Abstract
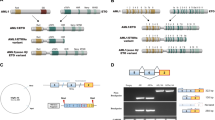
The t(8;21)(q22;q22) rearrangement represents the most common chromosomal translocation in acute myeloid leukemia (AML). It results in a transcript encoding for the fusion protein AML1-ETO (AE) with transcription factor activity. AE is considered to be an attractive target for treating t(8;21) leukemia. However, AE expression alone is insufficient to cause transformation, and thus the potential of such therapy remains unclear. Several genes are deregulated in AML cells, including KIT that encodes a tyrosine kinase receptor. Here, we show that AML cells transduced with short hairpin RNA vector targeting AE mRNAs have a dramatic decrease in growth rate that is caused by induction of apoptosis and deregulation of the cell cycle. A reduction in KIT mRNA levels was also observed in AE-silenced cells, but silencing KIT expression reduced cell growth but did not induce apoptosis. Transcription profiling of cells that escape cell death revealed activation of a number of signaling pathways involved in cell survival and proliferation. In particular, we find that the extracellular signal-regulated kinase 2 (ERK2; also known as mitogen-activated protein kinase 1 (MAPK1)) protein could mediate activation of 23 out of 29 (79%) of these upregulated pathways and thus may be regarded as the key player in establishing the t(8;21)-positive leukemic cells resistant to AE suppression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chang KS, Fan YH, Stass SA, Estey EH, Wang G, Trujillo JM et al. Expression of AML1-ETO fusion transcripts and detection of minimal residual disease in t(8;21)-positive acute myeloid leukemia. Oncogene 1993; 8: 983–988.
Licht JD . AML1 and the AML1-ETO fusion protein in the pathogenesis of t(8;21) AML. Oncogene 2001; 20: 5660–5679.
Guerrasio A, Rosso C, Martinelli G, Lo Coco F, Pampinella M, Santoro A et al. Polyclonal haemopoieses associated with long-term persistence of the AML1-ETO transcript in patients with FAB M2 acute myeloid leukaemia in continous clinical remission. Br J Haematol 1995; 90: 364–368.
Wang J, Saunthararajah Y, Redner RL, Liu JM . Inhibitors of histone deacetylase relieve ETO-mediated repression and induce differentiation of AML1-ETO leukemia cells. Cancer Res 1999; 59: 2766–2769.
Hug BA, Lazar MA . ETO interacting proteins. Oncogene 2004; 23: 4270–4274.
Minucci S, Nervi C, Lo Coco F, Pelicci PG . Histone deacetylases: a common molecular target for differentiation treatment of acute myeloid leukemias? Oncogene 2001; 20: 3110–3115.
Wang J, Hoshino T, Redner RL, Kajigaya S, JM Liu . ETO, fusion partner in t(8;21) acute myeloid leukemia, represses transcription by interaction with the human N-CoR/mSin3/HDAC1 complex. Proc Natl Acad Sci USA 1998; 95: 10860–10865.
Zhen T, Wu CF, Liu P, Wu HY, Zhou GB, Lu Y et al. Targeting of AML1-ETO in t(8;21) leukemia by oridonin generates a tumor suppressor-like protein. Sci Transl Med 2012; 4: 127ra138.
Yuan Y, Zhou L, Miyamoto T, Iwasaki H, Harakawa N, Hetherington CJ et al. AML1-ETO expression is directly involved in the development of acute myeloid leukemia in the presence of additional mutations. Proc Natl Acad Sci USA 2001; 98: 10398–10403.
Muller AM, Duque J, Shizuru JA, Lubbert M . Complementing mutations in core binding factor leukemias: from mouse models to clinical applications. Oncogene 2008; 27: 5759–5773.
Schessl C, Rawat VP, Cusan M, Deshpande A, Kohl TM, Rosten PM et al. The AML1-ETO fusion gene and the FLT3 length mutation collaborate in inducing acute leukemia in mice. J Clin Invest 2005; 115: 2159–2168.
Kuchenbauer F, Feuring-Buske M, Buske C . AML1-ETO needs a partner: new insights into the pathogenesis of t(8;21) leukemia. Cell Cycle 2005; 4: 1716–1718.
de Guzman CG, Warren AJ, Zhang Z, Gartland L, Erickson P, Drabkin H et al. Hematopoietic stem cell expansion and distinct myeloid developmental abnormalities in a murine model of the AML1-ETO translocation. Mol Cell Biol 2002; 22: 5506–5517.
Schwieger M, Lohler J, Friel J, Scheller M, Horak I, Stocking C . AML1-ETO inhibits maturation of multiple lymphohematopoietic lineages and induces myeloblast transformation in synergy with ICSBP deficiency. J Exp Med 2002; 196: 1227–1240.
Zheng X, Oancea C, Henschler R, Ruthardt M . Cooperation between constitutively activated c-Kit signaling and leukemogenic transcription factors in the determination of the leukemic phenotype in murine hematopoietic stem cells. Int J Oncol 2009; 34: 1521–1531.
Cammenga J, Horn S, Bergholz U, Sommer G, Besmer P, Fiedler W et al. Extracellular KIT receptor mutants, commonly found in core binding factor AML, are constitutively active and respond to imatinib mesylate. Blood 2005; 106: 3958–3961.
Care RS, Valk PJ, Goodeve AC, Abu-Duhier FM, Geertsma-Kleinekoort WM, Wilson GA et al. Incidence and prognosis of c-KIT and FLT3 mutations in core binding factor (CBF) acute myeloid leukaemias. Br J Haematol 2003; 121: 775–777.
Beghini A, Larizza L, Cairoli R, Morra E . c-kit activating mutations and mast cell proliferation in human leukemia. Blood 1998; 92: 701–702.
Wang YY, Zhao LJ, Wu CF, Liu P, Shi L, Liang Y et al. C-KIT mutation cooperates with full-length AML1-ETO to induce acute myeloid leukemia in mice. Proc Natl Acad Sci USA 2011; 108: 2450–2455.
Ikeda H, Kanakura Y, Tamaki T, Kuriu A, Kitayama H, Ishikawa J et al. Expression and functional role of the proto-oncogene c-kit in acute myeloblastic leukemia cells. Blood 1991; 78: 2962–2968.
Cole SR, Aylett GW, Harvey NL, Cambareri AC, Ashman LK . Increased expression of c-Kit or its ligand Steel Factor is not a common feature of adult acute myeloid leukaemia. Leukemia 1996; 10: 288–296.
Reuss-Borst MA, Buhring HJ, Schmidt H, Muller CA . AML: immunophenotypic heterogeneity and prognostic significance of c-kit expression. Leukemia 1994; 8: 258–263.
Wang YY, Zhou GB, Yin T, Chen B, Shi JY, Liang WX et al. AML1-ETO and C-KIT mutation/overexpression in t(8;21) leukemia: implication in stepwise leukemogenesis and response to Gleevec. Proc Natl Acad Sci USA 2005; 102: 1104–1109.
Brioschi M, Fischer J, Cairoli R, Rossetti S, Pezzetti L, Nichelatti M et al. Down-regulation of microRNAs 222/221 in acute myelogenous leukemia with deranged core-binding factor subunits. Neoplasia 2010; 12: 866–876.
Krosl G, He G, Lefrancois M, Charron F, Romeo PH, Jolicoeur P et al. Transcription factor SCL is required for c-kit expression and c-Kit function in hemopoietic cells. J Exp Med 1998; 188: 439–450.
Lecuyer E, Herblot S, Saint-Denis M, Martin R, Begley CG, Porcher C et al. The SCL complex regulates c-kit expression in hematopoietic cells through functional interaction with Sp1. Blood 2002; 100: 2430–2440.
Munugalavadla V, Dore LC, Tan BL, Hong L, Vishnu M, Weiss MJ et al. Repression of c-kit and its downstream substrates by GATA-1 inhibits cell proliferation during erythroid maturation. Mol Cell Biol 2005; 25: 6747–6759.
Choi Y, Elagib KE, Delehanty LL, Goldfarb AN . Erythroid inhibition by the leukemic fusion AML1-ETO is associated with impaired acetylation of the major erythroid transcription factor GATA-1. Cancer Res 2006; 66: 2990–2996.
Hatlen MA, Wang L, Nimer SD . AML1-ETO driven acute leukemia: insights into pathogenesis and potential therapeutic approaches. Front Med 2012; 6: 248–262.
Spirin PV, Baskaran F, Orlova NN, Rulina AV, Nikitenko NA, Chernolovskaia EL et al. [Downregulation of activated leukemic oncogenes AML1-ETO and RUNX1(K83N) expression with RNA-interference]. Mol Biol (Mosk) 2010; 44: 876–888.
Spirin PV, Nikitenko NA, Lebedev TD, Rubtsov PM, Stocking C, Prasolov VS . [Modulation of activated oncogene c-kit expression with RNA-interference]. Mol Biol (Mosk) 2011; 45: 1036–1045.
Mitkevich VA, Petrushanko IY, Spirin PV, Fedorova TV, Kretova OV, Tchurikov NA et al. Sensitivity of acute myeloid leukemia Kasumi-1 cells to binase toxic action depends on the expression of KIT and AML1-ETO oncogenes. Cell Cycle 2011; 10: 4090–4097.
Larizza L, Magnani I, Beghini A . The Kasumi-1 cell line: a t(8;21)-kit mutant model for acute myeloid leukemia. Leuk Lymphoma 2005; 46: 247–255.
Heidenreich O, Krauter J, Riehle H, Hadwiger P, John M, Heil G et al. AML1/MTG8 oncogene suppression by small interfering RNAs supports myeloid differentiation of t(8;21)-positive leukemic cells. Blood 2003; 101: 3157–3163.
Okuda T, Cai Z, Yang S, Lenny N, Lyu CJ, van Deursen JM et al. Expression of a knocked-in AML1-ETO leukemia gene inhibits the establishment of normal definitive hematopoiesis and directly generates dysplastic hematopoietic progenitors. Blood 1998; 91: 3134–3143.
Bessard A, Frémin C, Ezan F, Fautrel A, Gailhouste L, Baffet G . RNAi-mediated ERK2 knockdown inhibits growth of tumor cells in vitro and in vivo. Oncogene 2008; 27: 5315–5325.
Lefloch R, Pouysségur J, Lenormand P . Total ERK1/2 activity regulates cell proliferation. Cell Cycle 2009; 8: 705–711.
Acknowledgements
This work was supported by the Programs of the Presidium of the Russian Academy of Sciences: ‘Molecular and Cell Biology’, ‘Fundamental Research Basics in Nanotechnology and Nanomaterials’, ‘Dynamics and Conservation of Genomes’ and the Russian Foundation for Basic Research (grant nos. 13-04-00599-a, 14-04-00821-a, 14-04-32108, 12-0433094 and 10-04-00593-a). We thank ‘UMA Foundation’ for their support in preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Spirin, P., Lebedev, T., Orlova, N. et al. Silencing AML1-ETO gene expression leads to simultaneous activation of both pro-apoptotic and proliferation signaling. Leukemia 28, 2222–2228 (2014). https://doi.org/10.1038/leu.2014.130
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.130
This article is cited by
-
MicroRNA let-7b downregulates AML1-ETO oncogene expression in t(8;21) AML by targeting its 3′UTR
Experimental Hematology & Oncology (2021)
-
Growth factor signaling predicts therapy resistance mechanisms and defines neuroblastoma subtypes
Oncogene (2021)
-
TAF1 plays a critical role in AML1-ETO driven leukemogenesis
Nature Communications (2019)