Abstract
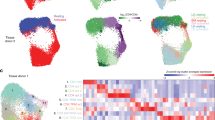
Myeloperoxidase (MPO) has been associated with both a myeloid lineage commitment and favorable prognosis in patients with acute myeloid leukemia (AML). DNA methyltransferase inhibitors (decitabine and zeburaline) induced MPO gene promoter demethylation and MPO gene transcription in AML cells with low MPO activity. Therefore, MPO gene transcription was directly and indirectly regulated by DNA methylation. A DNA methylation microarray subsequently revealed a distinct methylation pattern in 33 genes, including DNA methyltransferase 3 beta (DNMT3B), in CD34-positive cells obtained from AML patients with a high percentage of MPO-positive blasts. Based on the inverse relationship between the methylation status of DNMT3B and MPO, we found an inverse relationship between DNMT3B and MPO transcription levels in CD34-positive AML cells (P=0.0283). In addition, a distinct methylation pattern was observed in five genes related to myeloid differentiation or therapeutic sensitivity in CD34-positive cells from AML patients with a high percentage of MPO-positive blasts. Taken together, the results of the present study indicate that MPO may serve as an informative marker for identifying a distinct and crucial DNA methylation profile in CD34-positive AML cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hoyle CF, Gray RG, Wheatley K, Wheatley K, Swirsky D, de Bastos M et al. Prognostic importance of Sudan Black positivity: a study of bone marrow slides from 1386 patients with de novo acute myeloid leukemia. Br J Haematol 1991; 79: 398–407.
Matsuo T, Cox C, Bennett JM . Prognostic significance of myeloperoxidase positivity of blast cells in acute myeloblastic leukemia without maturation (FAB: M1): an ECOG study. Hematol Pathol 1989; 3: 153–158.
Matsuo T, Kuriyama K, Miyazaki Y, Yoshida S, Tomonaga M, Emi N et al. The percentage of myeloperoxidase-positive blast cells is a strong independent prognostic factor in acute myeloid leukemia, even in the patients with normal karyotype. Leukemia 2003; 17: 1538–1543.
Miyawaki S, Sakamaki H, Ohtake S, Emi N, Yagasaki F, Mitani K et al. A randomized, postremission comparison of four courses of standard-dose consolidation therapy without maintenance therapy versus three courses of standard-dose consolidation with maintenance therapy in adults with acute myeloid leukemia. Cancer 2005; 104: 2726–2734.
Ohtake S, Miyawaki S, Fujita H, Kiyoi H, Shinagawa K, Usui N et al. Randomized study of induction therapy comparing standard-dose idarubicin with high-dose daunorubicin in adult patients with previously untreated acute myeloid leukemia: the JALSG AML201 Study. Blood 2011; 117: 2358–2365.
Sawayama Y, Miyazaki Y, Ando K, Horio K, Tsutsumi C, Imanishi D et al. Expression of myeloperoxidase enhances the chemosensitivity of leukemia cells through the generation of reactive species and the nitration of protein. Leukemia 2008; 22: 956–964.
Nakazato T, Sagawa M, Yamato K, Xian M, Yamamoto T, Suematsu M et al. Myeloperoxidase is a key regulator of oxidative stress mediated apoptosis in myeloid leukemic cells. Clin Cancer Res 2007; 13: 5436–5445.
Taguchi J, Miyazaki Y, Tsutsumi C, Sawayama Y, Ando K, Tsushima H et al. Expression of the myeloperoxidase gene in AC133 positive leukemia cells relates to the prognosis of acute myeloid leukemia. Leuk Res 2006; 30: 1105–1112.
Austin GE, Zhao W-G, Zhang W, Austin ED, Findley HW, Murtagh JJ . Identification and characterization of the human myeloperoxidase promoter. Leukemia 1995; 9: 848–857.
Zhao W-G, Regmi A, Austin ED, Braun JE, Racine M, Austin GE . Cis-elements in the promoter region of the human myeloperoxidase gene. Leukemia 1996; 10: 1089–1103.
Zhao W-G, Lu J-P, Austin GE . Enhancement of myeloperoxidase promoter function by PEBP2/CBF or inhibition by an Sp1-containing repressor requires the participation of a third element DP4. Blood 1996; 88 (Suppl 1): 47a.
Zhao W-G, Regmi A, Lu J-P, Austin GE . Identification and functional analysis of multiple murine myeloperoxidase (MPO) promoters and comparison with the human MPO promoter region. Leukemia 1997; 11: 97–105.
Austin GE, Zhao WG, Regmi A, Lu JP, Braun J . Identification of an upstream enhancer containing an AML1 site in the human myeloperoxidase (MPO) gene. Leuk Res 1998; 22: 1037–1048.
Yao C, Qin Z, Works KN, Austin GE, Young AN . C/EBP and C-Myb sites are important for the functional activity of the human myeloperoxidase upstream enhancer. Biochem Biophys Res Commun 2008; 371: 309–314.
Khoury H, Dalal BI, Nantel SH, Horsman DE, Lavoie JC, Shepherd JD et al. Correlation between karyotype and quantitative immunophenotype in acute myelogenous leukemia with t(8;21). Mod Pathol 2004; 17: 1211–1216.
Shimada H, Ichikawa H, Ohki M . Potential involvement of the AML1-MTG8 fusion protein in the granulocytic maturation characteristic of the t(8;21) acute myelogenous leukemia revealed by microarray analysis. Leukemia 2002; 16: 874–885.
Meyers S, Lenny N, Hiebert SW . The t(8;21) fusion protein interferes with AML-1B-dependent transcriptional activation. Mol Cell Biol 1995; 15: 1974–1982.
Tominaga-Sato S, Tsushima H, Ando K, Itonaga H, Imaizumi Y, Imanishi D et al. Expression of myeloperoxidase and gene mutations in AML patients with normal karyotype: double CEBPA mutations are associated with high percentage of MPO positivity in leukemic blasts. Int J Hematol 2011; 94: 81–89.
Kato N, Kitamura J, Doki N, Komeno Y, Watanabe-Okochi N, Togami K et al. Two types of C/EBPα mutations play distinct but collaborative roles in leukemogenesis; lessons from clinical data and BMT models. Blood 2011; 117: 221–233.
Momparler RL, Bovenzi V . DNA methylation and cancer. J Cell Physiol 2000; 183: 145–154.
Issa JP . Methylation and prognosis: of molecular clocks and hypermethylation phenotypes. Clin Cancer Res 2003; 9: 2879–2881.
Jones PA, Baylin SB . The epigenetic of cancer. Cell 2007; 128: 683–692.
Schmelz K, Sattler N, Wagner M, Lubbert M, Dorken B, Tamm I . Induction of gene expression by 5-Aza-2’-deoxycytidine in acute myeloid leukemia (AML) and myelodysplastic syndrome (MDS) but not epithelial cells by DNA-methylation-dependent and –independent mechanisms. Leukemia 2005; 19: 103–111.
Figueroa ME, Lugthart S, Li Y, Erpelinck-Verschueren C, Deng X, Christos PJ et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell 2010; 17: 13–27.
Bibikova M, Le J, Barnes B, Saedinia-Melnyk S, Zhou L, Shen R et al. Genome-wide DNA methylation profiling using infinium assay. Epigenomics 2009; 1: 177–200.
Gunderson KL, Steemers FJ, Ren H, Ng P, Zhou L, Tsan C et al. Whole-genome genotyping. Methods Enzymol 2006; 410: 359–376.
Steemers FJ, Gunderson KL . Whole genome genotyping technologies on the BeadArray platform. Biotechnol J 2007; 2: 41–49.
Yang AS, Estecio MR, Doshi K, Kondo Y, Tajara EH, Issa JP . A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res 2004; 32: e38.
Okano M, Xie S, Li E . Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet 1998; 19: 219–220.
Aimiuwu J, Wang H, Chen P, Xie Z, Wang J, Liu S et al. RNA-dependent inhibition of ribonucleotide reductase is a major pathway for 5-azacitidine activity in acute myeloid leukemia. Blood 2012; 119: 5229–5238.
Aoyama S, Nakano H, Danbara M, Higashihara M, Harigae H, Takahashi S . The differentiation and apoptotic effects of 2-aza-5’deoxycytidine are dependent on the PU.1 expression level in PU.1-transgenic K562 cells. Biochem Biophys Res Commun 2012; 420: 775–781.
Curik N, Burda P, Vargova K, Pospisil V, Belickova M, Vlckova P et al. 5-azacitidine in aggressive myelodysplastic syndromes regulates chromatin structure at PU.1 gene and cell differentiation capacity. Leukemia 2012; 26: 1804–1811.
Kulis M, Heath S, Bibikova M, Queiros AC, Navarro A, Clot G et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat Genet 2012; 44: 1236–1242.
Razin A, Riggs AD . DNA methylation and gene function. Science 1980; 210: 604–610.
Robertson KD . DNA methylation, methyltransferases, and cancer. Oncogene 2001; 20: 3139–3155.
Arand J, Spieler D, Karius T, Branco MR, Meilinger D, Meissner A et al. In vitro control of CpG and non-CpG DNA methylation by DNA methyltransferases. PLoS Genet 2012; 8: e1002750.
Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med 2010; 363: 2424–2433.
Marcucci G, Metzeler KH, Schwind S, Becker H, Maharry K, Mrozek K et al. Age-related prognostic impact of different types of DNMT3A mutations in adults with primary cytogenetically normal acute myeloid leukemia. J Clin Oncol 2012; 30: 742–750.
Yan XJ, Xu J, Gu ZH, Pan CM, Lu G, Shen Y et al. Exome sequencing identifies somatic mutations of DNA methyltransferase gene DNMT3A in acute monocytic leukemia. Nat Genet 2011; 43: 309–315.
Yamashita Y, Yuan J, Suetake I, Suzuki H, Ishikawa Y, Choi YL et al. Array-based genomic resequencing of human leukemia. Oncogene 2010; 29: 3723–3731.
Kim SJ, Zhao H, Hardikar S, Singh AK, Goodell MA, Chen T . A DNMT3A mutation common in AML exhibits dominant-negative effects in murine ES cells. Blood 2013; 122: 4086–4089.
Thol F, Damm F, Ludeking A, Winschel C, Wagner K, Morgan M et al. Incidence and prognostic influence of DNMT3A mutations in acute myeloid leukemia. J Clin Oncol 2011; 29: 2889–2896.
Robertson KD, Uzvolgyi E, Liang G, Talmadge C, Sumegi J, Gonzales FA et al. The human DNA methyltransferase (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissue and overexpression in tumors. Nucleic Acids Res 1999; 27: 2291–2298.
Girault I, Tozlu S, Lidereau R, Bieche I . Expression analysis of DNA methyltransferase 1, 3A, and 3B in sporadic breast carcinomas. Clin Cancer Res 2003; 9: 4415–4422.
Roll JD, Rivenbark AG, Jones WD, Coleman WB . DNMT3b overexpression contributes to a hypermethylator phenotype in human breast cancer cell lines. Mol Cancer 2008; 7: 15.
Mizuno S, Chijiwa T, Okamura T, Akashi K, Fukumaki Y, Niho Y et al. Expression of DNA methyltransferases DNMT1, 3A, and 3B in normal hematopoiesis and in acute and chronic myelogenous leukemia. Blood 2001; 97: 1172–1179.
Hayette S, Thomas X, Jallades L, Chabane K, Charlot C, Tigaud I et al. High DNA methyltransferase DNMT3B levels: a poor prognostic marker in acute myeloid leukemia. PLoS One 2012; 7: e51527.
Shah MY, Vasanthankumar A, Barnes NY, Figueroa ME, Kamp A, Hendrick C et al. DNMT3B7, a truncated DNMT3B isoform expressed in human tumors, disrupts embryonic development and accelerates lymphomagenesis. Cancer Res 2010; 70: 5840–5850.
Gopalakrishnan S, Van Emburgh BO, Shan J, Su Z, Fields CR, Vieweg J et al. A novel DNMT3B splice variant expressed in tumor and pluripotent cells modulates genomic DNA methylation patterns and displays altered DNA binding. Mol Cancer Rs 2009; 7: 1622–1634.
Van Emburgh BO, Robertson KD . Modulation of DNMT3b function in vitro by interactions with Dnmt3L, Dnmt3a and Dnmt3b splice variants. Nucleic Acids Res 2001; 39: 4984–5002.
Gopalakrishna-Pilai S, Iverson LE . A DNMT3B alternatively spliced exon and encoded peptide are novel biomarkers of human pluripotent stem cells. PLoS One 2011; 6: e20663.
Wu CH, Gordon J, Rastegar M, Ogretmen B, Safa AR . Proteinase-3, a serine protease which mediates doxorubicin-induced apoptosis in the HL-60 leukemia cell line, is downregulated in its doxorubicin-resistant variant. Oncogene 2002; 21: 5160–5174.
Aqirre X, Roman-Gomez J, Jimenez-Velasco A, Garate L, Montiel-Duarte C, Navarro G et al. ASPP1, a common activator of TP53, is inactivated by aberrant methylation of its promoter in acute lymphoblastic leukemia. Oncogene 2006; 25: 1862–1870.
Fazi F, Rosa A, Fatica A, Gelmetti V, De Marchis ML, Nervi C et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell 2005; 123: 819–831.
Taketani T, Taki T, Shibuya N, Kikuchi A, Hanada R, Hayashi Y . Novel NUP98-HOXC11 fusion gene resulted from a chromosomal break within exon 1 of HOXC11 in acute myeloid leukemia with t(11;12)(p15;q13). Cancer Res 2002; 62: 4571–4574.
Lin M, Sutherland DR, Horsfall W, Totty N, Yeo E, Nayar R et al. Cell surface antigen CD109 is a novel member of the alpha(2) macroglobulin/C3, C4, C5 family of thioester-containing proteins. Blood 2002; 99: 1683–1691.
Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Masse A et al. Mutation in TET2 in myeloid cancers. N Engl J Med 2009; 360: 2289–2301.
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med 2009; 361: 1058–1066.
Carbuccia N, Trouplin V, Gelsi-Boyer V, Murati A, Rocquain J, Adelaide J et al. Mutuali exclusion of ASXL1 and NPM1 mutations in a series of acute myeloid leukemias. Leukemia 2010; 24: 469–473.
Shih AH, Abdel-Wahab O, Patel JP, Levine RL . The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer 2012; 12: 599–612.
Acknowledgements
We thank Ms M Yamaguchi and Ms H Urakami for their technical assistance. This work was partly supported by the grant from the Ministry of Health, Labour and Welfare of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Author contributions
HI and Y Miyazaki conceived and designed the study; HI, DI, WYF, SS, KA, YS, DS, KT, HH, YI, JT, HT, SY, TF, TH, Y Moriuchi, KY and Y Miyazaki collected and analyzed the samples and data; HI and Y Miyazaki performed the statistical analysis, wrote the manuscript and created the figures and tables; and all authors critically reviewed the manuscript and read and approved the final version.
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Itonaga, H., Imanishi, D., Wong, YF. et al. Expression of myeloperoxidase in acute myeloid leukemia blasts mirrors the distinct DNA methylation pattern involving the downregulation of DNA methyltransferase DNMT3B. Leukemia 28, 1459–1466 (2014). https://doi.org/10.1038/leu.2014.15
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.15
Keywords
This article is cited by
-
Comprehensive comparison between azacytidine and decitabine treatment in an acute myeloid leukemia cell line
Clinical Epigenetics (2022)
-
DNA methylation: a saga of genome maintenance in hematological perspective
Human Cell (2022)
-
Super-enhancer profiling identifies novel critical and targetable cancer survival gene LYL1 in pediatric acute myeloid leukemia
Journal of Experimental & Clinical Cancer Research (2022)
-
LINC00649 underexpression is an adverse prognostic marker in acute myeloid leukemia
BMC Cancer (2020)
-
Demethylator phenotypes in acute myeloid leukemia
Leukemia (2018)