Abstract
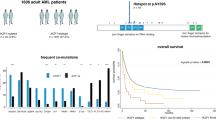
RUNX1-mutated acute myeloid leukemia (AML) show a distinct pattern of genetic abnormalities and an adverse prognosis. We analyzed the impact of multiple RUNX1 mutations and RUNX1 wild-type (WT) loss in 467 AML with RUNX1 mutations (mut): (1) RUNX1 WT loss (n=53), (2) >1 RUNX1mut (n=94) and (3) 1 RUNX1mut (n=323). In 1 RUNX1mut, +8 was most frequent, whereas in WT loss +13 was the most abundant trisomy (+8: 66% vs 31%, P=0.022; +13: 15% vs 62%, P<0.001). Analyses of 28 genes in 163 selected cases revealed SRSF2 (39%), ASXL1 (36%), DNMT3A (19%), IDH2 (17%) and SF3B1 (17%) as most frequently mutated genes. RUNX1 WT loss showed a higher frequency of ASXL1mut compared with the other cases (50% vs 29%, P=0.009). Median overall survival (OS) in the total cohort was 14 months. WT loss (OS: 5 months) and >1 RUNX1mut (14 months) showed an adverse impact on prognosis compared with 1 RUNX1mut (22 months; P=0.002 and 0.048, respectively). Mutations in ASXL1 and ⩾2 additional mutations correlated with shorter OS (10 vs 18 months, P=0.028; 12 vs 20 months, P=0.017). Thus, the number of RUNX1mut, RUNX1 WT loss and the number and type of additional mutations is biologically and clinically relevant.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Grimwade D, Hills RK, Moorman AV, Walker H, Chatters S, Goldstone AH et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood 2010; 116: 354–365.
Döhner H, Weisdorf DJ, Bloomfield CD . Acute myeloid leukemia. N Engl J Med 2015; 373: 1136–1152.
Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, Robertson A et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368: 2059–2074, Cancer Genome Atlas Research Network.
Döhner H, Gaidzik VI . Impact of genetic features on treatment decisions in AML. Hematology Am Soc Hematol Educ Program 2011; 2011: 36–42.
Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H et al. WHO Classification of Tumours of Haematopoietic and Lympoid Tissues. IARC: Lyon, France, 2008.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM et al. The 2016 revision to the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia. Blood 2016; 127: 2391–2405.
Okuda T, Nishimura M, Nakao M, Fujita Y . RUNX1/AML1: a central player in hematopoiesis. Int J Hematol 2001; 74: 252–257.
Berardi MJ, Sun C, Zehr M . The Ig fold of the core binding factor alpha Runt domain is a member of a family of structurally and functionally related Ig-fold DNA-binding domains. Structure 1999; 7: 1247–1256.
Bravo J, Li Z, Speck NA, Warren AJ . The leukemia-associated AML1 (Runx1)-CBF beta complex functions as a DNA-induced molecular clamp. Nat Struct Biol 2001; 8: 371–378.
Bowers SR, Calero-Nieto FJ, Valeaux S, Fernandez-Fuentes N, Cockerill PN . 'Runx1 binds as a dimeric complex to overlapping Runx1 sites within a palindromic element in the human GM-CSF enhancer'. Nucleic Acids Res 2010; 38: 6124–6134.
Gaidzik VI, Bullinger L, Schlenk RF, Zimmermann AS, Röck J, Paschka P et al. RUNX1 mutations in acute myeloid leukemia: results from a comprehensive genetic and clinical analysis from the AML Study Group. J Clin Oncol 2011; 29: 1364–1372.
Gaidzik VI, Teleanu V, Papaemmanuil E, Weber D, Paschka P, Hahn J et al. RUNX1 mutations in acute myeloid leukemia are associated with distinct clinico-pathologic and genetic features. Leukemia 2016; 30: 2160–2168.
Osato M, Asou N, Abdalla E, Hoshino K, Yamasaki H, Okubo T et al. Biallelic and heterozygous point mutations in the runt domain of the AML1/PEBP2alphaB gene associated with myeloblastic leukemias. Blood 1999; 93: 1817–1824.
Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia 2014; 28: 241–247.
Grossmann V, Kern W, Harbich S, Alpermann T, Jeromin S, Schnittger S et al. Prognostic relevance of RUNX1 mutations in T-cell acute lymphoblastic leukemia. Haematologica 2011; 96: 1874–1877.
Gelsi-Boyer V, Trouplin V, Adélaïde J, Aceto N, Remy V, Pinson S et al. Genome profiling of chronic myelomonocytic leukemia: frequent alterations of RAS and RUNX1 genes. BMC Cancer 2008; 8: 299.
Imai Y, Kurokawa M, Izutsu K, Hangaishi A, Takeuchi K, Maki K et al. Mutations of the AML1 gene in myelodysplastic syndrome and their functional implications in leukemogenesis. Blood 2000; 96: 3154–3160.
Owen CJ, Toze CL, Koochin A, Forrest DL, Smith CA, Stevens JM et al. Five new pedigrees with inherited RUNX1 mutations causing familial platelet disorder with propensity to myeloid malignancy (FPD/AML). Blood 2008; 112: 4639–4645.
Haferlach T, Stengel A, Eckstein S, Perglerová K, Alpermann T, Kern W et al. The new provisional WHO entity 'RUNX1 mutated AML' shows specific genetics but no prognostic influence of dysplasia. Leukemia 2016; 30: 2109–2112.
Quentin S, Cuccuini W, Ceccaldi R, Nibourel O, Pondarre C, Pagès MP et al. Myelodysplasia and leukemia of Fanconi anemia are associated with a specific pattern of genomic abnormalities that includes cryptic RUNX1/AML1 lesions. Blood 2011; 117: e161–e170.
Skokowa J, Steinemann D, Katsman-Kuipers JE, Zeidler C, Klimenkova O, Unalan M et al. Cooperativity of RUNX1 and CSF3R mutations in severe congenital neutropenia: a unique pathway in myeloid leukemogenesis. Blood 2014; 123: 2229–2237.
Harada H, Harada Y, Tanaka H, Kimura A, Inaba T . Implications of somatic mutations in the AML1 gene in radiation-associated and therapy-related myelodysplastic syndrome/acute myeloid leukemia. Blood 2003; 101: 673–680.
Jongmans MC, Kuiper RP, Carmichael CL, Wilkins EJ, Dors N, Carmagnac A et al. Novel RUNX1 mutations in familial platelet disorder with enhanced risk for acute myeloid leukemia: clues for improved identification of the FPD/AML syndrome. Leukemia 2010; 24: 242–246.
Song WJ, Sullivan MG, Legare RD, Hutchings S, Tan X, Kufrin D et al. Haploinsufficiency of CBFA2 causes familial thrombocytopenia with propensity to develop acute myelogenous leukaemia. Nat Genet 1999; 23: 166–175.
Haferlach T, Schoch C, Löffler H, Gassmann W, Kern W, Schnittger S et al. Morphologic dysplasia in de novo acute myeloid leukemia (AML) is related to unfavorable cytogenetics but has no independent prognostic relevance under the conditions of intensive induction therapy: results of a multiparameter analysis from the German AML Cooperative Group studies. J Clin Oncol 2003; 21: 256–265.
Kern W, Voskova D, Schoch C, Hiddemann W, Schnittger S, Haferlach T . Determination of relapse risk based on assessment of minimal residual disease during complete remission by multiparameter flow cytometry in unselected patients with acute myeloid leukemia. Blood 2004; 104: 3078–3085.
Bene MC, Castoldi G, Knapp W, Ludwig WD, Matutes E, Orfao E et al. Proposals for the immunological classification of acute leukemias. European Group for the Immunological Characterization of Leukemias (EGIL). Leukemia 1995; 9: 1783–1786.
Schoch C, Schnittger S, Bursch S, Gerstner D, Hochhaus A, Berger U et al. Comparison of chromosome banding analysis, interphase- and hypermetaphase-FISH, qualitative and quantitative PCR for diagnosis and for follow-up in chronic myeloid leukemia: a study on 350 cases. Leukemia 2002; 16: 53–59.
McGowan-Jordan J, Simons A, Schmid M . An International System for Human Cytogenetic Nomenclature. Karger: Basel, New York, 2016.
MacDonald JR, Ziman R, Yuen RK, Feuk L, Scherer SW . The Database of Genomic Variants: a curated collection of structural variation in the human genome. Nucleic Acids Res 2014; 42: D986–D992.
Grimwade D, Ivey A, Huntly BJ . Molecular landscape of acute myeloid leukemia in younger adults and its clinical relevance. Blood 2016; 127: 29–41.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med 2016; 374: 2209–2221.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P et alChronic Myeloid Disorders Working Group of the International Cancer Genome Consortium.. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 2013; 122: 3616–3627.
Bejar R, Stevenson KE, Caughey BA, Abdel-Wahab O, Steensma DP, Galili N et al. Validation of a prognostic model and the impact of mutations in patients with lower-risk myelodysplastic syndromes. J Clin Oncol 2012; 30: 3376–3382.
Milosevic JD, Puda A, Malcovati L, Berg T, Hofbauer M, Stukalov A et al. Clinical significance of genetic aberrations in secondary acute myeloid leukemia. Am J Hematol 2012; 87: 1010–1016.
Dicker F, Haferlach C, Sundermann J, Wendland N, Weiss T, Kern W et al. Mutation analysis for RUNX1, MLL-PTD, FLT3-ITD, NPM1 and NRAS in 269 patients with MDS or secondary AML. Leukemia 2010; 24: 1528–1532.
Mesa RA, Hanson CA, Ketterling RP, Schwager S, Knudson RA, Tefferi A . Trisomy 13: prevalence and clinicopathologic correlates of another potentially lenalidomide-sensitive cytogenetic abnormality. Blood 2009; 113: 1200–1201.
Dicker F, Haferlach C, Kern W, Haferlach T, Schnittger S . Trisomy 13 is strongly associated with AML1/RUNX1 mutations and increased FLT3 expression in acute myeloid leukemia. Blood 2007; 110: 1308–1316.
Acknowledgements
We thank all co-workers at the MLL Munich Leukemia Laboratory for approaching together many aspects in the field of leukemia diagnostics and research by their dedicated work. We would like to thank all physicians for providing and caring for patients as well as collecting data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
CH, WK and TH declare part ownership of Munich Leukemia Laboratory (MLL). AS, MM and NN are employed by the MLL Munich Leukemia Laboratory. KP is employed by MLL2.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Stengel, A., Kern, W., Meggendorfer, M. et al. Number of RUNX1 mutations, wild-type allele loss and additional mutations impact on prognosis in adult RUNX1-mutated AML. Leukemia 32, 295–302 (2018). https://doi.org/10.1038/leu.2017.239
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.239
This article is cited by
-
Genomic and clinical findings in myeloid neoplasms with PDGFRB rearrangement
Annals of Hematology (2022)
-
Allogeneic stem cell transplantation for AML patients with RUNX1 mutation in first complete remission: a study on behalf of the acute leukemia working party of the EBMT
Bone Marrow Transplantation (2021)
-
RUNX1 mutations in blast-phase chronic myeloid leukemia associate with distinct phenotypes, transcriptional profiles, and drug responses
Leukemia (2021)
-
Spectrum of abnormalities and clonal transformation in germline RUNX1 familial platelet disorder and a genomic comparative analysis with somatic RUNX1 mutations in MDS/MPN overlap neoplasms
Leukemia (2020)
-
Molecular landscape and targeted therapy of acute myeloid leukemia
Biomarker Research (2018)