Abstract
Current prognostication in primary myelofibrosis (PMF) is based on the dynamic international prognostic scoring system (DIPSS)-plus, which employs clinical and cytogenetic variables. We recently reported DIPSS-plus independent prognostic significance for calreticulin (CALR) (favorable) and ASXL1 (unfavorable) mutations. In the current study, 570 PMF patients were recruited for derivation (n=277) and validation (n=293) of a molecular prognostic model based on these two mutations. Survival was the longest in CALR+ASXL1− (median 10.4 years) and shortest in CALR−ASXL1+ patients (median, 2.3 years; hazard ratio (HR), 5.9; 95% confidence interval (CI), 3.5–10.0). CALR+ASXL1+ and CALR–ASXL1− patients had similar survival and were grouped together in an intermediate-risk category (median survival, 5.8 years; HR, 2.5; 95% CI, 1.5–4.0). The CALR/ASXL1 mutations-based prognostic model was DIPSS-plus independent (P<0.0001) and effective in identifying low-/intermediate-1-risk patients with shorter (median, 4 years) or longer (median 20 years) survival and high-/intermediate-2-risk patients with shorter (median, 2.3 years) survival. Multivariable analysis distinguished CALR−ASXL1+ mutational status as the most significant risk factor for survival: HR 3.7 vs 2.8 for age >65 years vs 2.7 for unfavorable karyotype. These observations signify immediate clinical relevance and warrant i) CALR and ASXL1 mutation determination in all patients with PMF and ii) molecular revision of DIPSS-plus.
Similar content being viewed by others
Introduction
Karyotype and somatic mutations have a major part in disease prognostication and management of patients with myeloid malignancies. For example, the presence of unfavorable karyotype is prognostically detrimental in both acute myeloid leukemia (AML) and chronic myeloid neoplasms and is often an indication for treatment with allogeneic stem cell transplant (ASCT). The latter is also the preferred treatment of choice in AML associated with FLT3-ITD, whereas chemotherapy alone might be adequate for AML patients expressing NPM1 mutations without FLT3-ITD.1 Similarly, the prognostic relevance of mutations in myelodysplastic syndromes2 and primary myelofibrosis (PMF)3 has been recognized but not yet implemented in clinical practice.
Somatic mutations have now been incorporated into formal diagnostic criteria in myeloproliferative neoplasms (MPNs).4 JAK2 or MPL mutations are found in 50–70% of patients with PMF or essential thrombocythemia (ET) and calreticulin (CALR) mutations account for the majority of the remaining cases;5, 6 in strictly World Health Organization-defined disease, CALR mutations were seen in 49% of ET and 74% of PMF patients not expressing mutant JAK2 or MPL.7, 8 In ET, CALR mutations correlated with male sex, younger age, lower leukocyte count, lower hemoglobin level and higher platelet count8 and in PMF with younger age, higher platelet count and lower incidences of anemia, leukocytosis and spliceosome mutations.7 Furthermore, CALR mutations in ET were associated with longer thrombosis-free survival8, 9 and in PMF with longer overall survival.7
Before the discovery of CALR mutations in ET and PMF, we had identified mutant ASXL1 as dynamic international prognostic scoring system (DIPSS)-plus10 and IPSS11 independent risk factor for survival in PMF.3 More recently, we discovered the prognostic synergism between CALR and ASXL1 mutations in PMF and highlighted the inferior survival associated with ‘CALR−ASXL1+’ mutational status.7 The main objective of the current study was to further explore the prognostic interaction between CALR and ASXL1 mutations in PMF with the intent to derive (using a patient cohort from the Mayo Clinic, USA) and validate (using a patient cohort from the University of Florence, Italy) a molecular prognostic model, in the context both DIPSS-plus and IPSS.
Materials and methods
The current study was approved by the institutional review boards of the Mayo Clinic, Rochester, MN, USA and University of Florence, Florence, Italy. All patients provided informed written consent for study sample collection as well as permission for its use in research. Inclusion to the current study required availability of archived peripheral blood or bone marrow sample collected at the time of diagnosis or first referral; a total of 277 patients from the Mayo Clinic (the Mayo cohort) and 293 from the University of Florence (the Florence cohort) met these stipulations. The diagnoses of PMF and leukemic transformation were according to World Health Organization criteria.12 Unfavorable karyotype designation and DIPSS-plus or IPSS risk categorization were as previously described.10, 11, 13
We used published methods to screen for CALR, JAK2, MPL, ASXL1, EZH2, IDH1, IDH2 and spliceosome (SRSF2, SF3B1and U2AF1) mutations.7, 8, 14, 15, 16, 17, 18 All statistical analyses considered clinical and laboratory parameters obtained at time of diagnosis (Florence cohort) or first referral (Mayo cohort). Differences in the distribution of continuous variables between categories were analyzed by either Mann–Whitney (for comparison of two groups) or Kruskal–Wallis (comparison of three or more groups) test. Patient groups with nominal variables were compared by chi-square test. Overall survival analysis was considered from the date of diagnosis (Florence cohort) or first referral (Mayo cohort) to date of death (uncensored) or last contact (censored). Date of leukemic transformation replaced date of death, as the uncensored variable, for estimating leukemia-free survival. Overall and leukemia-free survival curves were prepared by the Kaplan–Meier method and compared by the log-rank test. Cox proportional hazard regression model was used for multivariable analysis. P-values<0.05 were considered significant. The Stat View (SAS Institute, Cary, NC, USA) statistical package was used for all calculations for the Mayo cohort and SPSS software was used for the Florence cohort.8
Results
A total of 570 study patients were recruited from two centers: 277 from the Mayo Clinic (derivation cohort) and 293 from the University of Florence (validation cohort). Tables 1 and 2 summarize the presenting clinical and laboratory characteristics of the two patient cohorts, respectively.
Mayo Clinic patients (derivation cohort)
Phenotypic correlates
The information in Table 1 confirms our previous observations regarding the phenotypic correlates of CALR mutations in PMF, including younger age, higher platelet count, lower leukocyte count, lower frequency of anemia, lower DIPSS-plus risk distribution and lower incidence of spliceosome mutations.7 The presence of ASXL1 mutations in CALR-mutated patients was associated with the male gender (P=0.03), lower platelet count (P=0.03), lower hemoglobin level (P=0.01), higher transfusion requirement (P=0.04), higher circulating blast percentage (P=0.02), higher incidence of constitutional symptoms (P=0.007) and higher DIPSS-plus risk distribution (P=0.04). In other words, the CALR phenotype in PMF appeared to be adversely affected by the presence of ASXL1 mutations.
Prognostic evaluation of CALR, ASXL1 and other relevant mutations
In multivariable analysis that included all prognostically relevant mutations,3 absence of CALR (hazard ratio (HR), 2.6; 95% confidence interval (CI), 1.8–3.9), presence of ASXL1 (HR, 2.0; 95% CI, 1.5–2.7) and presence of SRSF2 (HR, 1.7; 95% CI, 1.1–2.8) mutations were significantly associated with shortened survival; EZH2 (P=0.07) or IDH (P=0.48) mutations were not significant. Absence of CALR (HR, 2.3; 95% CI, 1.5–3.5) and presence of ASXL1 (HR, 1.5; 95% CI, 1.1–2.1) mutations remained significant when DIPSS-plus risk stratification was added into the multivariable model. CALR and ASXL1 mutational status were then considered together to further enhance their prognostic contribution; the longest survival was seen in CALR+ASXL1− patients (median, 10.4 years), which was significantly better than that of CALR−ASXL1+ patients (median, 2.3 years; HR, 5.9; 95% CI, 3.5–10.0) and CALR−ASXL1− patients (median, 5.4 years; HR, 2.3, 95% CI, 1.6–4.2). The difference in survival between CALR+ASXL1− and CALR+ASXL1+ patients (median, 7.8 years; HR, 1.7; 95% CI, 0.9–3.6) was of borderline significance (P=0.13). On the other hand, survival was not significantly different between patients with CALR+ASXL1+ and CALR−ASXL1− mutational status (P=0.2).
Construction of a prognostic model based on CALR and ASXL1 mutational status
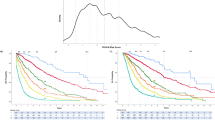
Based on the above-stated observations, we constructed a three-tier molecular risk stratification on the basis of CALR/ASXL1 mutational status: low risk included CALR+ASXL1− patients (median survival, 10.4 years); high risk included CALR−ASXL1+ patients (median survival, 2.3 years; HR, 5.9; 95% CI, 3.5–10.0); and intermediate risk included either CALR+ASXL1+ or CALR−ASXL1 (median survival, 5.8 years; HR, 2.5; 95% CI, 1.5–4.0). This CALR/ASXL1 mutations-based prognostic model (Figure 1) was DIPSS-plus independent (P<0.0001) and highly effective in identifying short- (median survival, 4 years; HR, 10.3; 95% CI, 3.3–31.9) and long-term (median survival, 20 years) survivors with DIPSS-plus low-/intermediate-1-risk disease (Figure 2) and short-term survivors with high-/intermediate-2-risk disease (median survival, 2.0 years; HR 2.0; 95% CI, 1.1–3.7) (Figure 3). Conversely, DIPSS-plus risk stratification showed added value in molecularly low (P=0.0005) or intermediate (P<0.0001) risk disease but not in high-risk disease (P=0.21).
Kaplan–Meier estimates of overall survival in 88 Mayo Clinic patients with low- or intermediate-1-risk PMF, according to the DIPSS-plus,10 stratified by the presence or absence of CALR and ASXL1 mutations.
Kaplan–Meier estimates of overall survival in 189 Mayo Clinic patients with high- or intermediate-2-risk PMF, according to the DIPSS-plus,8 stratified by the presence or absence of CALR and ASXL1 mutations.
Multivariable analysis that included the CALR/ASXL1 mutations-based molecular risk stratification along with all eight DIPSS-plus risk factors distinguished CALR−ASXL1+ mutational status as the most significant risk factor for survival (HR, 3.7) followed by age >65 years (HR, 2.8) and unfavorable karyotype (HR, 2.7). In this model, constitutional symptoms and circulating blast percentage lost their significance, whereas anemia, leukocytosis and thrombocytopenia remained significant. CALR−ASXL1+ mutational status was also associated with inferior leukemia-free survival (P=0.04; HR, 3.3; 95% CI, 1.1–10.0) and its significance was reduced to borderline status (P=0.06) during multivariable analysis that included unfavorable karyotype and platelet count <100 × 109/l, which are previously recognized risk factors for leukemic transformation.10
University of Florence patients (validation cohort)
The phenotypic correlates of CALR mutations in the Florence cohort (Table 2) were mostly similar to those noted in the Mayo cohort (Table 1), but also showed some differences. As was the case in the Mayo cohort, CALR-mutated Florence patients were younger and displayed higher platelet count and lower IPSS risk scores, whereas the associations with leukocyte count, hemoglobin level and constitutional symptoms were less evident. As was the case in the Mayo cohort, the presence of ASXL1 mutations in CALR-mutated cases was detrimental and associated with higher rate of marked leukocytosis, circulating blast percentage and thrombocytopenia. In multivariable analysis that included IPSS risk scores, presence of mutant CALR had a favorable (HR, 0.5; 95% CI, 0.2–0.98) and mutant ASXL1 an unfavorable (HR, 2.3; 95% CI, 1.4–3.8) impact on survival. A similar IPSS-inclusive multivariable analysis confirmed the independent prognostic value of the three-tier CALR/ASXL1 risk stratification: HR (95% CI) were 6.4 (2.2–18.9) for CALR−ASXL1+ patients and 3.4 (1.2–9.4) for patients with CALR+ASXL1+ or CALR−ASXL1− mutational status.
Application of the Mayo cohort-derived CALR/ASXL1 mutations-based prognostic model was equally effective in delineating Florence patients with significant survival differences (Figure 4); median survivals were ‘not reached’ in low-risk patients (CALR+ASXL1−), 11.5 years (HR, 3.2; 95% CI, 1.3–8.1) in intermediate-risk patients (CALR+ASXL1+ or CALR−ASXL1) and 3.2 years (HR, 8.7; 95% CI, 3.3–23.1) in high-risk patients (CALR−ASXL1+). The CALR/ASXL1 mutations-based prognostic model was also effective in identifying short- (median survival 6 years; HR, 18.7; 95% CI, 2.3–153.9) and long-term (median survival not reached) survivors with IPSS low-/intermediate-1-risk disease (Figure 5) and short-term survivors with high-/intermediate-2-risk disease (median survival 3.1 years for molecular high risk and 3.7 years for molecular intermediate risk; HR 3.9, 95% CI 1.1–13.4 for molecular high risk and 3.0, 95% CI 0.9–9.8 for molecular intermediate risk) (Figure 6). In the Florence cohort, both CALR−ASXL1+ (HR, 8.0; 95% CI, 1.7–38.2) and CALR+ASXL1+ or CALR−ASXL1− (HR, 4.5; 95% CI, 1.04–19.1) patients had inferior leukemia-free survival, compared with CALR+ASXL1− patients.
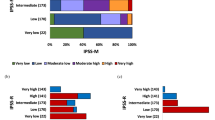
Kaplan–Meier estimates of overall survival in patients of the Italian series (n=293) stratified according to the mutational status of CALR and ASXL1. The three tiers included CALR+/ASXL1− (n=45), CALR−/ASXL1+ (n=46) and CALR−/ASXL1− or CALR+/ASXL1+ (n=202). The different survival curves were statistically significant, P<0.0001. The HR is presented together with the 95% CI, using CALR+/ASXL1− as the reference population (HR=1.0).
Kaplan–Meier estimates of overall survival in patients of the Italian series belonging to IPSS11 low- and intermediate-1-risk category (n=180) stratified according to the mutational status of CALR and ASXL1. The three tiers included CALR+/ASXL1− (n=30), CALR−/ASXL1+ (n=16) and CALR−/ASXL1− or CALR+/ASXL1+ (n=134). The different survival curves were statistically significant, P=0.001. The HR is presented together with the 95% CI, using CALR+/ASXL1− as the reference population (HR=1.0).
Kaplan–Meier estimates of overall survival in patients of the Italian series belonging to the IPSS11 intermediate-2- and high-risk category (n=92) stratified according to the mutational status of CALR and ASXL1. The three tiers included CALR+/ASXL1− (n=11), CALR−/ASXL1+ (n=28) and CALR−/ASXL1− or CALR+/ASXL1+ (n=53). The different survival curves were trend significant, P=0.07 The HR is presented together with the 95% CI, using CALR+/ASXL1− as the reference population (HR=1.0).
Discussion
Our observations from the current study of 570 patients with PMF confirm the prognostic advantage of harboring a CALR mutation, especially in the absence of a concomitantly occurring ASXL1 mutation. Survival was best in the presence of CALR and absence of ASXL1 mutation (that is, CALR+ASXL1−) and worst otherwise (that is, CALR−ASXL1+). This is somewhat similar to the scenario in AML with NPM1+FLT3-ITD− vs NPM1−FLT3-ITD+ mutational status, respectively.1 Furthermore, the presence of CALR mutations appears to attenuate, but not fully overcome, the unfavorable prognosis in ASXL1-mutated patients (that is, CALR+ASXL1+). Conversely, the absence of ASXL1 mutations is associated with better survival, even in CALR-unmutated cases (that is, CALR−ASXL1−). DIPSS-plus had limited added value in molecularly defined high-risk patients (that is, CALR−ASXL1+) but was effective in identifying short-lived patients among the molecularly low- (that is, CALR+ASXL1−) and intermediate-risk (that is, CALR−ASXL1− or CALR+ASXL1+) groups. On the other hand, although the value of molecular risk stratification was most evident in DIPSS-plus low-/intermediate-1-risk patients, it was also apparent in high-/intermediate-2-risk patients. These results were validated in an independent patient cohort and performed similarly in the context of IPSS.
How does one translate this new information into clinical practice? First, our observations provide strong evidence for the prognostic relevance of performing CALR and ASXL1 mutation determination in all patients with PMF. Second, survival in PMF is significantly compromised (median <2.5 years) in the presence of either CALR−ASXL1+ mutation status or DIPSS-plus high-risk score. In other words, a high-risk molecular signature identifies DIPSS-plus low- or intermediate-1-risk patients whose survival might not be better than those with DIPSS-plus high-risk disease. Therefore, the risk of aggressive therapy, including ASCT, might be justified not only in DIPSS-plus high or intermediate-2-risk disease but also in low or intermediate-1-risk patients who harbor a high-risk molecular signature (that is, CALR−ASXL1+). On the other hand, aggressive therapy might be less urgent in DIPSS-plus intermediate-2-risk patients who do not express high-risk mutational profile (that is, CALR−ASXL1+). We were particularly gratified with the highly indolent nature of the disease (median survival ∼20 years) in DIPSS-plus low/intermediate-1-risk patients with low-risk molecular profile (that is, CALR+ASXL1−) and the risk of ASCT or investigational drug therapy may not be justified in such patients. A similar scenario was confirmed in an independent cohort of patients, in the context of IPSS.
Drug therapy in PMF remains inadequate and there is no firm evidence yet that, beyond their value in reducing splenomegaly and improving symptoms, JAK inhibitors provide survival advantage.19 In other words, there is a dire need for new drugs with disease-modifying activity and continuing value for ASCT. To that effect, accurate disease prognostication for the selection of appropriate patients for specific treatment strategies is of paramount importance. The International Working Group for MPN research and treatment (IWG-MRT) has led the effort in this regard with the development of the IPSS in 2009.11 IPSS was first revised to dynamic IPSS (DIPSS)20 and subsequently to DIPSS-plus.10 A key element of DIPSS-plus is karyotype, whose prognostic value was further highlighted by the more recent identification of ‘very high risk’ cytogenetic abnormalities, which included monosomal karyotype and inv(3)/i(17q) abnormalities.21 The observations from the current study signify the additional importance of molecular information in disease prognostication in PMF and provide the basis for revising DIPSS-plus accordingly.
It is difficult and somewhat premature to speculate on the biological explanation for our observation because of our limited insight on the pathogenetic contribution of either CALR or ASXL1 mutations. The adverse prognostic effect of ASXL1 mutations is not specific to PMF and has been realized across different types of myeloid malignancies. Mutant ASXL1 or loss of ASXL1 induces myelodysplastic syndrome-like disease in mice, putatively through loss of PRC2-mediated histone methylation, which in turn leads to increased expression of HOXA9 and microRNA-125a.22, 23 Mutant CALR displays a significantly modified C terminal that is less acidic and lacks the endoplasmic reticulum retention motif. Currently, little is known about the consequences of CALR mutations on the function of wild-type protein, which includes regulation of intracellular calcium homeostasis and the control of protein folding/misfolding.24 Transfection of mutant CALR into a Ba/F3 murine cell line was shown to induce IL-3-independent growth that was associated with increased STAT5 phosphorylation.6 However, the particular observation needs to be validated and further elaborated upon. Regardless, it is possible that a CALR, compared with a JAK2-mutated clone is genetically more stable and biologically more capable of handling the epigenetic alteration associated with mutant ASXL1.
References
Hou HA, Lin CC, Chou WC, Liu CY, Chen CY, Tang JL et al. Integration of cytogenetic and molecular alterations in risk stratification of 318 patients with de novo non-M3 acute myeloid leukemia. Leukemia 2014; 28: 50–58.
Bejar R, Stevenson K, Abdel-Wahab O, Galili N, Nilsson B, Garcia-Manero G et al. Clinical effect of point mutations in myelodysplastic syndromes. New Engl J Med 2011; 364: 2496–2506.
Vannucchi AM, Lasho TL, Guglielmelli P, Biamonte F, Pardanani A, Pereira A et al. Mutations and prognosis in primary myelofibrosis. Leukemia 2013; 27: 1861–1869.
Tefferi A, Thiele J, Vannucchi AM, Barbui T . An overview on CALR and CSF3R mutations and a proposal for revision of WHO diagnostic criteria for myeloproliferative neoplasms. Leukemia 2014; e-pub ahead of print 20 January 2014; doi:10.1038/leu.2014.35.
Nangalia J, Massie CE, Baxter EJ, Nice FL, Gundem G, Wedge DC et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. New Engl J Med 2013; 369: 2391–2405.
Klampfl T, Gisslinger H, Harutyunyan AS, Nivarthi H, Rumi E, Milosevic JD et al. Somatic mutations of calreticulin in myeloproliferative neoplasms. New Engl J Med 2013; 369: 2379–2390.
Tefferi A, Lasho TL, Finke CM, Knudson RA, Ketterling R, Hanson CH et al. CALR vs JAK2 vs MPL mutated or triple-negative myelofibrosis: clinical, cytogenetic and molecular comparisons. Leukemia 2014; e-pub ahead of print 9 January 2014; doi:10.1038/leu.2014.3.
Rotunno G, Mannarelli C, Guglielmelli P, Pacilli A, Pancrazzi A, Pieri L et al. Impact of calreticulin mutations on clinical and hematological phenotype and outcome in essential thrombocythemia. Blood 2013; e-pub ahead of print 26 December 2013.
Rumi E, Pietra D, Ferretti V, Klampfl T, Harutyunyan AS, Milosevic JD et al. JAK2 or CALR mutation status defines subtypes of essential thrombocythemia with substantially different clinical course and outcomes. Blood 2013; e-pub ahead of print 23 December 2013.
Gangat N, Caramazza D, Vaidya R, George G, Begna K, Schwager S et al. DIPSS plus: a refined Dynamic International Prognostic Scoring System for primary myelofibrosis that incorporates prognostic information from karyotype, platelet count, and transfusion status. J Clin Oncol 2011; 29: 392–397.
Cervantes F, Dupriez B, Pereira A, Passamonti F, Reilly JT, Morra E et al. New prognostic scoring system for primary myelofibrosis based on a study of the International Working Group for Myelofibrosis Research and Treatment. Blood 2009; 113: 2895–2901.
Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood 2009; 114: 937–951.
Caramazza D, Begna KH, Gangat N, Vaidya R, Siragusa S, Van Dyke DL et al. Refined cytogenetic-risk categorization for overall and leukemia-free survival in primary myelofibrosis: a single center study of 433 patients. Leukemia 2011; 25: 82–88.
Patnaik MM, Padron E, LaBorde RR, Lasho TL, Finke CM, Hanson CA et al. Mayo prognostic model for WHO-defined chronic myelomonocytic leukemia: ASXL1 and spliceosome component mutations and outcomes. Leukemia 2013; 27: 1504–1510.
Patnaik MM, Lasho TL, Finke CM, Hanson CA, Hodnefield JM, Knudson RA et al. Spliceosome mutations involving SRSF2, SF3B1, and U2AF35 in chronic myelomonocytic leukemia: prevalence, clinical correlates, and prognostic relevance. American Journal of Hematology 2013; 88: 201–206.
Tefferi A, Finke CM, Lasho TL, Wassie EA, Knudson R, Ketterling RP et al. U2AF1 mutations in primary myelofibrosis are strongly associated with anemia and thrombocytopenia despite clustering with JAK2V617F and normal karyotype. Leukemia 2013; 28: 431–433.
Lasho TL, Jimma T, Finke CM, Patnaik M, Hanson CA, Ketterling RP et al. SRSF2 mutations in primary myelofibrosis: significant clustering with IDH mutations and independent association with inferior overall and leukemia-free survival. Blood 2012; 120: 4168–4171.
Tefferi A, Jimma T, Sulai NH, Lasho TL, Finke CM, Knudson RA et al. IDH mutations in primary myelofibrosis predict leukemic transformation and shortened survival: clinical evidence for leukemogenic collaboration with JAK2V617F. Leukemia 2012; 26: 475–480.
Pardanani A, Vannucchi AM, Passamonti F, Cervantes F, Barbui T, Tefferi A . JAK inhibitor therapy for myelofibrosis: critical assessment of value and limitations. Leukemia 2011; 25: 218–225.
Passamonti F, Cervantes F, Vannucchi AM, Morra E, Rumi E, Pereira A et al. A dynamic prognostic model to predict survival in primary myelofibrosis: a study by the IWG-MRT (International Working Group for Myeloproliferative Neoplasms Research and Treatment). Blood 2010; 115: 1703–1708.
Tefferi A, Jimma T, Gangat N, Vaidya R, Begna KH, Hanson CA et al. Predictors of greater than 80% 2-year mortality in primary myelofibrosis: a Mayo Clinic study of 884 karyotypically annotated patients. Blood 2011; 118: 4595–4598.
Inoue D, Kitaura J, Togami K, Nishimura K, Enomoto Y, Uchida T et al. Myelodysplastic syndromes are induced by histone methylation-altering ASXL1 mutations. J Clin Invest 2013; 123: 4627–4640.
Abdel-Wahab O, Gao J, Adli M, Dey A, Trimarchi T, Chung YR et al. Deletion of Asxl1 results in myelodysplasia and severe developmental defects in vivo. J Exp Med 2013; 210: 2641–2659.
Eggleton P, Michalak M . Calreticulin for better or for worse, in sickness and in health, until death do us part. Cell Calcium 2013; 54: 126–131.
Acknowledgements
This study was supported by the Mayo Clinic Harvey-Yulman Charitable Foundation for Myelofibrosis Tissue Bank and Clinical Database of Molecular and Biological Abnormalities and by a special grant from Associazione Italiana per la Ricerca sul Cancro-‘AIRC 5 per Mille’- to AGIMM, ‘AIRC-Gruppo Italiano Malattie Mieloproliferative’ (no. 1005) to AMV; for a description of the AGIMM project, see at http://www.progettoagimm.it). Partially supported by Ministero della Università e Ricerca (MIUR; FIRB project #RBAP11CZLK and PRIN 2010NYKNS7 to AMV).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Tefferi, A., Guglielmelli, P., Lasho, T. et al. CALR and ASXL1 mutations-based molecular prognostication in primary myelofibrosis: an international study of 570 patients. Leukemia 28, 1494–1500 (2014). https://doi.org/10.1038/leu.2014.57
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.57
This article is cited by
-
CALR mutations possess unique prognostic relevance in myelofibrosis—before and after transplant
Bone Marrow Transplantation (2024)
-
Evolutionary signatures of human cancers revealed via genomic analysis of over 35,000 patients
Nature Communications (2023)
-
Biological drivers of clinical phenotype in myelofibrosis
Leukemia (2023)
-
Improving allogeneic stem cell transplantation in myelofibrosis
International Journal of Hematology (2022)
-
Prognostic value of ASXL1 mutations in patients with primary myelofibrosis and its relationship with clinical features: a meta-analysis
Annals of Hematology (2021)