Abstract
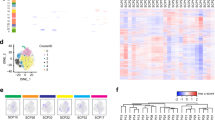
Genetic heterogeneity is common in tumors, explicable by the development of subclones with distinct genetic and epigenetic alterations. We describe an in vitro model for cancer heterogeneity, comprising the diffuse large B-cell lymphoma cell line U-2932 which expresses two sets of cell surface markers representing twin populations flow-sorted by CD20 vs CD38 expression. U-2932 populations were traced to subclones of the original tumor with clone-specific immunoglobulin IgVH4–39 hypermutation patterns. BCL6 was overexpressed in one subpopulation (R1), MYC in the other (R2), both clones overexpressed BCL2. According to the combined results of immunoglobulin hypermutation and cytogenetic analysis, R1 and R2 derive from a mother clone with genomic BCL2 amplification, which acquired secondary rearrangements leading to the overexpression of BCL6 (R1) or MYC (R2). Some 200 genes were differentially expressed in R1/R2 microarrays including transcriptional targets of the aberrantly expressed oncogenes. Other genes were regulated by epigenetic means as shown by DNA methylation analysis. Ectopic expression of BCL6 in R2 variously modulated new candidate target genes, confirming dual silencing and activating functions. In summary, stable retention of genetically distinct subclones in U-2932 models tumor heterogeneity in vitro permitting functional analysis of oncogenes against a syngenic background.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Greaves M, Maley CC . Clonal evolution in cancer. Nature 2012; 481: 306–313.
Notta F, Mulligham CG, Wang JCY, Peoppl A, Doulatov S, Phillips LA et al. Evolution of human BCR-ABL1 lymphoblastic leukaemia—initiating cells. Nature 2011; 469: 362–367.
Anderson K, Lutz C, vanDelft FW, Bateman CM, Guo Y, Colman SM et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 2011; 469: 356–361.
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 2012; 481: 506–509.
Klein CA . Parallel progression of primary tumours and metastases. Nature Rev Cancer 2009; 9: 302–312.
Navin N, Kendall J, Troge J, Andrews P, Rodgers L, McIndoo J et al. Tumour evolution inferred by single-cell sequencing. Nature 2011; 472: 90–94.
Klein U, Dalla-Favera R . Germinal centres: role in B-cell physiology and malignancy. Nature Rev Immunol 2008; 8: 22–33.
Drexler HG . Guide to leukemia-lymphoma cell lines. 2nd Edn. Braunschweig, 2010.
MacLeod RA, Kaufmann M, Drexler HG . Cytogenetic harvesting of commonly used tumour cell lines. Nat Protoc 2007; 2: 372–382.
MacLeod RA, Kaufmann M, Drexler HG . Cytogenetic analysis of cancer cell lines. Methods Mol Biol 2011; 731: 57–78.
Quentmeier H, Schneider B, Röhrs S, Romani J, Zaborski M, MacLeod RAF et al. SET-NUP214 fusion in acute myeloid leukemia- and T-cell acute lymphoblastic leukemia-derived cell lines. J Hematol Oncol 2009; 2: 3.
Rosenquist LAH, Forestier E, Holmberg D, Lindh J, Löfvenberg E, Roos G . Clonal rearragements in childhood and adult precursor B acute lymphoblastic leukemia: a comparative polymerase chain reaction study using multiple sets of primers. Eur J Haematol 1999; 63: 211–218.
Peng HZ, Du MQ, Koulis A, Aiello A, Dogan A, Pan LX et al. Nonimmunoglobulin gene hypermutation in germinal center B cells. Blood 1999; 93: 2167–2172.
Amini RM, Berglund M, Rosenquist R, von Heideman A, Lagercrantz S, Thunberg U et al. A novel B-cell line (U-2932) established from a patient with diffuse large B-cell lymphoma following Hodgkin lymphoma. Leukemia Lymphoma 2002; 43: 2179–2189.
Basso K, Dalla-Favera R . BCL6: master regulator of the germinal center reaction and key oncogene in B cell lymphomagenesis. Adv Immunol 2010; 105: 193–210.
Barrans SL, Evans PAS, O'Connor SJM, Kendall SJ, Owen RG, Haynes AP et al. The t(14;18) is associated with germinal center-derived diffuse large B-cell lymphoma and is a strong predictor of outcome. Clin Cancer Res 2003; 9: 2133–2139.
Kobayashi T, Tsutsumi Y, Sakamoto N, Nagoshi H, Yamamoto-Sugitani M, Shimura Y et al. Double-hit lymphomas constitute a highly aggressive subgroup in diffuse large B-cell lymphomas in the era of rituximab. Jpn J Clin Oncol 2012; 42: 1035–1042.
Victora GD, Dominguez-Sola D, Holmes AB, Deroubaix S, Dalla-Favera R, Nussenzweig MC . Identification of human germinal center light and dark zone cells and their relationship to human B-cell lymphomas. Blood 2012; 120: 2240–2248.
Shaffer AL, Yu X, He Y, Boldrick J, Chan EP, Staudt LM . BCL-6 represses genes that function in lymphocyte differentiation, inflammation and cell cycle control. Immunity 2000; 13: 199–212.
Phan RT, Dalla-Favera R . The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. Nature 2004; 432: 635–639.
Phan RT, Saito M, Basso K, Niu H, Dalla-Favera R . BCL6 interacts with the transcription factor Miz-1 to suppress the cyclin-dependent kinase inhibitor p21 and cell cycle arrest in germinal center B cells. Nature Immunol 2005; 6: 1054–1060.
Ci W, Polo JM, Cerchietti L, Shaknovich R, Wang L, Yang SN et al. The BCL6 transcriptional program features repression of multiple oncogenes in primary B cells and is deregulated in DLBCL. Blood 2009; 113: 5536–5547.
Warburg O . On the origin of cancer cells. Science 1956; 123: 309–314.
Siegmund KD, Marjoram P, Woo YJ, Tavare S, Shibata D . Inferring clonal expansion and cancer stem cell dynamics from DNA methylation patterns in colorectal cancers. Proc Natl Acad Sci USA 2009; 106: 4828–4833.
Jung S, Yi L, Kim J, Jeong D, Oh T, Kim CH et al. The role of vimentin as a methylation biomarker for early diagnosis of cervical cancer. Mol Cells 2011; 31: 405–411.
Carey JPW, Asirvatham AJ, Galm O, Ghogomu TA, Chaudhary J . Inhibitor of differentiation 4 (Id4) is a potential tumor suppressor in prostate cancer. BMC Cancer 2009; 9: 173.
Snuderl M, Fazlollahi L, Le LP, Nitta M, Zhelyazkova BH, Davicson CJ et al. Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 2011; 20: 810–817.
Acknowledgements
We thank Mrs Karin Battmer (MHH, Hannover, Germany) for technical help in retroviral transduction experiments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Quentmeier, H., Amini, R., Berglund, M. et al. U-2932: two clones in one cell line, a tool for the study of clonal evolution. Leukemia 27, 1155–1164 (2013). https://doi.org/10.1038/leu.2012.358
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2012.358