Abstract
Apart from its unique histopathological appearance with rare tumor cells embedded in an inflammatory background of bystander cells, classical Hodgkin lymphoma (cHL) is characterized by an unusual activation of a broad range of signaling pathways involved in cellular activation. This includes constitutive high-level activity of nuclear factor-κB (NF-κB), Janus kinase/signal transducer and activator of transcription (JAK/STAT), activator protein-1 (AP-1) and interferon regulatory factor (IRF) transcription factors (TFs) that are physiologically only transiently activated. Here, we demonstrate that inactivation of the putative ubiquitin E3-ligase PDLIM2 contributes to this TF activation. PDLIM2 expression is lost at the mRNA and protein levels in the majority of cHL cell lines and Hodgkin and Reed–Sternberg (HRS) cells of nearly all cHL primary samples. This loss is associated with PDLIM2 genomic alterations, promoter methylation and altered splicing. Reconstitution of PDLIM2 in HRS cell lines inhibits proliferation, blocks NF-κB transcriptional activity and contributes to cHL-specific gene expression. In non-Hodgkin B-cell lines, small interfering RNA-mediated PDLIM2 knockdown results in superactivation of TFs NF-κB and AP-1 following phorbol 12-myristate 13-acetate (PMA) stimulation. Furthermore, expression of PDLIM2 is lost in anaplastic large cell lymphoma (ALCL) that shares key biological aspects with cHL. We conclude that inactivation of PDLIM2 is a recurrent finding in cHL and ALCL, promotes activation of inflammatory signaling pathways and thereby contributes to their pathogenesis.
Similar content being viewed by others
Introduction
Classical Hodgkin lymphoma (cHL) is characterized by a unique histopathological appearance in the affected lymph nodes demonstrating only a minority of tumor cells, the so-called Hodgkin and Reed–Sternberg (HRS) cells, that are embedded in an abundant inflammatory microenvironment.1 It is assumed that HRS cells attract these inflammatory bystander cells by high-level production of cytokines and chemokines, and that the interaction between HRS cells and their surrounding cells supports their growth, survival and immune escape.1, 2, 3 The abundant production of inflammatory mediators by HRS cells reflects their highly activated phenotype. In line with such a phenotype, HRS cells show constitutive activation of a plethora of transcription factors (TFs) that are physiologically only transiently activated, and that are implicated in cellular activation and immediate-early gene induction. Among these TFs, nuclear factor-κB (NF-κB), signal transducer and activator of transcription (STAT) 3, 5 and 6, activator protein-1 (AP-1) and interferon regulatory factor-5 (IRF5) play key roles in HL pathogenesis.4, 5, 6, 7, 8
The mechanisms leading to activation of these TFs in HRS cells are complex and affect various layers of control of respective pathway components.1 These include deleterious mutations of negative regulators of receptor-proximal activation modules,9 of negative regulators of more downstream located signaling components (for example, inhibitor of NF-κB (IκB)α mutations in the NF-κB pathway),10, 11 genomic alterations of respective TFs12, 13 and the constitutive activation of upstream kinases.14 The strength and plethora of pathway activation in HRS cells is exceptional among lymphoid malignancies. Interestingly, despite their complex transcriptional alterations including long-terminal repeat activation,15, 16 HRS cells demonstrate a surprisingly constant phenotype. Our recent data on the pivotal role of IRF5 together with NF-κB regarding the orchestration of the HL-specific gene expression program7 suggest that only a limited number of pathogenic events coordinate the cHL phenotype.
Given the number of high-level activated TFs in HRS cells involved in immediate-early gene regulation, we postulated that alterations of the ubiquitin system contribute to their simultaneous activation. Based on a literature search in combination with gene expression data of HRS and non-Hodgkin (NH) B cell-derived cell lines,17 we focused on PDLIM2 (also known as mystique or SLIM).18, 19, 20 PDLIM2 has been shown to act as ubiquitin E3-ligase inducing inactivation and degradation of TFs.19, 21 In addition, ubiquitin-independent mechanisms of PDLIM2 have been described.19 By these mechanisms, PDLIM2 controls the activity of TFs, such as NF-κB-p65, STAT1 and STAT3,19, 21, 22, 23 that are highly activated in HRS cells. Furthermore, alterations of PDLIM2 gene expression have been observed in malignancies,23, 24, 25 and the PDLIM2 gene locus in 8p21.3 is in the minimal commonly deleted region in 8p21.3 in several B-cell non-Hodgkin lymphomas.26, 27
Here, we describe an inactivation of the putative ubiquitin E3-ligase PDLIM2 as unifying defect of HRS cells. PDLIM2 loss of expression is associated with various mechanisms, including genomic alterations and promoter DNA methylation. Functionally, loss of PDLIM2 promotes growth of HRS cells, facilitates the activation of inflammatory TFs and results in deregulation of differentially expressed cHL-associated genes, suggesting a pathogenic role in cHL. Furthermore, we demonstrate loss of PDLIM2 expression in anaplastic large cell lymphoma (ALCL) that shares key biological aspects with cHL.
Materials and methods
Cell lines and culture conditions
HRS (L428, L1236, KM-H2, L591 (EBV+), U-HO1, SUP-HD1, HDLM-2, L540, L540Cy), pro-B lymphoblastic leukemia (Reh), Burkitt lymphoma (Namalwa, BL-60, BJAB), diffuse large B-cell lymphoma (SU-DHL-4), ALCL (K299, SU-DHL-1, DEL, JB6, all anaplastic lymphoma kinase (ALK) positive; Mac-1, Mac-2A, FE-PD, DL40, all ALK negative), T-cell leukemia-derived (Jurkat, KE-37, Molt-14, H9) and HEK293 cells were cultured as previously described.28 Cells were electroporated in OPTI-MEM I using Gene-Pulser Xcell (Bio-Rad, Munich, Germany) with 950 μF and 0.18 kV (L540Cy, HEK293) or 500 μF and 0.3 kV (Reh, L591). Transfection efficiency was determined by pEGFP-N3 (Clontech Laboratories, Mountain View, CA, USA) co-transfection. L591 and L540Cy cells were transfected with 40 μg of a pcDNA3-PDLIM2 transcript variant 2 (tv 2) construct along with 10 μg pEGFP-N3. GFP+ cells were enriched by fluorescence-activated cell sorting. Reh cells were transfected with two pSUPER plasmid29-based siPDLIM2 expression plasmids (each 20 μg) or 40 μg of scrambled small interfering RNA (siRNA) controls along with 10 μg pEGFP-N3 and enriched as previously described.28 For reporter assays, L591, L540Cy or Reh cells were transfected with 1 μg of 6xκBLuc or 15 μg 5xTRE reporter constructs, together with 50, 25 or 200 ng pRL-TKLuc as internal control. Where indicated, cells were additionally transfected with 40 μg pcDNA3-IκBαΔN or pcDNA3-PDLIM2 constructs or controls. The ratio of the pRL-TKLuc and 6xκBLuc or 15 μg 5xTRE luciferase activities was determined (Dual luciferase kit; Promega, Mannheim, Germany). Where indicated, cells were stimulated with 200 ng/ml phorbol 12-myristate 13-acetate (PMA; Sigma-Aldrich, Taufkirchen, Germany) or treated with 25 μg/ml cycloheximide (Sigma-Aldrich).
DNA constructs
pcDNA3-IκBαΔN and luciferase constructs pGL3-6xκBLuc and pGL3-5xTRE-TATA (kindly provided by Peter Angel, Heidelberg, Germany) have been previously described.30, 31 PDLIM2 plasmids encoding tv 1 (NM_176871.2; 1100 bp), variant 2 (NM_021630.4; 1058 bp) or PDLIM2_ΔLIM were generated by cloning corresponding sequences through 5′ BamHI and 3′ XhoI (tv 1; ΔLIM) or EcoRI (tv 2; varL1236) into pcDNA3.1(+) (Invitrogen, Karlsruhe, Germany). Primers used for amplification of PDLIM2 variants were PDLIM2 BamHI s 5′-ACACGGATCCTCCACCATGGCGTTGACGGTGGATGTG-3′ and PDLIM2 XhoI as 5′-ACACGAATTCTCACCAGCTTGGCTGGATGGG-3′ (tv 1) or PDLIM2 EcoRI as 5′-ACACGAATCCTCAGGCCCGAGAGCTGAG-3′ (tv 2). PDLIM2_ΔLIM was amplified according to Tanaka et al.22 using primers PDLIM2 BamHI s and PDLIM2_ΔLIM XhoI as 5′-GCCTCGAGTCAGGTGGCCAGGGCCCTG-3′. The truncated PDLIM2 variant from L1236 cells was amplified using primers PDLIM2_varL1236 BamHI s 5′-GCGGATCCGCGTTGACGGTGGATGTGGCC-3′ and PDLIM2 EcoRI as. The, in comparison with the wt PDLIM2 amplificate, smaller product was excised after gel electrophoresis and cloned into pcDNA3 with N-terminal FLAG. Sequences for the siPDLIM2 constructs were selected from the Dharmacon PDLIM2 ON-TARGETplus SMARTpool (J-010731; Dharmacon, Lafayette, CO, USA). Target sequences 5′-GGACAGCTCCTTGGAAGTG-3′ or 5′-ACATAATCGTGGCCATCAA-3′ were cloned through BglII and HindIII restriction sites into pSUPER.29 The scrambled siRNA construct has been previously described.32 All DNA constructs were verified by sequencing.
RNA preparation and reverse transcriptase-PCR analyses
Total RNA preparation, complementary DNA synthesis and semiquantitative PCR and real-time quantitative PCR analyses were performed as previously described.17, 28 Primers are listed in Supplementary Table 1. All PCR products were verified by sequencing.
Analysis of apoptosis and proliferation
The percentage of viable and apoptotic cells was determined by Annexin V-fluorescein isothiocyanate/propidium iodide double staining (Bender MedSystems, Vienna, Austria) and flow cytometry as previously described.7 Proliferation of cells was determined by measurement of [3H]-thymidine incorporation using standard protocols.
Western blot analyses
Whole-cell and nuclear extract preparation and western blotting were performed as described5 with 30 μg whole-cell or 20 μg nuclear extracts. Primary antibodies were: PDLIM2 (sc-79988), IκBα (sc-371; both from Santa Cruz, Heidelberg, Germany), PDLIM2 (ab68220; Abcam, Cambridge, UK) and β-actin (A5316, Sigma-Aldrich).
Immunohistochemistry
Immunohistochemical stainings of 20 primary cHL, 10 follicular lymphoma, 15 diffuse large B-cell lymphoma and 10 Burkitt lymphoma cases were performed as previously described.33 Primary antibodies (PDLIM2, PA5-31484, Thermo Scientific, Darmstadt, Germany; PDLIM2, ab68220, Abcam; IκBα, sc-371, Santa Cruz) were applied in a dilution of 1:100 (PDLIM2 antibodies) or 1:500 (IκBα) overnight.
Interphase cytogenetics
Fluorescence in situ hybridization and FICTION (fluorescence immunophenotyping and interphase cytogenetic as a tool for investigation of neoplasia) techniques were used to analyze copy number changes of the PDLIM2 gene in cHL cell lines and primary samples on cryosections from the Department of Pathology, Hematopathology Section and Lymph Node Registry, University Hospital Schleswig-Holstein, Kiel, Germany. The anonymized use of these patient samples followed the regulations approved by the institutional review board of the Medical Faculty of the Christian-Albrechts-University Kiel (decision D447/10). For FICTION, the procedure included pretreatment with anti-CD30 antibody (BerH2 generated in-house34) and detection with Alexa-594-conjugated antibody (Molecular Probes, Leiden, The Netherlands). Two hybridization probes were prepared as previously described.35 They contained DNA from the 157 423 kb large BAC RP11-382J24 (NCBI AC087854.11; chr8:22 225 035–22 382 458 genomic position based on the BAC and sequences mapped to the hg19 assembly) containing the SLC39A14 and PPP3CC genes (Invitrogen, GmbH, Darmstadt, Germany) labeled in SpectrumGreen (Abbott/Vysis, Downers Grove, IL, USA) located in the minimally deleted region in B-cell lymphomas 8p21.3(ref. 27) ∼55 kbp telomeric of the PDLIM2 gene. This probe was combined either with the centromeric CEP10 SpectrumAqua probe (assay 1) or the centromeric CEP8 SpectrumOrange probe (assay 2) (both Abbott/Vysis) that served as internal controls. Both assays were used independently to the analyzed samples. For evaluation, a Zeiss fluorescence microscope (Göttingen, Germany) was used. For each case, at least 5 large CD30+ cells were evaluated independently by two observers. We estimated the ploidy levels of the analyzed cases by taking median signal numbers for the centromeric probes CEP6,(ref. 9) CEP10 (this study), CEP16(ref. 15) and CEP17.36 For detection of PDLIM2 deletions, the signal number of the RP11-382J24 probe (PDLIM2) was compared with the ploidy of the respective case separately for both used assays (Supplementary Table 2). Lower number of RP11-382J24 probe signals than the ploidy in at least 30% of the nuclei of the case observed in both assays was indicative for a deletion.
Bisulfite pyrosequencing
Bisulfite pyrosequencing of two PDLIM2 promoter regions called ‘promoter 1’ (P1; promoter region of NM_021630) and ‘promoter 2’ (P2; promoter region of NM_198042, NM_176871) and a previously analyzed region (Qu et al.24, 25 called ‘pub. reg.’) was performed according to standard protocols with few modifications.15 Briefly, genomic DNA was bisulfite converted using the EpiTect Bisulfite Conversion Kit (Qiagen, Hilden, Germany). PCR amplification was performed with locus-specific primers with one primer biotinylated at the 5′ end. For PDLIM2 amplification reactions, PyroMark PCR Kit (Qiagen) was used. After PCR analyses, single strands were prepared followed by a denaturation step at 85 °C for 2 min and final sequencing primer hybridization. Pyrosequencing was performed using Pyrosequencer ID and DNA methylation analysis software Pyro Q-CpG 1.0.9 (Biotage, Uppsala, Sweden) that was also used to evaluate the ratio T:C (mC:C) at CpG sites analyzed. All assays were optimized and validated using commercially available completely methylated DNA (Millipore, Darmstadt, Germany) and DNA isolated from peripheral blood of 10 healthy male and female controls, respectively. The use of human material was approved by the Ethikkommision of the Charité–Universitätsmedizin Berlin and performed in accordance with the Declaration of Helsinki. Primers used are listed in Supplementary Table 3.
Statistical analyses of experimental and microarray data
All statistical analyses were done in R v2.9.1 (http://www.r-project.org/) and GraphPad Prism Version 5.0. One-sided Welch’s t-test was used to analyze real-time quantitative PCR experiments in Figures 4d and 5a; two-sided Welch’s t-test was used to analyze data from proliferation experiments, luciferase assays as well as real-time quantitative PCR experiments. Gene expression profiles were determined with Illumina HT-12 v 4.0 Expression BeadChips (Illumina, San Diego, CA, USA). Hybridization was performed in duplicates from independent transfection experiments. Raw microarray data were background corrected with Illumina’s proprietary background correction routine and quantile normalization were applied. Subsequently, data were log2 transformed. LIMMA was used for determination of significantly differentially expressed genes. Listed are all genes with a log2 fold change of at least −/+ 0.5 at 72 h (Supplementary Table 6). Microarray data are available from the Gene Expression Omnibus of the National Center for Biotechnology Information (GSE86845).
Results
Loss of PDLIM2 expression in HRS cell lines
Given the plethora of activated TFs in cHL and a hypothesized involvement of the ubiquitin system in this TF activation, we started our analysis with a literature search on genes associated with the ubiquitin system, combined with the analysis of microarray-based gene expression data of HRS and NH cell lines.17 One of the most interesting candidates was PDLIM2 that is involved in the control of TF degradation,19, 22 and that showed a loss of expression in several of the HRS compared with NH cell lines (data not shown). To confirm these data, we analyzed PDLIM2 mRNA and protein expression levels in an extended panel of cell lines (Figure 1), including various HRS and NH cell lines reflecting different B-cell differentiation stages. In contrast to the robust mRNA expression in the analyzed NH cell lines, PDLIM2 mRNA expression was substantially reduced or lost in seven out of nine HRS cell lines (L1236, L591, HDLM-2, L540, L540Cy, U-HO1 and SUP-HD1; Figures 1a and b). Interestingly, maintenance of PDLIM2 expression in L428 and KM-H2 cells correlated with the presence of deleterious IκB mutations.10, 37
Loss of PDLIM2 expression in HRS cell lines. (a) PDLIM2 mRNA expression analyses by quantitative (upper panel) and semiquantitative (lower panel) PCR analyses in various HRS and NH cell lines, as indicated. (a, upper panel) PDLIM2 mRNA expression was assessed by real-time quantitative PCR (qPCR) and calculated using the 2−ΔΔCt method. Expression in Reh cells was set as 1. Error bars denote 95% confidence intervals. One out of three independent experiments is shown. ***P<0.001. (a, lower panel) Analysis of PDLIM2 and, as a control, GAPDH mRNA expression in the various cell lines by semiquantitative PCR (sqPCR). One of three independent experiments is shown. (b) Analysis of PDLIM2 mRNA (upper panel) and protein (lower panel) expression in HRS (L428, HDLM-2, U-HO1, SUP-HD1) and NH (BJAB, SU-DHL-4) cell lines, as indicated. Expression of GAPDH and PARP1 are shown as controls. One of three independent experiments is shown. WB, western blot analysis. (c) Analysis of PDLIM2 and IκBα protein expression in various cell lines, as indicated. Nuclear extracts were analyzed for PDLIM2 (upper panel), and whole-cell extracts for IκBα (lower panel) protein expression; expression of PARP1 and β-actin are shown as controls. One of four independent experiments is shown. Note that L428 and KM-H2 cells lack IκBα protein expression because of deleterious NFKBIA mutations or deletions.10
At the protein level, we confirmed the loss of PDLIM2 expression in most of the HRS cell lines (Figures 1b and c). At least three isoforms of PDLIM2 are known with two major isoforms expressed in the lymphoid compartment.20 These two isoforms were cloned as controls for PDLIM2 protein expression in immunoblotting experiments (Supplementary Figure 1). In nearly all NH B cell lines, robust expression of PDLIM2 was detectable (Figures 1b and c), whereas among HRS cell lines PDLIM2 protein expression was only maintained in L428 and KM-H2 cells, for both of which the correlation with a loss of IκBα protein expression was confirmed (Figure 1c). If expressed, the cell lines nearly exclusively expressed PDLIM2 tv 2 (Supplementary Figure 1). Interestingly, despite PDLIM2 mRNA expression, L1236 cells virtually lacked PDLIM2 protein expression.
Lack of PDLIM2 expression in HRS cells of primary cHL samples
Next, we determined PDLIM2 protein expression in primary cHL tissue samples as well as normal lymphoid tissue (Figure 2 and Supplementary Table 4a). We established immunohistochemical analyses with two different antibodies that showed consistent results. In normal lymphoid tissue, PDLIM2 was expressed in virtually all lymphoid cells with however reduced expression levels in germinal center B cells. Furthermore, various B-cell non-Hodgkin lymphoma cases including 10 follicular lymphoma, 15 diffuse large B-cell lymphoma and 10 Burkitt lymphoma cases showed PDLIM2 expression in all cases, in the vast majority of them with a strong staining pattern (Supplementary Figure 2). In contrast, among 20 cHL cases of various subtypes with or without Epstein–Barr virus infection, HRS cells of 19 cases lacked PDLIM2 expression (Figure 2 and Supplementary Table 4a). In all cases, PDLIM2 expression was readily detectable in surrounding cells that served as internal control. Only HRS cells of one case expressed PDLIM2, and this was however predominantly observed at the cell membrane (Figure 2g). To investigate a possible correlation with loss of IκBα expression suggested by the cell line analyses, we established immunohistochemical analyses of IκBα using an antibody recognizing the C-terminus of the protein. cHL cases lacking IκBα expression due to deleterious truncating mutations would therefore be expected to be negative.10 In 4 of 20 cHL cases (20%), IκBα protein expression was not detectable, and this is in line with the estimated percentage of deleterious NFKBIA mutations in cHL in other studies;11 all 4 cases lacked PDLIM2 expression. We conclude that loss of PDLIM2 expression is a unifying defect of HRS cells independent of IκB alterations.
HRS cells of primary classical HL cases rarely express PDLIM2 and show a recurrent loss of IκBα protein expression. (a, b) Naive mantle zone B cells present an intense PDLIM2 protein expression, whereas PDLIM2 expression is weak in germinal center B cells (a, × 100; b, × 200). (c, d) Representative cHL mixed cellularity case with lost PDLIM2 expression (c, × 200, insert, × 400) and lost IκBα expression (d, × 200, insert, × 400). (e, f) Representative cHL nodular sclerosis case showing lost PDLIM2 expression (e, × 200, insert, × 400). This case shows weak IκBα protein expression in the HRS cells (f, × 200, insert, × 400). (g, h) In this exceptional cHL mixed cellularity case a membrane-bound PDLIM2 expression pattern occurred (g, × 200, insert, × 400). The IκBα protein expression was typically weak in the HRS cells of this case (h, × 200, insert, × 400). For all PDLIM2 stainings shown in this figure, a rabbit anti-PDLIM2 antibody was used.
Genomic loss of the PDLIM2 gene locus, aberrant PDLIM2 promoter methylation and altered splicing are associated with loss of PDLIM2 expression in HRS cells
To reveal the underlying molecular mechanisms of loss of PDLIM2 expression in cHL cells, we screened for genomic alterations, DNA methylation changes and splicing defects. The analysis of previously published SNP-CHIP data9, 38 and subsequent fluorescence in situ hybridization analyses (Supplementary Table 2 and Supplementary Figure 3) indicated deletions of the PDLIM2 locus in 3 out of 4 cHL cell lines, leading us to analyze primary HRS cells for chromosomal losses targeting the PDLIM2 gene. For the primary cases we used FICTION analysis that combined anti-CD30 immunophenotyping with the fluorescence in situ hybridization probe for 8p21.3—where PDLIM2 is located—and internal centromere controls. In 5 out of 20 (25%) cases, we observed a deletion of the PDLIM2 gene in at least 30% of the analyzed HRS cell nuclei (Supplementary Table 2). Remarkably, in one case with a deletion targeting the PDLIM2 locus, in some HRS cells the signal pattern was characteristic for a homozygous deletion. In this case, an average of 15% of the HRS cell nuclei lacked the fluorescent probe signal corresponding to the 8p21.3 region harboring PDLIM2 but showed at least one fluorescent signal for the CEP probes. Together, these results are well in line with previous array comparative genomic hybridization studies on primary HRS cells13, 39, 40 (for a detailed summary see also Supplementary Table 5) and suggest that genomic deletions contribute to the loss of PDLIM2 expression in HRS cells.
An important alternative mechanism of gene silencing in tumor cells is DNA methylation. We therefore investigated the methylation status of CpG elements of three different regions in the vicinity of two major PDLIM2 transcription start sites, including one previously published region24, 25 (Figure 3a and Supplementary Figure 4). In HRS cell lines without PDLIM2 expression, two DNA regions (designated as promoter 1 (P1): NM_021630 and promoter 2 (P2): NM_176871) were strongly methylated at the investigated CpG elements (Figure 3a). The same regions were unmethylated in DNA from cells with PDLIM2 expression including NH cell lines, peripheral blood cells from healthy donors or NH mature B cell-derived lymphomas, as shown for follicular lymphoma (Figures 3a and 6e and Supplementary Figure 4a). In the previously published region (located further upstream in the 5′ region of PDLIM2; designated as pub. reg.) that has been associated with PDLIM2 gene methylation in cancer cells,24 we did not observe alterations of PDLIM2 DNA methylation in our samples irrespective of the PDLIM2 expression status (Supplementary Figure 4b). We conclude that aberrant DNA methylation of the P1 and P2 regions is associated with the loss of PDLIM2 expression in HRS cells.
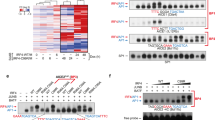
DNA methylation of the PDLIM2 promoters in HRS cell lines; altered splicing of PDLIM2 mRNA in L1236 cells. (a, upper panel) Overview of the regions analyzed (amplicons) for DNA methylation. P1, amplicon of this region corresponds to the region of the transcription start site (TSS) of the PDLIM2 transcript variant 2 (NM_021630); P2, amplicon of this region corresponds to the region of the TSS of the PDLIM2 transcript variant 1 (NM_176871); pub. reg., a region previously analyzed for methylation.24, 25 The percentage GC content of the DNA is shown in five-base windows. The exon–intron structures of the PDLIM2 transcript variants are diagrammed underneath. (a, lower panel) Result of bisulfite pyrosequencing in various cell lines, as indicated. The percentage of DNA methylation for each individual CpG element in each amplicon is represented by a color code (red, CpG site fully methylated (100%); green, CpG site fully unmethylated (0%)). As controls, commercially available completely methylated DNA (mean methyl. control) and unmethylated pooled DNA extracted from peripheral blood (mean pool pB) were included. (b) PDLIM2 altered splicing in L1236 cells. (b, left panel) mRNA sequence of a PDLIM2 transcript amplified in L1236 cells. Shown is the PDLIM2 mRNA sequence with skipping of exon 7 (end of exon 6 is underlined in black, beginning of exon 8 is underlined in red; according to NM_021630.4) that results in a reading frame shift and generation of a premature STOP (TGA, marked in red). The wild-type (wt) STOP codon is marked in bold black. (b, right panel) Structure of wt PDLIM2 and the predicted truncated L1236 variant that results in a C-terminally truncated protein with deletion of the LIM domain (black box, protein sequence out of frame). AA numbers of the proteins are indicated. (c) PDLIM2 protein variants. HEK293 cells were transfected with empty plasmid (Mock) as a control, or plasmids encoding PDLIM2 tv2, the L1236 splicing variant (varL1236) or a C-terminally truncated version lacking the LIM domain.22 Western blot (WB) was performed using antibody ab68220. Expression of PARP1 is shown as control. (d) The PDLIM2 L1236 splicing variant is instable. HEK293 cells were transfected with control plasmid (−) or plasmids encoding PDLIM2 tv2 or the L1236 splicing variant (PDLIM2 varL1236). At 48 h after transfection, cells were treated with cycloheximide (CHX). At the indicated times, whole-cell lysates were prepared and analyzed for expression of the respective PDLIM2 proteins. Expression of PARP1 was analyzed as a control. One of three independent experiments is shown. Note that the expression level of PDLIM2 varL1236 rapidly decreases following CHX treatment, whereas the expression level of PDLIM2 tv 2 remains stable even after 6 h of CHX treatment.
Furthermore, in L1236 HRS cells we observed a discrepancy between the—although low—PDLIM2 mRNA but absent protein expression (see Figure 1). We therefore sequenced the L1236 PDLIM2 mRNA to search for sequence alterations (Figure 3b). Apart from expression of PDLIM2 wt transcripts, we revealed in a fraction of transcripts skipping of exon 7 (according to NM_021630.4; skipping of 77 bp), resulting in a frameshift and generation of a premature termination codon in exon 8 with a predicted protein size of ∼10 kDa smaller than the wt variant and lacking the C-terminally located LIM domain (Figure 3b), suggesting a loss of function alteration.22 We cloned this variant (PDLIM2 varL1236) and confirmed a protein size of ∼28 kDa by western blot analysis (Figure 3c). Remarkably, by treatment of the cells with cycloheximide we revealed a strongly reduced half-life of this truncated PDLIM2 variant (Figure 3d), indicating reduced protein stability. Sequencing at the DNA level did not show a genomic alteration in L1236 cells (data not shown). This splicing variant was not detected in NH cell lines or Epstein–Barr virus-positive lymphoblastoid cells (data not shown). Together, our studies indicate that various mechanisms are associated with the loss of PDLIM2 in HRS cells, suggesting a selection pressure for PDLIM2 loss in cHL.
Reconstitution of PDLIM2 in HRS cell lines results in reduction of cell growth, inhibition of NF-κB activity and alteration of differentially expressed genes
To investigate the biological consequences of the PDLIM2 loss in HRS cells, we reconstituted PDLIM2 expression in the cell lines L591, L540Cy and SUP-HD1 (Figure 4a and Supplementary Figure 5a). In these assays, we observed a significant reduction of proliferation following PDLIM2 reconstitution, in L591 cells by ∼50% and in L540Cy by ∼25% (Figure 4b). We did not observe an alteration of cell viability (data not shown). Given the known role of PDLIM2 for termination of NF-κB activity22 and of NF-κB for growth and survival of HRS cells,4, 41 we analyzed a possible inhibition of the NF-κB system following PDLIM2 reconstitution in HRS cells using NF-κB-dependent reporter gene studies (Figure 4c and Supplementary Figure 5b). We included the NF-κB super-repressor IκBαΔN14 as a positive control. As expected, in these cell lines the constitutive NF-κB activity was strongly reduced by IκBαΔN. In accordance with an NF-κB inhibitory function,22 PDLIM2 wt also suppressed the NF-κB activity in HRS cell lines. In line with a putative loss of function, the truncated PDLIM2 varL1236 was clearly less effective regarding NF-κB inhibition, and a previously characterized loss-of-function ΔLIM variant lacking the LIM domain22 did not alter NF-κB activity (Supplementary Figure 5b). Together, the loss of PDLIM2 contributes to the enhanced NF-κB activity characteristic for HRS cells.
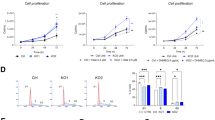
PDLIM2 reconstitution in HRS cell lines reduces proliferation and suppresses NF-κB transcriptional activity; analysis of PDLIM2 regulated, differentially expressed genes. (a) The HRS cell lines L591 and L540Cy were transiently transfected with empty plasmid as a control (Mock) or PDLIM2 (tv 2) expression plasmid (PDLIM2) along with a pEGFP expression plasmid. At 48 h after transfection, transfected GFP+ cells were enriched and protein expression of PDLIM2 and, as a control, β-actin, was analyzed by western blot (WB). Reh cells with and Mac-2A cells without PDLIM2 expression (see also Figure 6) as well as empty plasmid (Mock) or PDLIM2-transfected HEK293 cells were included as controls. (b) PDLIM2 reconstitution reduces proliferation of HRS cell lines. Cells were transfected and enriched as described in (a). At 24, 48 and 72 h after enrichment, cells were pulsed with 1 μCi [3H]-thymidine per well for a further 20 h, and [3H]-thymidine incorporation was determined. c.p.m., counts per min. (c) PDLIM2 reconstitution suppresses NF-κB transcriptional activity in HRS cell lines. (c, upper panel) pGL3-basic or 6xκBLuc-constructs were transfected into HRS cell lines together with empty plasmid (Mock; filled bars) or PDLIM2 (gray bars) or IκBαΔN (open bars) expression constructs. Luciferase activity is shown as fold activation compared with pGL3-basic (set 1). Data are represented as mean±s.d. (c, lower panel) Expression of PDLIM2, IκBαΔN and, as a control, β-actin in transfected cells was analyzed by WB. Note that IκBαΔN is significantly smaller than full-length IκBα. One out of five independent experiments is shown. *P<0.05; **P<0.01; ***P<0.001. (d) Analysis of PDLIM2-regulated genes in L591 HRS cells. L591 cells were transfected and enriched as described in (a). (d, upper panel) FBXO32, IL10RA, MAK and IER5 mRNA expression was assessed by real-time PCR and calculated using the 2−ΔΔCt method. Expression in Mock-transfected cells was set as 1. Error bars denote 95% confidence intervals. One out of three independent experiments in shown. **P<0.01; ***P<0.001. (d, lower panel) Analysis of FBXO32, IL10RA, MAK, IER5 and, as a control, GAPDH mRNA expression in the various cell lines by reverse transcriptase-PCR (RT-PCR). One of three independent experiments is shown.
To gain insights into PDLIM2-induced transcriptional changes in HRS cells, we performed genome-wide gene expression profiling of L591 cells following PDLIM2 reconstitution by Illumina microarrays. In total, 80 genes showed significant expression changes following PDLIM2 reconstitution (Supplementary Table 6). Genes downregulated by PDLIM2 expression included FBXO32, MAK, IL10RA and IER5 (Figure 4d, upper panel), and their differential gene expression pattern between the HL and NH cell lines (Figure 4d, lower panel) was verified by PCR analyses. These data indicate that the loss of PDLIM2 in HRS cells contributes to the deregulation of HRS cell-specific genes.
Superactivation of NF-κB and AP-1 transcriptional activity following downregulation of PDLIM2 in NH cells
Next, we aimed to investigate the consequences of PDLIM2 downregulation on the transcriptional activity of the NF-κB and AP-1 TFs in transformed B lymphoid cells (Figure 5). We transfected Reh and SU-DHL-4 cells with two different PDLIM2-specific siRNAs (Figure 5a and data not shown). Having established the PDLIM2 knockdown, we performed NF-κB reporter assays with these cells. As we hypothesized that the effect of PDLIM2 downregulation might be most pronounced following concomitant stimulation of the respective signaling pathways, we included stimulation experiments with the phorbol ester PMA in our reporter gene analyses (Figures 5b and c). As expected, PMA stimulation alone led to a moderate but significant increase of NF-κB reporter activity. Importantly, PMA-induced NF-κB stimulation in Reh cells was further increased approximately fivefold following PDLIM2 knockdown (Figure 5b). Similar results were obtained with SU-DHL-4 cells (Figure 5c). Furthermore, we investigated the effect of PDLIM2-modulation on AP-1 reporter activity in Reh cells (Figure 5d). As observed for NF-κB, the PMA-induced AP-1 activity was significantly increased following PDLIM2 knockdown. These data indicated that in this experimental setting PDLIM2 controls inducible TF activities and that loss of PDLIM2 enhances the activity of NF-κB and AP-1 TFs.
Knockdown of PDLIM2 in NH cells results in NF-κB and AP-1 superactivation. (a) Knockdown of PDLIM2 in Reh cells. Reh cells were transfected with control (siCTL) or a combination of two PDLIM2-specific siRNA (siPDLIM2 #1+#2/siPDLIM2) constructs, and enriched transfected cells were analyzed for PDLIM2 mRNA (left) and protein (right) expression. (a, left) PDLIM2 mRNA expression was assessed by real-time PCR and calculated using the 2−ΔΔCt method. Expression in control-transfected Reh cells was set as 1. Error bars denote 95% confidence intervals. ***P<0.001. (a, right) Analysis of PDLIM2 protein expression in siCTL- and siPDLIM2-transfected Reh cells. Whole-cell extracts were analyzed for protein expression of PDLIM2 and, as a control, β-actin. Untransfected Reh (positive control) and L591 (negative control) cells as well as Mock- or PDLIM2-transfected HEK293 cells are shown as controls. (b) Reh cells were transiently transfected with pGL3-basic or 6xκBLuc-reporter gene constructs together with control (siCTL) or a combination of two PDLIM2-specific siRNA (siPDLIM2 #1+#2 / siPDLIM2) constructs. At 48 h after transfection, cells were left untreated or treated twice with PMA. Thereafter, cells were analyzed for luciferase activity. Luciferase activity is shown as fold activation compared with pGL3-basic- and siCTL-transfected and unstimulated cells (set 1). Data are represented as mean±s.d. One of four independent experiments is shown. ***P<0.001. (c) SU-DHL-4 cells were transiently transfected and analyzed as described in (c). One of three independent experiments is shown. *P<0.05; **P<0.01; ***P<0.001. (d) Reh cells were transiently transfected with pGL3-basic or 5xTRE-Luc-(AP-1) reporter gene constructs together with control (siCTL) or a combination of two PDLIM2-specific siRNA (siPDLIM2 #1+#2/siPDLIM2) constructs. At 48 h after transfection, cells were left untreated or treated twice with PMA. Thereafter, cells were analyzed for luciferase activity. Luciferase activity is shown as fold activation compared with pGL3-basic- and siCTL-transfected and unstimulated cells (set 1). Data are represented as mean±s.d. One of four independent experiments is shown. ***P<0.001.
Downregulation of PDLIM2 in ALCL
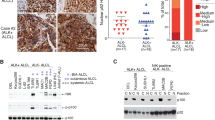
ALCL shares several key biological aspects with cHL.42, 43, 44, 45 We therefore analyzed PDLIM2 expression in ALCL (Figure 6). We revealed a striking downregulation or loss of PDLIM2 expression in ALCL cell lines at the mRNA and protein levels (Figures 6a and b). Similarly, 10 out of 10 ALK-positive and 11 out of 11 ALK-negative primary ALCL cases lacked PDLIM2 protein expression as assessed by immunohistochemistry (Figure 6c and Supplementary Table 4b). We did not observe chromosomal aberrations of the PDLIM2 gene locus in ALCL (data not shown). However, the PDLIM2 locus displayed strong methylation at the P1 and P2 sites in ALCL cell lines lacking PDLIM2 expression (Figure 6d). Furthermore, we observed hypermethylation mainly of the P2 site in the majority of primary ALCL cases analyzed (Figure 6e).
Loss of PDLIM2 expression in ALCL. (a) PDLIM2 mRNA expression analyses by quantitative (upper panel) and semi-quantitative (lower panel) PCR analyses in ALK-positive (K299, SU-DHL-1, DEL, JB6) and ALK-negative (FE-PD, Mac-1, Mac-2A, DL40) ALCL as well as T-cell leukemia-derived (Jurkat, KE-37, Molt-14, H9) control cell lines. (a, upper panel) PDLIM2 mRNA expression was assessed by real-time quantitative PCR and calculated using the 2−ΔΔCt method. Expression in Jurkat cells was set as 1. Error bars denote 95% confidence intervals. One out of three independent experiments in shown. ***P<0.001. (a, lower panel) Analysis of PDLIM2 and, as a control, GAPDH mRNA expression by sqPCR. One of three independent experiments is shown. (b) Analysis of PDLIM2 and IκBα protein expression in the various cell lines. Whole-cell extracts were analyzed for protein expression of PDLIM2 (upper panel), IκBα (center) and, as a control, β-actin (lower panel). One of four independent experiments is shown. (c) Lack of PDLIM2 expression in primary ALCL. Representative immunohistochemical (IHC) stainings of one ALK-positive (left) and one ALK-negative (right) ALCL case. Note that the large ALCL cells lack PDLIM2 protein expression, whereas infiltrating hematopoietic cells express PDLIM2. (d) Methylation of the PDLIM2 promoter regions in ALCL cell lines. Result of bisulfite pyrosequencing in various cell lines, as indicated; for the analyzed regions refer to Figure 3a. The percentage of DNA methylation for each individual CpG element in each amplicon is represented by a color code (red, CpG site fully methylated (100%), green, CpG site fully unmethylated (0%)). As controls, commercially available completely methylated DNA (mean methyl. control) and unmethylated pooled DNA extracted from peripheral blood (mean pool pb) were included. (e) Methylation of the PDLIM2 promoter regions in primary ALK-positive ALCL analyzed by bisulfite pyrosequencing and follicular lymphoma (FL) samples analyzed by whole methylome sequencing.52 Results are depicted as described in (d). NA, data not available; TCC, tumor cell content.
Discussion
Here we describe and functionally characterize the loss of the putative ubiquitin-E3 ligase PDLIM2 as unifying defect in distinct lymphoma entities, cHL and ALCL. PDLIM2 has been linked to tumorigenesis, in particular in colon and breast cancer or HTLV1-induced adult T-cell leukemia.24, 25, 46 In these malignancies, PDLIM2 expression levels are consistently reduced, in part because of aberrant DNA methylation.24, 25 Importantly, reconstitution of PDLIM2 reduced the tumor growth, suggesting a tumor suppressor function of PDLIM2.24, 25, 46 However, for most of these malignancies data on PDLIM2 in primary samples are lacking, and cancer cell lines used for these studies display significant PDLIM2 baseline expression levels.24, 25 Our data for cHL and ALCL show the most prominent loss of PDLIM2 expression in malignancies reported so far, and they support a tumor suppressor role of PDLIM2: (1) in most HRS and ALCL cells PDLIM2 expression is undetectable, reflecting a functional PDLIM2 deletion; (2) several mechanisms contribute to PDLIM2 silencing, including aberrant methylation, genomic deletion and altered splicing, all arguing for selective pressure on this gene locus; (3) the biological effects observed in HRS cell lines argue for the functional relevance of lost PDLIM2 expression.
In more detail, in line with the work of Tanaka et al.,22 we demonstrate that PDLIM2 is ubiquitously expressed in lymphoid cells, with however clearly reduced expression levels in germinal center B cells, as demonstrated in our work. In agreement with the expression in lymphoid cells, the analyzed PDLIM2 promoter regions were unmethylated in peripheral mononuclear cells from healthy donors. In contrast, PDLIM2 expression was lost and both analyzed PDLIM2 promoter regions were strongly methylated in HRS cell lines. Thereby, the already low PDLIM2 expression in germinal center B cells, the putative cells of origin of cHL,1 might facilitate PDLIM2 silencing in HRS cells. Of note, we did not observe an altered methylation in a region previously proposed to be involved in PDLIM2 silencing in cancer cells,24, 25 and this might be because of different epigenetic regulations of PDLIM2 between cancer and lymphoid cells, or methodical differences. Importantly, the PDLIM2 gene locus is located in a minimal deleted region of 600 kb in 8p21.3 in B-cell non-Hodgkin lymphoma, and TRAIL-R1 and -R2 (tumor necrosis factor-related apoptosis-inducing ligand receptors 1 and 2) have been suggested as candidate genes of this deletion.26, 27 Our data suggest that PDLIM2 might be a further candidate gene targeted by this deletion. Finally, we detected in one HRS cell line a non-DNA-templated altered splicing pattern leading to an instable, C-terminally truncated protein lacking the LIM domain. Even though the relevance of this finding for primary cHL remains unclear, similarly truncated mutant constructs—generated for biological studies—lose their putative tumor suppressor function and the ability to ubiquitinate NF-κB-p65.20, 22, 24 Together, our data add the loss of PDLIM2 to the list of defects that target inducible signaling pathways in cHL.1
We propose that the major implication of the loss of PDLIM2 expression in HRS cells is the generation of a hyperresponsive cellular background by promoting the sustained activation of TF activities due to disturbed degradation. This fits with the observation that HRS cells share hallmark gene expression patterns with highly activated lymphoid cells and that they are characterized by the abundant and constitutive activity of TFs such as NF-κB, AP-1 and STAT that are usually transiently activated and rapidly downregulated following their induction in the physiological context.1, 5, 7 Characteristic for HRS cells is thereby, apart from the strength of the activation, the plethora of concomitantly activated TF activities. Because of the involvement of PDLIM2 in the termination of TFs including NF-κB,22 STAT factors19, 21 and, as suggested by our work, AP-1, the loss of PDLIM2 expression in HRS cells provides a molecular mechanism that contributes to this phenomenon. Our data in Reh cells as well as previously published data19, 22 indicate that knockdown of PDLIM2 per se is not sufficient for induction of TF activities, that is, the respective signaling pathways need to be stimulated for example by ligand–receptor interactions or constitutive activation of upstream signaling components. Up to now, several genomic and molecular defects of such upstream signaling pathway components involved in NF-κB, Janus kinase (JAK)/STAT and AP-1 activation are known in HRS cells.1 Whereas most of these defects target receptor-associated regulatory proteins and rather proximal located kinase modules, the loss of PDLIM2 in HRS cells adds a defect at the far end of the respective TF activation cascades by disabling their nuclear brakes.
Functionally, following PDLIM2 reconstitution in HRS cells the biological consequences were weaker than previously demonstrated for, for example, stringent NF-κB inhibition.41 This is most likely explained by the plethora of upstream defects in the respective signaling cascades in HRS cells1 and the fact that PDLIM2 is certainly not the only factor mediating post-translational modifications of these TFs.47 Furthermore, experimentally, interfering with TF activities at the far end of their activation cascade is much more sluggish in comparison with direct intervention with super-repressors like IκBαΔN. However, HRS cells will provide an ideal tool to further investigate the function and mechanisms of action of PDLIM2 in future studies. This is already demonstrated by the identification of PDLIM2 target genes in our study, the function of which in cHL has to be investigated in future studies. In addition, future work will be required to elucidate the exact mechanism of how PDLIM2 interferes with TF activation in HRS cells, as ubiquitin-dependent and -independent mechanisms have been described.19
Remarkably, we identified ALCL as a second lymphoid malignancy with a consistent loss of PDLIM2 expression. Importantly, ALCL shares biological similarities with cHL, including the loss of cellular identity,1, 43 a highly activated phenotype (as exemplified by high-level CD30 expression)42, 44 and activation of AP-1, STAT and NF-κB TFs.5, 44, 48 Suppression of PDLIM2 occurs independent of the presence or absence of ALK translocations. Even though ALK activation contributes to the induction of several of the TF activities mentioned above,49 the loss of PDLIM2 expression most likely enhances TF activation in ALCL in a similar manner as described in our work in cHL, and this has to be investigated in future studies.
Taken together, we have identified the loss of PDLIM2 as a unifying defect of cHL and ALCL that contributes to the permanent activation of a plethora of key TFs, thereby promoting cellular growth and differential gene regulation. These data increase the complexity of molecular and genomic defects already known in these lymphoid malignancies. Finally, such defects of the ubiquitin system might offer an explanation for the rather disappointing clinical activity of proteasome inhibitors in cHL.50, 51
References
Küppers R, Engert A, Hansmann ML . Hodgkin lymphoma. J Clin Invest 2012; 122: 3439–3447.
Steidl C, Connors JM, Gascoyne RD . Molecular pathogenesis of Hodgkin's lymphoma: increasing evidence of the importance of the microenvironment. J Clin Oncol 2011; 29: 1812–1826.
Ansell SM, Lesokhin AM, Borrello I, Halwani A, Scott EC, Gutierrez M et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin's lymphoma. N Engl J Med 2015; 372: 311–319.
Bargou RC, Emmerich F, Krappmann D, Bommert K, Mapara MY, Arnold W et al. Constitutive nuclear factor-kappaB-RelA activation is required for proliferation and survival of Hodgkin's disease tumor cells. J Clin Invest 1997; 100: 2961–2969.
Mathas S, Hinz M, Anagnostopoulos I, Krappmann D, Lietz A, Jundt F et al. Aberrantly expressed c-Jun and JunB are a hallmark of Hodgkin lymphoma cells, stimulate proliferation and synergize with NF-kappa B. EMBO J 2002; 21: 4104–4113.
Scheeren FA, Diehl SA, Smit LA, Beaumont T, Naspetti M, Bende RJ et al. IL-21 is expressed in Hodgkin lymphoma and activates STAT5; evidence that activated STAT5 is required for Hodgkin lymphomagenesis. Blood 2008; 111: 4706–4715.
Kreher S, Bouhlel MA, Cauchy P, Lamprecht B, Li S, Grau M et al. Mapping of transcription factor motifs in active chromatin identifies IRF5 as key regulator in classical Hodgkin lymphoma. Proc Natl Acad Sci USA 2014; 111: E4513–E4522.
Kube D, Holtick U, Vockerodt M, Ahmadi T, Haier B, Behrmann I et al. STAT3 is constitutively activated in Hodgkin cell lines. Blood 2001; 98: 762–770.
Schmitz R, Hansmann ML, Bohle V, Martin-Subero JI, Hartmann S, Mechtersheimer G et al. TNFAIP3 (A20) is a tumor suppressor gene in Hodgkin lymphoma and primary mediastinal B cell lymphoma. J Exp Med 2009; 206: 981–989.
Emmerich F, Meiser M, Hummel M, Demel G, Foss HD, Jundt F et al. Overexpression of I kappa B alpha without inhibition of NF-kappaB activity and mutations in the I kappa B alpha gene in Reed-Sternberg cells. Blood 1999; 94: 3129–3134.
Lake A, Shield LA, Cordano P, Chui DT, Osborne J, Crae S et al. Mutations of NFKBIA, encoding IkappaB alpha, are a recurrent finding in classical Hodgkin lymphoma but are not a unifying feature of non-EBV-associated cases. Int J Cancer 2009; 125: 1334–1342.
Joos S, Granzow M, Holtgreve-Grez H, Siebert R, Harder L, Martin-Subero JI et al. Hodgkin's lymphoma cell lines are characterized by frequent aberrations on chromosomes 2p and 9p including REL and JAK2. Int J Cancer 2003; 103: 489–495.
Hartmann S, Martin-Subero JI, Gesk S, Hüsken J, Giefing M, Nagel I et al. Detection of genomic imbalances in microdissected Hodgkin- and Reed-Sternberg cells of classical Hodgkin lymphoma by array based comparative genomic hybridization. Haematologica 2008; 93: 1318–1326.
Krappmann D, Emmerich F, Kordes U, Scharschmidt E, Dörken B, Scheidereit C . Molecular mechanisms of constitutive NF-kappaB/Rel activation in Hodgkin/Reed-Sternberg cells. Oncogene 1999; 18: 943–953.
Lamprecht B, Walter K, Kreher S, Kumar R, Hummel M, Lenze D et al. Derepression of an endogenous long terminal repeat activates the CSF1R proto-oncogene in human lymphoma. Nat Med 2010; 16: 571–579, 571p following 579.
Babaian A, Romanish MT, Gagnier L, Kuo LY, Karimi MM, Steidl C et al. Onco-exaptation of an endogenous retroviral LTR drives IRF5 expression in Hodgkin lymphoma. Oncogene 2015; 35: 2542–2546.
Köchert K, Ullrich K, Kreher S, Aster JC, Kitagawa M, Jöhrens K et al. High-level expression of Mastermind-like 2 contributes to aberrant activation of the NOTCH signaling pathway in human lymphomas. Oncogene 2011; 30: 1831–1840.
Torrado M, Senatorov VV, Trivedi R, Fariss RN, Tomarev SI . Pdlim2, a novel PDZ-LIM domain protein, interacts with alpha-actinins and filamin A. Invest Ophthalmol Vis Sci 2004; 45: 3955–3963.
Tanaka T, Soriano MA, Grusby MJ . SLIM is a nuclear ubiquitin E3 ligase that negatively regulates STAT signaling. Immunity 2005; 22: 729–736.
Loughran G, Healy NC, Kiely PA, Huigsloot M, Kedersha NL, O'Connor R . Mystique is a new insulin-like growth factor-I-regulated PDZ-LIM domain protein that promotes cell attachment and migration and suppresses anchorage-independent growth. Mol Biol Cell 2005; 16: 1811–1822.
Tanaka T, Yamamoto Y, Muromoto R, Ikeda O, Sekine Y, Grusby MJ et al. PDLIM2 inhibits T helper 17 cell development and granulomatous inflammation through degradation of STAT3. Sci Signal 2011; 4: ra85.
Tanaka T, Grusby MJ, Kaisho T . PDLIM2-mediated termination of transcription factor NF-kappaB activation by intranuclear sequestration and degradation of the p65 subunit. Nat Immunol 2007; 8: 584–591.
Sun F, Xiao Y, Qu Z . Oncovirus Kaposi sarcoma herpesvirus (KSHV) represses tumor suppressor PDLIM2 to persistently activate nuclear factor kappaB (NF-kappaB) and STAT3 transcription factors for tumorigenesis and tumor maintenance. J Biol Chem 2015; 290: 7362–7368.
Qu Z, Fu J, Yan P, Hu J, Cheng SY, Xiao G . Epigenetic repression of PDZ-LIM domain-containing protein 2: implications for the biology and treatment of breast cancer. J Biol Chem 2010; 285: 11786–11792.
Qu Z, Yan P, Fu J, Jiang J, Grusby MJ, Smithgall TE et al. DNA methylation-dependent repression of PDZ-LIM domain-containing protein 2 in colon cancer and its role as a potential therapeutic target. Cancer Res 2010; 70: 1766–1772.
Martinez-Climent JA, Vizcarra E, Sanchez D, Blesa D, Marugan I, Benet I et al. Loss of a novel tumor suppressor gene locus at chromosome 8p is associated with leukemic mantle cell lymphoma. Blood 2001; 98: 3479–3482.
Rubio-Moscardo F, Blesa D, Mestre C, Siebert R, Balasas T, Benito A et al. Characterization of 8p21.3 chromosomal deletions in B-cell lymphoma: TRAIL-R1 and TRAIL-R2 as candidate dosage-dependent tumor suppressor genes. Blood 2005; 106: 3214–3222.
Lamprecht B, Bonifer C, Mathas S . Repeat-element driven activation of proto-oncogenes in human malignancies. Cell Cycle 2010; 9: 4276–4281.
Brummelkamp TR, Bernards R, Agami R . A system for stable expression of short interfering RNAs in mammalian cells. Science 2002; 296: 550–553.
Schwarzer N, Nost R, Seybold J, Parida SK, Fuhrmann O, Krull M et al. Two distinct phospholipases C of Listeria monocytogenes induce ceramide generation, nuclear factor-kappa B activation, and E-selectin expression in human endothelial cells. J Immunol 1998; 161: 3010–3018.
Peterziel H, Müller J, Danner A, Barbus S, Liu HK, Radlwimmer B et al. Expression of podoplanin in human astrocytic brain tumors is controlled by the PI3K-AKT-AP-1 signaling pathway and promoter methylation. Neuro Oncol 2012; 14: 426–439.
Janz M, Hummel M, Truss M, Wollert-Wulf B, Mathas S, Jöhrens K et al. Classical Hodgkin lymphoma is characterized by high constitutive expression of activating transcription factor 3 (ATF3), which promotes viability of Hodgkin/Reed-Sternberg cells. Blood 2006; 107: 2536–2539.
Hartmann S, Agostinelli C, Klapper W, Korkolopoulou P, Koch K, Marafioti T et al. Revising the historical collection of epithelioid cell-rich lymphomas of the Kiel Lymph Node Registry: what is Lennert's lymphoma nowadays? Histopathology 2011; 59: 1173–1182.
Schwarting R, Gerdes J, Dürkop H, Falini B, Pileri S, Stein H . BER-H2: a new anti-Ki-1 (CD30) monoclonal antibody directed at a formol-resistant epitope. Blood 1989; 74: 1678–1689.
Giefing M, Siebert R . FISH and FICTION to detect chromosomal aberrations in lymphomas. Methods Mol Biol 2013; 971: 227–244.
Otto C, Giefing M, Massow A, Vater I, Gesk S, Schlesner M et al. Genetic lesions of the TRAF3 and MAP3K14 genes in classical Hodgkin lymphoma. Br J Haematol 2012; 157: 702–708.
Emmerich F, Theurich S, Hummel M, Haeffker A, Vry MS, Döhner K et al. Inactivating I kappa B epsilon mutations in Hodgkin/Reed-Sternberg cells. J Pathol 2003; 201: 413–420.
Giefing M, Winoto-Morbach S, Sosna J, Döring C, Klapper W, Küppers R et al. Hodgkin-Reed-Sternberg cells in classical Hodgkin lymphoma show alterations of genes encoding the NADPH oxidase complex and impaired reactive oxygen species synthesis capacity. PLoS One 2013; 8: e84928.
Steidl C, Telenius A, Shah SP, Farinha P, Barclay L, Boyle M et al. Genome-wide copy number analysis of Hodgkin Reed-Sternberg cells identifies recurrent imbalances with correlations to treatment outcome. Blood 2010; 116: 418–427.
Slovak ML, Bedell V, Hsu YH, Estrine DB, Nowak NJ, Delioukina ML et al. Molecular karyotypes of Hodgkin and Reed-Sternberg cells at disease onset reveal distinct copy number alterations in chemosensitive versus refractory Hodgkin lymphoma. Clin Cancer Res 2011; 17: 3443–3454.
Hinz M, Lemke P, Anagnostopoulos I, Hacker C, Krappmann D, Mathas S et al. Nuclear factor kappaB-dependent gene expression profiling of Hodgkin's disease tumor cells, pathogenetic significance, and link to constitutive signal transducer and activator of transcription 5a activity. J Exp Med 2002; 196: 605–617.
Stein H, Foss HD, Dürkop H, Marafioti T, Delsol G, Pulford K et al. CD30(+) anaplastic large cell lymphoma: a review of its histopathologic, genetic, and clinical features. Blood 2000; 96: 3681–3695.
Bonzheim I, Geissinger E, Roth S, Zettl A, Marx A, Rosenwald A et al. Anaplastic large cell lymphomas lack the expression of T-cell receptor molecules or molecules of proximal T-cell receptor signaling. Blood 2004; 104: 3358–3360.
Eckerle S, Brune V, Döring C, Tiacci E, Bohle V, Sundstrom C et al. Gene expression profiling of isolated tumour cells from anaplastic large cell lymphomas: insights into its cellular origin, pathogenesis and relation to Hodgkin lymphoma. Leukemia 2009; 23: 2129–2138.
Mathas S, Jöhrens K, Joos S, Lietz A, Hummel F, Janz M et al. Elevated NF-kappaB p50 complex formation and Bcl-3 expression in classical Hodgkin, anaplastic large-cell, and other peripheral T-cell lymphomas. Blood 2005; 106: 4287–4293.
Yan P, Fu J, Qu Z, Li S, Tanaka T, Grusby MJ et al. PDLIM2 suppresses human T-cell leukemia virus type I Tax-mediated tumorigenesis by targeting Tax into the nuclear matrix for proteasomal degradation. Blood 2009; 113: 4370–4380.
Huang B, Yang XD, Lamb A, Chen LF . Posttranslational modifications of NF-kappaB: another layer of regulation for NF-kappaB signaling pathway. Cell Signal 2010; 22: 1282–1290.
Crescenzo R, Abate F, Lasorsa E, Tabbo F, Gaudiano M, Chiesa N et al. Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell 2015; 27: 516–532.
Chiarle R, Voena C, Ambrogio C, Piva R, Inghirami G . The anaplastic lymphoma kinase in the pathogenesis of cancer. Nat Rev Cancer 2008; 8: 11–23.
Blum KA, Johnson JL, Niedzwiecki D, Canellos GP, Cheson BD, Bartlett NL . Single agent bortezomib in the treatment of relapsed and refractory Hodgkin lymphoma: cancer and leukemia Group B protocol 50206. Leuk Lymphoma 2007; 48: 1313–1319.
Younes A, Pro B, Fayad L . Experience with bortezomib for the treatment of patients with relapsed classical Hodgkin lymphoma. Blood 2006; 107: 1731–1732.
Kretzmer H, Bernhart SH, Wang W, Haake A, Weniger MA, Bergmann AK et al. DNA methylome analysis in Burkitt and follicular lymphomas identifies differentially methylated regions linked to somatic mutation and transcriptional control. Nat Genet 2015; 47: 1316–1325.
Acknowledgements
We thank Simone Kressmann and Brigitte Wollert-Wulf as well as the technical staff of the molecular cytogenetics and epigenetics laboratories at the Institute of Human Genetics in Kiel for excellent technical assistance. This work was supported in part by grants from the Deutsche Forschungsgemeinschaft to SM, MJ, GL and SH, the Experimental and Clinical Research Center, a joint cooperation between the Charité–Universitätsmedizin Berlin and the MDC as well as the German Cancer Consortium (DKTK). MG was supported by the FEBS ‘Long Term Fellowship’ and ‘Support for International Mobility of Scientists’ fellowship of the Polish Ministry of Science and Higher Education. RS and WK are supported by the KinderKrebsInitiative Buchholz/Holm-Seppensen and JR by the Dr Werner Jackstädt Foundation in the framework of a Junior Excellence Research Group (S134–10.100).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Wurster, K., Hummel, F., Richter, J. et al. Inactivation of the putative ubiquitin-E3 ligase PDLIM2 in classical Hodgkin and anaplastic large cell lymphoma. Leukemia 31, 602–613 (2017). https://doi.org/10.1038/leu.2016.238
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.238
This article is cited by
-
Tsc2 mutation rather than Tsc1 mutation dominantly causes a social deficit in a mouse model of tuberous sclerosis complex
Human Genomics (2023)
-
AP-1 family transcription factors: a diverse family of proteins that regulate varied cellular activities in classical hodgkin lymphoma and ALK+ ALCL
Experimental Hematology & Oncology (2021)