Abstract
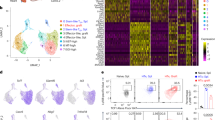
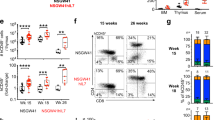
Chronic lymphocytic leukemia (CLL) cells require complex microenvironmental and immunologic interactions to survive and proliferate. Such interactions might be best recreated in animal models; however, this needs extensive verification. We therefore investigated the composition of the T-cell compartment in the Eμ-TCL1 transgenic mouse, currently the most widely used murine model for CLL. Immunophenotyping and transplant approaches were used to define T-cell subsets at various stages of CLL. Analogous to human CLL, we observed a skewing of T-cell subsets from naive to antigen-experienced memory T cells that was more pronounced in lymph nodes than in blood. Transplantation of CLL into non-transgenic recipients was feasible without immunosuppression in a pure C57BL/6 background and resulted in the prominent skewing of the T cells of the recipient mice. Both in spontaneously developed CLL and in the transplantation setting, a loss in T-cell receptor diversity was observed, with a relevant number of clonal T-cell populations arising. This suggests that antigen-dependent differentiation toward the T memory pool is initiated by murine CLL cells. In summary, we validate the TCL1 transgenic mouse model for analysis of T-cell phenotypes and suggest a CLL-dependent antigen-driven skewing of T cells in these mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Caligaris-Cappio F, Ghia P . Novel insights in chronic lymphocytic leukemia: are we getting closer to understanding the pathogenesis of the disease? J Clin Oncol 2008; 26: 4497–4503.
Ghia P, Chiorazzi N, Stamatopoulos K . Microenvironmental influences in chronic lymphocytic leukaemia: the role of antigen stimulation. J Intern Med 2008; 264: 549–562.
Chiorazzi N, Ferrarini M . Evolving view of the in-vivo kinetics of chronic lymphocytic leukemia B cells. Hematol Am Soc Hematol Educ Program 2006; 273–278: 512.
Chiorazzi N . Cell proliferation and death: forgotten features of chronic lymphocytic leukemia B cells. Best Pract Res Clin Haematol 2007; 20: 399–413.
Messmer BT, Messmer D, Allen SL, Kolitz JE, Kudalkar P, Cesar D et al. In vivo measurements document the dynamic cellular kinetics of chronic lymphocytic leukemia B cells. J Clin Invest 2005; 115: 755–764.
Patten PE, Buggins AG, Richards J, Wotherspoon A, Salisbury J, Mufti GJ et al. CD38 expression in chronic lymphocytic leukemia is regulated by the tumor microenvironment. Blood 2008; 111: 5173–5181.
Mellstedt H, Choudhury A . T and B cells in B-chronic lymphocytic leukaemia: Faust, Mephistopheles and the pact with the Devil. Cancer Immunol Immunother 2006; 55: 210–220.
Tinhofer I, Marschitz I, Kos M, Henn T, Egle A, Villunger A et al. Differential sensitivity of CD4+ and CD8+ T lymphocytes to the killing efficacy of Fas (Apo-1/CD95) ligand+ tumor cells in B chronic lymphocytic leukemia. Blood 1998; 91: 4273–4281.
Ramsay AG, Johnson AJ, Lee AM, Gorgun G, Le Dieu R, Blum W et al. Chronic lymphocytic leukemia T cells show impaired immunological synapse formation that can be reversed with an immunomodulating drug. J Clin Invest 2008; 118: 2427–2437.
Tinhofer I, Weiss L, Gassner F, Rubenzer G, Holler C, Greil R . Difference in the relative distribution of CD4+ T-cell subsets in B-CLL with mutated and unmutated immunoglobulin (Ig) VH genes: implication for the course of disease. J Immunother 2009; 32: 302–309.
Durig J, Ebeling P, Grabellus F, Sorg UR, Mollmann M, Schutt P et al. A novel nonobese diabetic/severe combined immunodeficient xenograft model for chronic lymphocytic leukemia reflects important clinical characteristics of the disease. Cancer Res 2007; 67: 8653–8661.
Bichi R, Shinton SA, Martin ES, Koval A, Calin GA, Cesari R et al. Human chronic lymphocytic leukemia modeled in mouse by targeted TCL1 expression. Proc Natl Acad Sci USA 2002; 99: 6955–6960.
Noguchi M, Ropars V, Roumestand C, Suizu F . Proto-oncogene TCL1: more than just a coactivator for Akt. FASEB J 2007; 21: 2273–2284.
Teitell MA . The TCL1 family of oncoproteins: co-activators of transformation. Nat Rev Cancer 2005; 5: 640–648.
Herling M, Patel KA, Weit N, Lilienthal N, Hallek M, Keating MJ et al. High TCL1 levels are a marker of B-cell receptor pathway responsiveness and adverse outcome in chronic lymphocytic leukemia. Blood 2009; 114: 4675–4686.
Schlette E, Tam CS, Herling M, Buenso-Ramos C, Wierda W, Ferrajoli A et al. High T-cell lymphoma breakpoint 1 (TCL1) levels are associated with lack of response and inferior outcome following fludarabine, cyclophosphamide, rituximab (FCR) frontline therapy in patients with chronic lymphocytic leukemia (CLL). Blood (ASH Annual Meeting Abstracts). Ref Type: Abstract 2010; 112: 2086.
Yan XJ, Albesiano E, Zanesi N, Yancopoulos S, Sawyer A, Romano E et al. B cell receptors in TCL1 transgenic mice resemble those of aggressive, treatment-resistant human chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2006; 103: 11713–11718.
Chen SS, Sherman MH, Hertlein E, Johnson AJ, Teitell MA, Byrd JC et al. Epigenetic alterations in a murine model for chronic lymphocytic leukemia. Cell Cycle 2009; 8: 3663–3667.
Chen SS, Raval A, Johnson AJ, Hertlein E, Liu TH, Jin VX et al. Epigenetic changes during disease progression in a murine model of human chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2009; 106: 13433–13438.
Enzler T, Kater AP, Zhang W, Widhopf GF, Chuang HY, Lee J et al. Chronic lymphocytic leukemia of Emu-TCL1 transgenic mice undergoes rapid cell turnover that can be offset by extrinsic CD257 to accelerate disease progression. Blood 2009; 114: 4469–4476.
Holler C, Pinon JD, Denk U, Heyder C, Hofbauer S, Greil R et al. PKCbeta is essential for the development of chronic lymphocytic leukemia in the TCL1 transgenic mouse model: validation of PKCbeta as a therapeutic target in chronic lymphocytic leukemia. Blood 2009; 113: 2791–2794.
Sanchez-Aguilera A, Rattmann I, Drew DZ, Muller LU, Summey V, Lucas DM et al. Involvement of RhoH GTPase in the development of B-cell chronic lymphocytic leukemia. Leukemia 2010; 24: 97–104.
Gorgun G, Ramsay AG, Holderried TA, Zahrieh D, Le Dieu R, Liu F et al. E(mu)-TCL1 mice represent a model for immunotherapeutic reversal of chronic lymphocytic leukemia-induced T-cell dysfunction. Proc Natl Acad Sci USA 2009; 106: 6250–6255.
Pannetier C, Cochet M, Darche S, Casrouge A, Zoller M, Kourilsky P . The sizes of the CDR3 hypervariable regions of the murine T-cell receptor beta chains vary as a function of the recombined germ-line segments. Proc Natl Acad Sci USA 1993; 90: 4319–4323.
Garcia C, Rosen A, Kimby E, Aguilar-Santelises M, Jondal M, Bjorkhilm M et al. Higher T-cell imbalance and growth factor receptor expression in B-cell chronic lymphocytic leukemia (B-CLL) as compared to monoclonal B-cell lymphocytosis of undetermined significance (B-MLUS). Leuk Res 1989; 13: 31–37.
Kimby E, Mellstedt H, Nilsson B, Bjorkholm M, Holm G . T lymphocyte subpopulations in chronic lymphocytic leukemia of B cell type in relation to immunoglobulin isotype(s) on the leukemic clone and to clinical features. Eur J Haematol 1987; 38: 261–267.
Bingaman AW, Patke DS, Mane VR, Ahmadzadeh M, Ndejembi M, Bartlett ST et al. Novel phenotypes and migratory properties distinguish memory CD4 T cell subsets in lymphoid and lung tissue. Eur J Immunol 2005; 35: 3173–3186.
Dutton RW, Bradley LM, Swain SL . T cell memory. Annu Rev Immunol 1998; 16: 201–223.
Farber DL . T cell memory: heterogeneity and mechanisms. Clin Immunol 2000; 95: 173–181.
Rezvany MR, Jeddi-Tehrani M, Osterborg A, Kimby E, Wigzell H, Mellstedt H . Oligoclonal TCRBV gene usage in B-cell chronic lymphocytic leukemia: major perturbations are preferentially seen within the CD4 T-cell subset. Blood 1999; 94: 1063–1069.
Rezvany MR, Jeddi-Tehrani M, Wigzell H, Osterborg A, Mellstedt H . Leukemia-associated monoclonal and oligoclonal TCR-BV use in patients with B-cell chronic lymphocytic leukemia. Blood 2003; 101: 1063–1070.
Acknowledgements
This work was supported by Austrian FWF Grants P19481 and L488 to AE, Austrian National Bank Grant 10990 to AE and by the state of Salzburg and the SFB 021 to RG. TCR CDR3 spectratyping was performed at the Children's Hospital of the Federal Hospital of Salzburg.
Author contributions AE, JPH, IT and RG designed the research; JPH, ChH, UD, TK, ClH, DT and DA performed the experimental work; AE and JPH performed the analysis; AE and JPH wrote the paper. All authors were involved in critical discussion of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Hofbauer, J., Heyder, C., Denk, U. et al. Development of CLL in the TCL1 transgenic mouse model is associated with severe skewing of the T-cell compartment homologous to human CLL. Leukemia 25, 1452–1458 (2011). https://doi.org/10.1038/leu.2011.111
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2011.111
Keywords
This article is cited by
-
Comprehensive characterization of clonality of driver genes revealing their clinical relevance in colorectal cancer
Journal of Translational Medicine (2022)
-
IL4I1-driven AHR signature: a new avenue for cancer therapy
Signal Transduction and Targeted Therapy (2021)
-
Bone marrow dendritic cells support the survival of chronic lymphocytic leukemia cells in a CD84 dependent manner
Oncogene (2020)
-
TCL1 transgenic mice as a model for CD49d-high chronic lymphocytic leukemia
Leukemia (2020)
-
T-cells in chronic lymphocytic leukemia: Guardians or drivers of disease?
Leukemia (2020)