IL-21 and IFN-γ. In some T-cell clones, there were some very few cells that produced IL-22 (<1 to 2% for the highest producers) that could possibly be remaining feeder cells (Supplementary Figure S1 online and data not shown). None of the tested T-cell clones produced IL-17 as assessed by flow cytometry (Supplementary Figure S1 online).
Gluten-specific T-cell clones produce interleukin (IL)-21 and interferon-γ (IFN-γ), but not IL-17. T cells (∼5 × 105) from six T-cell clones and one T-cell line, specific for various gluten epitopes, were incubated with 5 × 105 irradiated DR3-DQ2 B-lymphoblastoid cells and transglutaminase 2 (TG2)-treated chymotrypsin-digested gluten (CT-gluten), appropriate antigen-peptide, medium, or phorbol 12-myristate 13-acetate (PMA)/ionomycin. Supernatant fluids were collected after 18–24 h and analyzed for IFN-γ, IL-17, IL-21, and IL-22 by Luminex assay or enzyme-linked immunosorbent assay (ELISA).
Gluten-reactive T-cell lines do not produce IL-17 after gluten stimulation
As in vitro culturing could affect cytokine production, we wanted to evaluate whether T cells in the gluten-reactive T-cell lines maintained their ability to produce IL-17. This would also allow us to study the T cells after a shorter period of culturing. We stimulated T-cell lines with TG2-treated CT-gluten, PMA/ionomycin, or anti-CD3/CD28 beads. Antigen stimulation led to an increased production of IFN-γ, and in some lines CT-gluten stimulation was found to be even more efficient than anti-CD3/CD28 and PMA/ionomycin stimulation. However, CT-gluten stimulation failed to induce production of IL-17 (Figure 2). In contrast, PMA/ionomycin and anti-CD3/CD28 stimulation led to an increased IL-17 production in most T-cell lines (although the concentrations varied considerably), with PMA/ionomycin being more efficient than anti-CD3/CD28 stimulation. In the T-cell lines, the IL-17 production originated from the CD4+ T cells (Figure 3a). Intracellular staining of cytokines in T-cell lines by flow cytometry confirmed that CT-gluten induced IFN-γ but not IL-17 production, whereas PMA/ionomycin could induce production of both cytokines. We also observed that most of the IL-17-producing T cells were double producers of IL-17 and IFN-γ (Figure 3b). Two of seven T-cell lines produced levels of IL-22 that could be detected in enzyme-linked immunosorbent assay (Figures 1 and 2). Only three out of seven of the T-cell lines had >10% of the cells with intracellular IL-22 and none of these cells stained for the DQ2-αII tetramer (Table 1). Interestingly, most of the IL-22-producing T cells in the T-cell lines also produced IFN-γ but less than half produced IL-17 (Supplementary Figure S2 online).
Gluten-reactive T-cell lines produce interferon-γ (IFN-γ) but not interleukin (IL)-17 after stimulation with transglutaminase 2 (TG2)-treated chymotrypsin-digested gluten (CT-gluten), but are able to produce IL-17 after phorbol 12-myristate 13-acetate (PMA)/ionomycin or anti-CD3/CD28 stimulation. Cells (∼5–10 × 104) of gluten-reactive T-cell lines were stimulated with anti-CD3/CD28 beads, PMA/ionomycin, or 3 × 105 DR3-DQ2-irradiated B-lymphoblastoid cells pre-pulsed with TG2-treated CT-gluten or medium. Supernatant fluids were collected after 18–24 h and analyzed for IFN-γ, IL-17, and IL-22 by a Luminex assay or enzyme-linked immunosorbent assay (ELISA).
Interleukin (IL)-17 induced by phorbol 12-myristate 13-acetate (PMA)/ionomycin stimulation of the gluten-reactive T-cell lines is produced by CD4+ cells that also produce interferon-γ (IFN-γ). (a) The gluten-reactive T-cell line KT CDE3 was stimulated with PMA/ionomycin and stained for CD4 and IL-17. (b) The gluten-reactive T-cell line KT CDE3 was stimulated with irradiated DR3-DQ2 B-lymphoblastoid cells pre-pulsed with chymotrypsin-digested gluten (CT-gluten) or medium, or PMA/ionomycin. Intracellular staining for IFN-γ and IL-17 was performed. Gating was performed on CD4+ T cells. The figure is representative of three independent experiments. Numbers in quadrants represent the percentage of cells in each.
DQ2-α-II-specific T cells, in contrast to CD4+T cells with other specificities in the T-cell lines, produce IL-21 and IFN-γ but not IL-17
The combination of PMA/ionomycin is a particularly potent activator of T cells, which could explain why this activation stimulus, but not CT-gluten, induced IL-17 production. To address this issue, we stimulated the T cells of several polyclonal T-cell lines (from celiac disease patients) reactive to the DQ2-α-II epitope, with PMA/ionomycin and combined the staining of intracellular IFN-γ, IL-17, IL-21, or IL-22 (a Th17 cytokine) with DQ2-αII tetramer staining. DQ2-CLIP2 tetramer was used as a negative control (Supplementary Figure S3 online). This allowed visualization of the DQ2-α-II-specific T cells in the T-cell line. In all IL-17- or IL-22-producing T-cell lines, the DQ2-α-II-specific T cells were able to produce IFN-γ and IL-21 but not IL-17 or IL-22. Importantly, production of IL-17 and IL-22 was observed in many of the other T cells in the T-cell lines (Figures 4 and 5, Table 1, and Supplementary Figure S1 online). One T-cell line (TCL KT CD3) with both a clear DQ2-αI and DQ2-αII tetramer-positive population was also stained for the DQ2-αI tetramer and showed the same pattern of cytokine production (data not shown). These results corroborate that the gluten-reactive T cells are double producers of IL-21 and IFN-γ, but do not produce IL-17 or IL-22.
DQ2-αII tetramer-positive T cells do not produce interleukin (IL)-17 after phorbol 12-myristate 13-acetate (PMA)/ionomycin stimulation, whereas many of the other cells in the T-cell lines produce this cytokine. The T-cell line KT CDE3 (a) and T-cell line 559.B.4.1 (b) were stimulated 18–24 h with medium or PMA/ionomycin and stained with DQ2-αII tetramer, anti-interferon-γ (IFN-γ), and anti-IL-17. The numbers in quadrants represent the percentage of cells in each, and the numbers in brackets represent the percentage of cytokine-positive cells among the cells staining with the tetramer.
DQ2-αII tetramer-positive T cells in polyclonal T-cell lines produce interleukin (IL)-21, but not IL-22. The T-cell line KT CDE3 (a) and T-cell line 559.B.4.1 (b) were stimulated 18–24 h with medium or phorbol 12-myristate 13-acetate (PMA)/ionomycin and stained with DQ2-αII tetramer, anti-IL-21, and anti-IL-22. The numbers in quadrants represent the percentage of cells in each, and the numbers in brackets represent the percentage of cytokine-positive cells among the cells staining with the tetramer.
mRNA expression of Th17 markers by gluten-reactive T cells
To confirm the data also at the mRNA level, three T-cell lines were activated by PMA/ionomycin and sorted into DQ2-αII tetramer-positive and tetramer-negative populations, followed by analysis of mRNA expression by real-time PCR. As expected, the expression of IL-17A, IL-17F, and RORC (the Th17 transcription factor) was clearly lower in the DQ2-αII tetramer-positive population when compared with the DQ2-αII-negative population (Figure 6).
DQ2-α-II-specific T cells express less interleukin (IL)-17A, IL-17F, and RAR-related orphan receptor C (RORC) mRNA than the other CD4+ T cells in the T-cell line. Three representative T-cell lines, TCL KTCD3 (upper panel), TCL 493.1.2 (middle panel), and TCL 465.1.4 (lower panel), were expanded for 1 week and the CD4+ T cells sorted into DQ2-αII tetramer-positive and -negative T cells. Expression of IL-17A, IL-17F, and RORC mRNA was normalized to the expression in tetramer-positive cells in each T-cell line (arbitrary set to one).
Cytokine production by DQ2-α-II-specific T cells in peripheral blood
To test gluten-reactive T cells that had not been subjected to possible confounding effects of in vitro culture, we investigated the cytokine production by gluten-reactive T cells that egress into the peripheral blood of treated celiac disease patients after an oral gluten challenge14, 15 using DQ2-αI and DQ2-αII tetramers (Figure 7). The cells were stimulated by PMA/ionomycin. The stimulated cells uniformly expressed the activation marker CD69 after stimulation, and approximately 8% of the cells produced either IFN-γ, IL-17, IL-5, or IL-10 (unpublished data, tested in one patient by M. Ráki). Analyzing the cytokine production in DQ2-α-I- or DQ2-α-II-specific T cells in three patients, we found that 5–8% of the cells stained for IFN-γ, whereas tenfold less cells stained for intracellular IL-17. The accuracy of these data is hampered by few events of cells that were concomitantly positive for tetramer and intracellular cytokines.
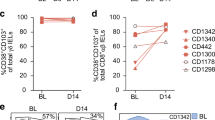
Cytokine production of gluten-reactive T cells in peripheral blood. Peripheral blood mononuclear cells (PBMCs) from three patients were isolated after oral gluten challenge. The cells were activated for 5 h with phorbol 12-myristate 13-acetate (PMA)/ionomycin, stained for surface markers, DQ2 tetramers and cytokines, and analyzed by flow cytometry. (a) Analyzed cells were gated on all CD4+ T cells. (b) Analyzed cells were gated on CD4+ T cells or DQ2-αI/DQ2-αII tetramer-positive cells as indicated. The depicted results are from one of the three patients (patient 3). Numbers in quadrants represent percentage of cells in each. (c) Percentage of tetramer-positive cells producing interferon-γ (IFN-γ) and/or interleukin (IL)-17 after activation for patients 1–3.
Discussion
We have investigated the production of IL-17, IL-21, and IL-22 by gluten-reactive CD4+ T cells in celiac disease patients. Using HLA-DQ2 tetramers, we were able to track antigen-specific cells directly in T-cell lines and in peripheral blood. We found that the gluten-reactive T cells, which are central in controlling the inflammatory reaction in celiac disease,8 produce IFN-γ and IL-21, but not IL-17.
One possible pitfall of the study may be that in vitro cultured and generated T-helper cells, and in particular Th17 cells, may lose their phenotype. This critique is relevant for our testing of the T-cell clones, but barely so for the results with the T-cell lines. Our gluten-reactive T-cell lines were cultured without antigen stimulation (except for the first 24 h, see Methods), thus providing similar growth conditions to gluten-specific and gluten-unspecific T cells. In experiments assessing cytokine production, we activated cells in the T-cell lines by PMA/ionomycin stimulation and then defined the gluten-specific T cells by tetramer staining. Many other cells with other unknown specificities were present. We found that the DQ2-α-II-specific T cells produced IFN-γ and IL-21, but not IL-17 or IL-22. Importantly, many of the other CD4+ cells of the lines produced IL-17. Our culture conditions thus allowed for expansion of IL-17-producing cells. In line with the finding at the protein level, the DQ2-α-II-specific T cells expressed less of the Th17 transcription RORC than the other T cells in the T-cell line. Moreover, the same protocol that we have used for the generation of gluten-specific T cells has been used to generate Mycobacterium avium subspecies paratuberculosis-specific T cells from intestinal biopsies of Crohn's disease patients, and many T-cell clones reactive to this bacterial antigen were found to produce IL-17.16 Finally, we find it unlikely that T cells reactive to other gluten epitopes than the DQ2-α-II epitope should produce IL-17. None of the gluten-specific T-cell clones that we tested, which cover a wide variety of gluten DQ2 epitopes, produced IL-17, and we were not able to detect IL-17 in cultures of T-cell lines stimulated with TG2-treated CT-gluten. In addition, one T-cell line stained with the DQ2-αI tetramer showed the same pattern of cytokine production as observed with the DQ2-αII tetramer.
IL-22 was found in the supernatant fluid of T-cell clones, but the cytokine was only produced by a very small percentage of cells (possibly remaining feeder cells). Only three out of seven of the T-cell lines produced significant amounts of IL-22 as assessed by flow cytometry. Co-staining for DQ2-αII tetramer-positive cells and intracellular cytokines in the IL-22-producing T-cell lines showed that the DQ2-αII tetramer-positive cells did not stain for IL-22. Taken together, the results suggest that the gluten-reactive T cells do not produce significant amounts of IL-22.
To analyze gluten-reactive T cells of celiac disease patients that have not been cultured in vitro, we attempted to stain for antigen-specific T cells with DQ2 tetramers in single-cell suspensions from fresh biopsies of celiac disease patients. This was unsuccessful as we were unable to identify a clear population of positive cells (data not shown). As an alternative approach, we tested the cytokine production of gluten-specific T cells that egress into the peripheral blood of gluten-free diet compliant celiac disease patients after a 3-day oral gluten challenge.14, 15 After stimulation with PMA/ionomycin most cells expressed the marker CD69; however, there were substantially less cells that were positive for intracellular IL-4, IFN-γ, or IL-17. For IFN-γ, we found 5–8% positive cells whereas the number of cells positive for IL-17 was approximately tenfold less. These results should be cautiously interpreted as they are based on analysis of few events in the flow cytometry analysis, and the phenotype of the cells in blood is not necessarily retained in the gut mucosa. Nevertheless, the results are in keeping with the findings of cultured T cells from biopsies indicating that gluten-reactive T cells produce IFN-γ but not IL-17.
Interestingly, we found that IFN-γ-producing gluten-specific T-cell clones also produced IL-21. IL-21 is a pleiotropic cytokine that has been shown to be produced by not only Th17 cells,17 but also follicular T helper cells and Th2 cells (shown in mice).18, 19 Our findings suggest that in humans, IFN-γ-producing T cells are able to produce significant amounts of IL-21. It has previously been shown that IL-21 is increased in the CD4+ population in celiac disease and that it helps induce production of IFN-γ and the Th1 transcription factor T-bet in the intestinal mucosa.20 Taken together with our findings, this suggests that in celiac disease, IL-21-producing gluten-reactive CD4+ T cells establish a positive feedback loop to further enhance production of IFN-γ.
In the small intestinal mucosa of untreated celiac disease patients, the mRNA for IL-17 has been found to be increased, and both T cells and non-T cells were found to contribute to increased production of IL-17.12, 13 In an organ culture system of gliadin-stimulated biopsies, increased production of IL-17 was found in both CD4+ and CD4+CD8+ T cells.13 IL-17 is produced by a large variety of different cells, and known sources of IL-17 in addition to Th17 cells include γδ T cells,21 neutrophils,22 invariant natural killer T cells, CD8+ T cells, monocytes, and eosinophils.17, 23 All these cell types are candidates for producing IL-17 in the celiac lesion. Importantly, in the previous studies the antigen specificity of the CD4+ cells producing IL-17 was not addressed. Our findings imply that gluten-reactive T cells are not producing IL-17 themselves. The gluten-reactive T cells may, however, indirectly affect IL-17 production. Neutralization of IL-21 with an antibody was found to diminish the induction of IL-17.13 Conceivably, IL-21 produced by gluten-reactive CD4+T cells can be important for induction of IL-17 by other cells.
We find it likely that IL-21 produced by gluten-reactive T cells enhances both IL-17 and IFN-γ production in the intestinal mucosa. IL-21 may have a wider role in celiac disease pathogenesis as this cytokine also targets several other cell types such as CD8+ T cells and natural killer cells. IL-21 also has a role in B-cell differentiation and plasma cell function and could possibly contribute to the production of celiac disease-associated antibodies.24 Notably, the IL-2/IL-21 locus is a genetic risk factor for celiac disease.25
In summary, in this study we have shown that the CD4+ T cells recognizing gluten peptides in the context of HLA-DQ2 do not produce IL-17. However, these CD4+ T cells, which have key roles in the inflammatory reaction, produce IL-21, a cytokine that can enhance production of both IFN-γ and IL-17 and stimulate B cells. This cytokine and its receptor are thus potential targets for therapeutic intervention in celiac disease.
Methods
Subjects
All patient material used (T-cell lines and T-cell clones) was obtained from HLA-DQ2+ (DQA1*0501/DQB1*0201; DQ2.5) celiac disease patients diagnosed according to the recommendations of the American Gastroenterological Association.26 Three of the patients (CD548, CD465, and CD559) were also DQ8+ (DQA1*03/DQB1*0302). For the experiments on peripheral blood, three treated DQ2+ celiac disease patients were recruited. The treated patients had improved clinically on gluten-free diet and histological examination showed normalized intestinal architecture. The study was approved by the regional committee for medical research ethics, and all patients gave written consent before participating.
Antigens. CT-gluten was prepared as previously described.3, 27 A synthetic 33-mer α-gliadin peptide with three glutamine residues converted to glutamate (LQLQPF(PQPELPY)3PQPQPF) was purchased from EZBiolabs (Carmel, IN). CT-gluten was treated by TG2 as previously described3, 27 and used at a final concentration of 0.1 mg ml–1. The final concentration of the 33-mer peptide was 2 μM. Peptides representing the DQ2-γ-I (YQQLPQPEQPQQSFPEQERPF) and DQ2-γ-II (GIIQPEQPAQL) epitopes were used at a final concentration of 10 μm.
T-cell lines and T-cell clones. Gluten-reactive T-cell lines were generated from duodenal mucosal specimens of treated celiac disease patients as previously described.27 It is noteworthy that in this protocol the T cells in the T-cell lines were exposed to antigen only for the first 24 h and subsequent stimulation was performed in an antigen-nonspecific manner with allogeneic feeders, phytohemagglutinin, IL-2, and IL-15. The lines were kept in culture for 2–4 weeks before testing. All T-cell lines were reactive to deamidated gluten and previously generated in our lab for other purposes. The T-cell clones were generated from T-cell lines by limited dilution as previously described.27 The T-cell clones tested were all DQ2.5 restricted, with known specificity.
T-cell assays. DR3-DQ2 Epstein–Barr virus-transformed B cells (CD 114) irradiated at 75 Gy were used as antigen-presenting cells and incubated with antigen for 18–24 h. Subsequently, T cells were added to each well. Alternatively, anti-CD3/CD28 Dynabeads (Invitrogen, Carlsbad, CA, 1 bead per cell) or a combination of PMA (50 ng ml–1) and ionomycin (2 μm) were added together with the T cells. Supernatant fluid was collected for cytokine analysis after 18–24 h of T-cell stimulation, or T cells were stimulated for 5 h and stained for intracellular cytokine production as described below.
Preparation of DQ2 tetramers. DQ2-αI and DQ2-αII tetramers were prepared as previously described.14 As negative control tetramer, DQ2 molecules covalently linked with the invariant chain peptide CLIP228 (MATPLLMQALPMGAL) (DQ2-CLIP2) was used. Tetramers were conjugated to allophycocyanin or phycoerythrin (PE) depending on the experimental setting.
Antibodies. Anti-CD4-allophycocyanin-H7 and anti-IFN-γ-fluorescein isothiocyanate antibodies were purchased from BD Biosciences (San Jose, CA). Anti-IL-17-Alexa 647, anti-CD11c-PE-Cy7, anti-CD19-PE-Cy7, anti-CD8-PE-Cy7, anti-CD14-peridinin chlorophyll protein (PerCP), anti-IL-22-PE, anti-IFN-γ-Pacific blue, and anti-IL-21-PE antibodies were purchased from eBioscience (San Diego, CA). Anti-CD3 antibody (OKT3) was conjugated to Pacific blue in our department. PerCP-conjugated anti-CD4 antibody was purchased from Diatec (Oslo, Norway).
Intracellular cytokine staining and tetramer staining of T-cell lines and clones. T-cell lines and T-cell clones were stimulated as described above in the presence of GolgiStop (BD Biosciences) and tetramer (10 μg ml–1) for 5 h. Cells were then collected and stained for CD4. The cells were fixed overnight in 1% paraformaldehyde, permeabilized for 20–30 min in 2% fetal calf serum/0.2% saponin in phosphate-buffered saline and then stained for intracellular cytokines. Cell fluorescence was analyzed on a FACSCalibur (BD) or on an LSRII flow cytometer (BD).
Analysis of cytokines in cell culture supernatants. The supernatant fluid of six T-cell clones and six T-cell lines was analyzed, after stimulation as indicated above. The concentrations of IL-17 and IFN-γ were measured by the Luminex technology using BioPlex singleplex kit (Bio-Rad, Hercules, CA) according to the manufacturer's instructions. The concentrations of IL-21 and IL-22 were measured using the human IL-21 and human IL-22 enzyme-linked immunosorbent assay Ready-SET-go! kit from eBioscience.
Characterization of gluten-specific T cells in blood in patients orally challenged with gluten. Bread challenge was performed as previously described.14, 29 In brief, patients ingested four slices of white bread daily for 3 days and blood was drawn on day 6 followed by peripheral blood mononuclear cell isolation by density gradient centrifugation. A total of 10 million peripheral blood mononuclear cells were stimulated with medium or PMA/ionomycin for 5 h and stained for tetramer, surface markers, and intracellular cytokine production (IFN-γ and IL-17) as described above. CD4+ T cells were identified as CD3+/CD8-/CD11c-/CD19- cells. Cell fluorescence was analyzed using an LSRII flow cytometer (BD).
Sorting of T cells and real-time PCR. Three gluten-reactive T-cell lines were expanded for an additional week after initial testing (as described above) to approximately 6 × 107 cells and thereafter activated for 4 h with PMA (50 ng ml–1) and ionomycin (2 μm). The cells were stained for CD4 and DQ2-αII tetramer or DQ2-CLIP (as negative control). Sorting of DQ2-αII tetramer-positive and -negative cells was performed on a BD FACS DiVa flow cytometer. RNA was isolated using the RNeasy mini-kit (Qiagen, Alameda, CA) and complementary DNA synthesized using Superscript II reverse transcriptase (Invitrogen) and oligo dT primers. Real-time PCR was performed using Taqman gene expression master mix and Taqman gene expression assays (Hs01076112_m1 (RORC), Hs99999905_m1 (GAPDH), Hs00369400_m1 (IL-17F), and Hs00174383_m1 (IL-17A)) and a StepOnePlus instrument (ABI, Carlsbad, CA). The expression of GAPDH was used as an endogenous control of input RNA in each sample and the relative expression of each gene (arbitrary units) was calculated using the ΔΔCT method.
References
Dickson, B.C., Streutker, C.J. & Chetty, R. Coeliac disease: an update for pathologists. J. Clin. Pathol. 59, 1008–1016 (2006).
Molberg, Ø. et al. Gliadin specific, HLA DQ2-restricted T cells are commonly found in small intestinal biopsies from coeliac disease patients, but not from controls. Scand. J. Immunol. 46, 103–108 (1997).
Tollefsen, S. et al. HLA-DQ2 and -DQ8 signatures of gluten T cell epitopes in celiac disease. J. Clin. Invest. 116, 2226–2236 (2006).
Qiao, S.W. et al. Refining the rules of gliadin T cell epitope binding to the disease-associated DQ2 molecule in celiac disease: importance of proline spacing and glutamine deamidation. J. Immunol. 175, 254–261 (2005).
Molberg, Ø. et al. Tissue transglutaminase selectively modifies gliadin peptides that are recognized by gut-derived T cells in celiac disease. Nat. Med. 4, 713–717 (1998).
van de Waal, Y. et al. Selective deamidation by tissue transglutaminase strongly enhances gliadin-specific T cell reactivity. J. Immunol. 161, 1585–1588 (1998).
Karell, K. et al. HLA types in celiac disease patients not carrying the DQA1⋆05-DQB1⋆02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease. Hum. Immunol. 64, 469–477 (2003).
Jabri, B. & Sollid, L.M. Mechanisms of disease: immunopathogenesis of celiac disease. Nat. Clin. Pract. Gastroenterol. Hepatol. 3, 516–525 (2006).
O'Shea, J.J. & Paul, W.E. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327, 1098–1102 (2010).
Nilsen, E.M. et al. Gluten specific, HLA-DQ restricted T cells from coeliac mucosa produce cytokines with Th1 or Th0 profile dominated by interferon gamma. Gut 37, 766–776 (1995).
Gianfrani, C. et al. Gliadin-specific type 1 regulatory T cells from the intestinal mucosa of treated celiac patients inhibit pathogenic T cells. J. Immunol. 177, 4178–4186 (2006).
Castellanos-Rubio, A. et al. TH17 (and TH1) signatures of intestinal biopsies of CD patients in response to gliadin. Autoimmunity 42, 69–73 (2009).
Monteleone, I. et al. Characterization of IL-17A-producing cells in celiac disease mucosa. J. Immunol. 184, 2211–2218 (2010).
Raki, M. et al. Tetramer visualization of gut-homing gluten-specific T cells in the peripheral blood of celiac disease patients. Proc. Natl. Acad. Sci. USA 104, 2831–2836 (2007).
Anderson, R.P. et al. T cells in peripheral blood after gluten challenge in coeliac disease. Gut 54, 1217–1223 (2005).
Olsen, I. et al. Isolation of Mycobacterium avium subspecies paratuberculosis reactive CD4T cells from intestinal biopsies of Crohn's disease patients. PLoS ONE 4, e5641 (2009).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V.K. IL-17 and Th17 cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Fazilleau, N., Mark, L., Heyzer-Williams, L.J. & Heyzer-Williams, M.G. Follicular helper T cells: lineage and location. Immunity 30, 324–335 (2009).
Suto, A. et al. Development and characterization of IL-21-producing CD4+ T cells. J. Exp. Med. 205, 1369–1379 (2008).
Fina, D. et al. Interleukin-21 contributes to the mucosal T helper cell type 1 response in celiac disease. Gut 57, 887–892 (2007).
Roark, C.L., Simonian, P.L., Fontenot, A.P., Born, W.K. & O'Brien, R.L. Gammadelta T cells: an important source of IL-17. Curr. Opin. Immunol. 20, 353–357 (2008).
Ferretti, S., Bonneau, O., Dubois, G.R., Jones, C.E. & Trifilieff, A. IL-17, produced by lymphocytes and neutrophils, is necessary for lipopolysaccharide-induced airway neutrophilia: IL-15 as a possible trigger. J. Immunol. 170, 2106–2112 (2003).
Bettelli, E., Korn, T., Oukka, M. & Kuchroo, V.K. Induction and effector functions of T(H)17 cells. Nature 453, 1051–1057 (2008).
De Nitto, D., Monteleone, I., Franze, E., Pallone, F. & Monteleone, G. Involvement of interleukin-15 and interleukin-21, two gamma-chain-related cytokines, in celiac disease. World J. Gastroenterol. 15, 4609–4614 (2009).
van Heel, D.A. et al. A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat. Genet. 39, 827–829 (2007).
Rostom, A., Murray, J.A. & Kagnoff, M.F. American Gastroenterological Association (AGA) Institute technical review on the diagnosis and management of celiac disease. Gastroenterology 131, 1981–2002 (2006).
Molberg, Ø., McAdam, S.N., Lundin, K.E. & Sollid, L.M. Studies of gliadin-specific T cells in celiac disease. In Celiac Disease. Methods and Protocols. (Marsh, M.N., ed) 105–124 ( Humana, Totowa, NJ, USA, 2000 ).
Fallang, L.E. et al. Complexes of two cohorts of CLIP peptides and HLA-DQ2 of the autoimmune DR3-DQ2 haplotype are poor substrates for HLA-DM. J. Immunol. 181, 5451–5461 (2008).
Anderson, R.P., Degano, P., Godkin, A.J., Jewell, D.P. & Hill, A.V. In vivo antigen challenge in celiac disease identifies a single transglutaminase-modified peptide as the dominant A-gliadin T-cell epitope. Nat. Med. 6, 337–342 (2000).
Acknowledgements
We thank the celiac disease patients who have donated biological material to this study. This work was supported by the Research Council of Norway and by EXTRA funds from the Norwegian Foundation for Health and Rehabilitation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
About this article
Cite this article
Bodd, M., Ráki, M., Tollefsen, S. et al. HLA-DQ2-restricted gluten-reactive T cells produce IL-21 but not IL-17 or IL-22. Mucosal Immunol 3, 594–601 (2010). https://doi.org/10.1038/mi.2010.36
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2010.36
This article is cited by
-
Interferon-Driven Immune Dysregulation in Common Variable Immunodeficiency–Associated Villous Atrophy and Norovirus Infection
Journal of Clinical Immunology (2023)
-
IL-15, gluten and HLA-DQ8 drive tissue destruction in coeliac disease
Nature (2020)
-
IL-10 signaling prevents gluten-dependent intraepithelial CD4+ cytotoxic T lymphocyte infiltration and epithelial damage in the small intestine
Mucosal Immunology (2019)
-
Pathogenesis of Enteropathy-Associated T Cell Lymphoma
Current Hematologic Malignancy Reports (2018)
-
Macrophage-mediated gliadin degradation and concomitant IL-27 production drive IL-10- and IFN-γ-secreting Tr1-like-cell differentiation in a murine model for gluten tolerance
Mucosal Immunology (2017)