Abstract
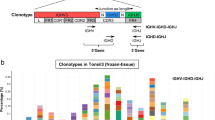
The immunoglobulin heavy-chain variable region gene (IgHV) mutational status is considered the gold standard of prognostication in chronic lymphocytic leukemia (CLL) and is currently determined by Sanger sequencing that allows the analysis of the major clone. Using next-generation sequencing (NGS), we sequenced the IgHV gene from two independent cohorts: (A) 270 consecutive patient samples obtained at diagnosis and (B) 227 patients from the UK ARCTIC-AdMIRe clinical trials. Using complementary DNA from purified CD19+CD5+ cells, we demonstrate the presence of multiple rearrangements in independent experiments and showed that 24.4% of CLL patients express multiple productive clonally unrelated IgHV rearrangements. On the basis of IgHV-NGS subclonal profiles, we defined five different categories: patients with (a) multiple hypermutated (M) clones, (b) 1 M clone, (c) a mix of M-unmutated (UM) clones, (d) 1 UM clone and (e) multiple UM clones. In population A, IgHV-NGS classification stratified patients into five different subgroups with median treatment-free survival (TFS) of >280(a), 131(b), 94(c), 29(d), 15(e) months (P<0.0001) and a median OS of >397(a), 292(b), 196(c), 137(d) and 100(e) months (P<0.0001). In population B, the poor prognosis of multiple UM patients was confirmed with a median TFS of 2 months (P=0.0038). In conclusion, IgHV-NGS highlighted one quarter of CLL patients with multiple productive IgHV subclones and improves disease stratification and raises important questions concerning the pre-leukemic cellular origin of CLL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hamblin T . Chronic lymphocytic leukaemia: one disease or two? Ann Hematol 2002; 81: 299–303.
Schuh A, Becq J, Humphray S, Alexa A, Burns A, Clifford R et al. Monitoring chronic lymphocytic leukemia progression by whole genome sequencing reveals heterogeneous clonal evolution patterns. Blood 2012; 120: 4191–4196.
Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 2013; 152: 714–726.
Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK . Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 1999; 94: 1848–1854.
Damle RN, Wasil T, Fais F, Ghiotto F, Valetto A, Allen SL et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood 1999; 94: 1840–1847.
Tobin G, Thunberg U, Johnson A, Thorn I, Soderberg O, Hultdin M et al. Somatically mutated Ig V(H)3-21 genes characterize a new subset of chronic lymphocytic leukemia. Blood 2002; 99: 2262–2264.
Thorselius M, Krober A, Murray F, Thunberg U, Tobin G, Buhler A et al. Strikingly homologous immunoglobulin gene rearrangements and poor outcome in VH3-21-using chronic lymphocytic leukemia patients independent of geographic origin and mutational status. Blood 2006; 107: 2889–2894.
Agathangelidis A, Darzentas N, Hadzidimitriou A, Brochet X, Murray F, Yan XJ et al. Stereotyped B-cell receptors in one-third of chronic lymphocytic leukemia: a molecular classification with implications for targeted therapies. Blood 2012; 119: 4467–4475.
Bomben R, Dal Bo M, Capello D, Forconi F, Maffei R, Laurenti L et al. Molecular and clinical features of chronic lymphocytic leukaemia with stereotyped B cell receptors: results from an Italian multicentre study. Br J Haematol 2009; 144: 492–506.
Ghia P, Stamatopoulos K, Belessi C, Moreno C, Stilgenbauer S, Stevenson F et al. ERIC recommendations on IGHV gene mutational status analysis in chronic lymphocytic leukemia. Leukemia 2007; 21: 1–3.
Krober A, Seiler T, Benner A, Bullinger L, Bruckle E, Lichter P et al. V(H) mutation status, CD38 expression level, genomic aberrations, and survival in chronic lymphocytic leukemia. Blood 2002; 100: 1410–1416.
Austen B, Powell JE, Alvi A, Edwards I, Hooper L, Starczynski J et al. Mutations in the ATM gene lead to impaired overall and treatment-free survival that is independent of IGVH mutation status in patients with B-CLL. Blood 2005; 106: 3175–3182.
Burger JA, Keating MJ, Wierda WG, Hartmann E, Hoellenriegel J, Rosin NY et al. Safety and activity of ibrutinib plus rituximab for patients with high-risk chronic lymphocytic leukaemia: a single-arm, phase 2 study. Lancet Oncol 2014; 15: 1090–1099.
Seifert M, Sellmann L, Bloehdorn J, Wein F, Stilgenbauer S, Durig J et al. Cellular origin and pathophysiology of chronic lymphocytic leukemia. J Exp Med 2012; 209: 2183–2198.
Chiorazzi N, Ferrarini M . Cellular origin(s) of chronic lymphocytic leukemia: cautionary notes and additional considerations and possibilities. Blood 2011; 117: 1781–1791.
Damm F, Mylonas E, Cosson A, Yoshida K, Della Valle V, Mouly E et al. Acquired initiating mutations in early hematopoietic cells of CLL patients. Cancer Discov 2014; 4: 1088–1101.
Kikushige Y, Ishikawa F, Miyamoto T, Shima T, Urata S, Yoshimoto G et al. Self-renewing hematopoietic stem cell is the primary target in pathogenesis of human chronic lymphocytic leukemia. Cancer Cell 2011; 20: 246–259.
Plevova K, Francova HS, Burckova K, Brychtova Y, Doubek M, Pavlova S et al. Multiple productive immunoglobulin heavy chain gene rearrangements in chronic lymphocytic leukemia are mostly derived from independent clones. Haematologica 2014; 99: 329–338.
Visco C, Moretta F, Falisi E, Facco M, Maura F, Novella E et al. Double productive immunoglobulin sequence rearrangements in patients with chronic lymphocytic leukemia. Am J Hematol 2013; 88: 277–282.
Rassenti LZ, Kipps TJ . Lack of allelic exclusion in B cell chronic lymphocytic leukemia. J Exp Med 1997; 185: 1435–1445.
Klinger M, Zheng J, Elenitoba-Johnson KSJ, Perkins SL, Faham M, Bahler DW . Next-generation IgVH sequencing CLL-like monoclonal B-cell lymphocytosis reveals frequent oligoclonality and ongoing hypermutation. Leukemia 2016; 30: 1055–1061.
Kriangkum J, Motz SN, Mack T, Beiggi S, Baigorri E, Kuppusamy H et al. Single-cell analysis and next-generation immuno-sequencing show that multiple clones persist in patients with chronic lymphocytic leukemia. PLoS One 2015; 10: e0137232.
Stamatopoulos B, Van Damme M, Crompot E, Dessars B, El Housni H, Mineur P et al. Opposite prognostic significance of cellular and serum circulating microRNA-150 in chronic lymphocytic leukemia patients. Mol Med 2015; 21: 123–133.
Stamatopoulos B, Meuleman N, Haibe-Kains B, Saussoy P, Van Den Neste E, Michaux L et al. microRNA-29c and microRNA-223 down-regulation has in vivo significance in chronic lymphocytic leukemia and improves disease risk stratification. Blood 2009; 113: 5237–5245.
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000; 343: 1910–1916.
Blachly JS, Ruppert AS, Zhao W, Long S, Flynn J, Flinn I et al. Immunoglobulin transcript sequence and somatic hypermutation computation from unselected RNA-seq reads in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2015; 112: 4322–4327.
Langerak AW, Davi F, Ghia P, Hadzidimitriou A, Murray F, Potter KN et al. Immunoglobulin sequence analysis and prognostication in CLL: guidelines from the ERIC review board for reliable interpretation of problematic cases. Leukemia 2011; 25: 979–984.
Chen C, Nagy Z, Prak EL, Weigert M . Immunoglobulin heavy chain gene replacement: a mechanism of receptor editing. Immunity 1995; 3: 747–755.
Mimmi S, Vecchio E, Iaccino E, Rossi M, Lupia A, Albano F et al. Evidence of shared epitopic reactivity among independent B-cell clones in chronic lymphocytic leukemia patients. Leukemia, 2016; 30: 2419–2422.
Fischer K, Bahlo J, Fink AM, Goede V, Herling CD, Cramer P et al. Long-term remissions after FCR chemoimmunotherapy in previously untreated patients with CLL: updated results of the CLL8 trial. Blood 2016; 127: 208–215.
Herishanu Y, Katz BZ, Lipsky A, Wiestner A . Biology of chronic lymphocytic leukemia in different microenvironments: clinical and therapeutic implications. Hematol Oncol Clin North Am 2013; 27: 173–206.
Burger JA . No cell is an island unto itself: the stromal microenvironment in chronic lymphocytic leukemia. Leuk Res 2007; 31: 887–888.
Acknowledgements
This research was supported by the 'Wallonie-Bruxelles International World' (WBI World), the IRIS-Recherche fund, and the Bekales Foundation and 'Les Amis de l’Institut Bordet'. BS is supported by a Télévie grant (FRS-FNRS—Fonds de la Recherche Scientifique—FNRS). AS, PR, AB, DV, PA and JM are supported by the NIHR Biomedical Research Centre (Oxford).
Author contributions
BS and AT were the principal investigators and took primary responsibility for the paper; BS and AT performed research and wrote the manuscript. DB, HD and JM provided help for the next-generation sequencing experiments. DV, LL and PA performed stereotyped receptor analysis, coverage checking and verified the statistics. TS performed the bioinformatics validation. RC, PR and AB participated to the scientific discussion and revised the manuscript; AS supervised and revised the manuscript and the research.
Disclaimer
The views expressed are those of the authors and not necessarily those of the Department of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Stamatopoulos, B., Timbs, A., Bruce, D. et al. Targeted deep sequencing reveals clinically relevant subclonal IgHV rearrangements in chronic lymphocytic leukemia. Leukemia 31, 837–845 (2017). https://doi.org/10.1038/leu.2016.307
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.307
This article is cited by
-
Immunoglobulin gene sequence analysis in chronic lymphocytic leukemia: the 2022 update of the recommendations by ERIC, the European Research Initiative on CLL
Leukemia (2022)
-
Nanopore sequencing approach for immunoglobulin gene analysis in chronic lymphocytic leukemia
Scientific Reports (2021)
-
Immunoglobulin gene analysis in chronic lymphocytic leukemia in the era of next generation sequencing
Leukemia (2020)
-
IgCaller for reconstructing immunoglobulin gene rearrangements and oncogenic translocations from whole-genome sequencing in lymphoid neoplasms
Nature Communications (2020)
-
Exome sequencing of the TCL1 mouse model for CLL reveals genetic heterogeneity and dynamics during disease development
Leukemia (2019)