Abstract
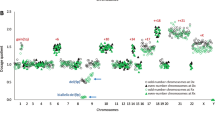
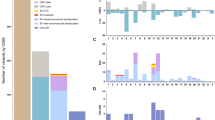
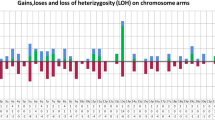
We have undertaken a genome-wide single nucleotide polymorphism (SNP) array analysis of 41 chronic myeloid leukemia (CML) patients. In total, 44 regions of uniparental disomy (UPD) >3 Mb were identified in 24 of 32 patients in chronic phase (CP), and 21 regions of UPD >3 Mb were identified in 13 of 21 patients in blast crisis (BC). Chromosome 8 had the highest frequency of UPD regions in both CP and BC samples. Eight recurrent regions of UPD were observed among the 41 patients, with chromosome 8 showing the highest frequency. Ten regions of copy number change (CNC) >3 Mb were observed in 4 of 21 patients in BC, whereas none were observed in CP. We have identified several recurrent regions of UPD and CNC in CML that may be of pathogenetic importance. Overrepresentation of genomic aberrations (UPD and copy number gain) mapping to chromosome 8 was observed. Selected candidate genes mapping within the aberrant genomic regions were sequenced and mutation of the TP53 gene was observed in one case in BC and of the ASXL1 gene in 6 of 41 cases in CP or BC. Mutation of ASXL1 represents an important new molecular abnormality in CML.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Hughes T, Goldman JM . Hematology, Basic Principles and Practice, 2nd edn. Churchill Livingstone: New York, 1995; 854–869.
Rowley JD . Letter: A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 1973; 243: 290–293.
Shet AS, Jahagirdar BN, Verfaillie CM . Chronic myelogenous leukemia: mechanisms underlying disease progression. Leukemia 2002; 16: 1402–1411.
Hughes TP, Kaeda J, Branford S, Rudzki Z, Hochhaus A, Hensley ML et al. Frequency of major molecular responses to imatinib or interferon alfa plus cytarabine in newly diagnosed chronic myeloid leukemia. N Engl J Med 2003; 349: 1423–1432.
Hochhaus A, O’Brien SG, Guilhot F, Druker BJ, Branford S, Foroni L et al. Six-year follow-up of patients receiving imatinib for the first-line treatment of chronic myeloid leukemia. Leukemia 2009; 23: 1054–1061.
Druker BJ, Sawyers CL, Kantarjian H, Resta DJ, Reese SF, Ford JM et al. Activity of a specific inhibitor of the BCR-ABL tyrosine kinase in the blast crisis of chronic myeloid leukemia and acute lymphoblastic leukemia with the Philadelphia chromosome. N Engl J Med 2001; 344: 1038–1042.
Volpe G, Panuzzo C, Ulisciani S, Cilloni D . Imatinib resistance in CML. Cancer Lett 2009; 274: 1–9.
Johansson B, Fioretos T, Mitelman F . Cytogenetic and molecular genetic evolution of chronic myeloid leukemia. Acta Haematol 2002; 107: 76–94.
Boultwood J, Peniket A, Watkins F, Shepherd P, McGale P, Richards S et al. Telomere length shortening in chronic myelogenous leukemia is associated with reduced time to accelerated phase. Blood 2000; 96: 358–361.
Campbell LJ, Fidler C, Eagleton H, Peniket A, Kusec R, Gal S et al. hTERT, the catalytic component of telomerase, is downregulated in the haematopoietic stem cells of patients with chronic myeloid leukaemia. Leukemia 2006; 20: 671–679.
Ahuja H, Bar-Eli M, Advani SH, Benchimol S, Cline MJ . Alterations in the p53 gene and the clonal evolution of the blast crisis of chronic myelocytic leukemia. Proc Natl Acad Sci USA 1989; 86: 6783–6787.
Calabretta B, Perrotti D . The biology of CML blast crisis. Blood 2004; 103: 4010–4022.
Mullighan CG, Miller CB, Radtke I, Phillips LA, Dalton J, Ma J et al. BCR-ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature 2008; 453: 110–114.
Wang L, Pearson K, Pillitteri L, Ferguson JE, Clark RE . Serial monitoring of BCR-ABL by peripheral blood real-time polymerase chain reaction predicts the marrow cytogenetic response to imatinib mesylate in chronic myeloid leukaemia. Br J Haematol 2002; 118: 771–777.
Hughes T, Deininger M, Hochhaus A, Branford S, Radich J, Kaeda J et al. Monitoring CML patients responding to treatment with tyrosine kinase inhibitors: review and recommendations for harmonizing current methodology for detecting BCR-ABL transcripts and kinase domain mutations and for expressing results. Blood 2006; 108: 28–37.
Wang L, Fidler C, Nadig N, Giagounidis A, Della Porta MG, Malcovati L et al. Genome-wide analysis of copy number changes and loss of heterozygosity in myelodysplastic syndrome with del(5q) using high-density single nucleotide polymorphism arrays. Haematologica 2008; 93: 994–1000.
Fidler C, Watkins F, Bowen DT, Littlewood TJ, Wainscoat JS, Boultwood J . NRAS, FLT3 and TP53 mutations in patients with myelodysplastic syndrome and a del(5q). Haematologica 2004; 89: 865–866.
Carbuccia N, Murati A, Trouplin V, Brecqueville M, Adelaide J, Rey J et al. Mutations of ASXL1 gene in myeloproliferative neoplasms. Leukemia 2009; 23: 2183–2186.
Gelsi-Boyer V, Trouplin V, Adelaide J, Bonansea J, Cervera N, Carbuccia N et al. Mutations of polycomb-associated gene ASXL1 in myelodysplastic syndromes and chronic myelomonocytic leukaemia. Br J Haematol 2009; 145: 788–800.
Mohamedali A, Gaken J, Twine NA, Ingram W, Westwood N, Lea NC et al. Prevalence and prognostic significance of allelic imbalance by single-nucleotide polymorphism analysis in low-risk myelodysplastic syndromes. Blood 2007; 110: 3365–3373.
Gibson J, Morton NE, Collins A . Extended tracts of homozygosity in outbred human populations. Hum Mol Genet 2006; 15: 789–795.
Petitjean A, Mathe E, Kato S, Ishioka C, Tavtigian SV, Hainaut P et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat 2007; 28: 622–629.
Fitzgibbon J, Smith LL, Raghavan M, Smith ML, Debernardi S, Skoulakis S et al. Association between acquired uniparental disomy and homozygous gene mutation in acute myeloid leukemias. Cancer Res 2005; 65: 9152–9154.
Raghavan M, Lillington DM, Skoulakis S, Debernardi S, Chaplin T, Foot NJ et al. Genome-wide single nucleotide polymorphism analysis reveals frequent partial uniparental disomy due to somatic recombination in acute myeloid leukemias. Cancer Res 2005; 65: 375–378.
Tagawa H, Karnan S, Kasugai Y, Tuzuki S, Suzuki R, Hosokawa Y et al. MASL1, a candidate oncogene found in amplification at 8p23.1, is translocated in immunoblastic B-cell lymphoma cell line OCI-LY8. Oncogene 2004; 23: 2576–2581.
Sakabe T, Shinomiya T, Mori T, Ariyama Y, Fukuda Y, Fujiwara T et al. Identification of a novel gene, MASL1, within an amplicon at 8p23.1 detected in malignant fibrous histiocytomas by comparative genomic hybridization. Cancer Res 1999; 59: 511–515.
Guo L, Zhong D, Lau S, Liu X, Dong XY, Sun X et al. Sox7 is an independent checkpoint for beta-catenin function in prostate and colon epithelial cells. Mol Cancer Res 2008; 6: 1421–1430.
Zhang Y, Huang S, Dong W, Li L, Feng Y, Pan L et al. SOX7, down-regulated in colorectal cancer, induces apoptosis and inhibits proliferation of colorectal cancer cells. Cancer Lett 2009; 277: 29–37.
Gribble SM, Reid AG, Roberts I, Grace C, Green AR, Nacheva EP . Genomic imbalances in CML blast crisis: 8q24.12-q24.13 segment identified as a common region of over-representation. Genes Chromosomes Cancer 2003; 37: 346–358.
Hosoya N, Sanada M, Nannya Y, Nakazaki K, Wang L, Hangaishi A et al. Genomewide screening of DNA copy number changes in chronic myelogenous leukemia with the use of high-resolution array-based comparative genomic hybridization. Genes Chromosomes Cancer 2006; 45: 482–494.
Gonzalez MA, Tachibana KE, Laskey RA, Coleman N . Control of DNA replication and its potential clinical exploitation. Nat Rev Cancer 2005; 5: 135–141.
Geay JF, Buet D, Zhang Y, Foudi A, Jarrier P, Berthebaud M et al. p210BCR-ABL inhibits SDF-1 chemotactic response via alteration of CXCR4 signaling and down-regulation of CXCR4 expression. Cancer Res 2005; 65: 2676–2683.
Jin L, Tabe Y, Konoplev S, Xu Y, Leysath CE, Lu H et al. CXCR4 up-regulation by imatinib induces chronic myelogenous leukemia (CML) cell migration to bone marrow stroma and promotes survival of quiescent CML cells. Mol Cancer Ther 2008; 7: 48–58.
Kagara N, Tanaka N, Noguchi S, Hirano T . Zinc and its transporter ZIP10 are involved in invasive behavior of breast cancer cells. Cancer Sci 2007; 98: 692–697.
Curtiss NP, Bonifas JM, Lauchle JO, Balkman JD, Kratz CP, Emerling BM et al. Isolation and analysis of candidate myeloid tumor suppressor genes from a commonly deleted segment of 7q22. Genomics 2005; 85: 600–607.
Bras M, Yuste VJ, Roue G, Barbier S, Sancho P, Virely C et al. Drp1 mediates caspase-independent type III cell death in normal and leukemic cells. Mol Cell Biol 2007; 27: 7073–7088.
Papagerakis S, Shabana AH, Depondt J, Gehanno P, Forest N . Immunohistochemical localization of plakophilins (PKP1, PKP2, PKP3, and p0071) in primary oropharyngeal tumors: correlation with clinical parameters. Hum Pathol 2003; 34: 565–572.
Mertens C, Kuhn C, Moll R, Schwetlick I, Franke WW . Desmosomal plakophilin 2 as a differentiation marker in normal and malignant tissues. Differentiation 1999; 64: 277–290.
Fisher CL, Randazzo F, Humphries RK, Brock HW . Characterization of Asxl1, a murine homolog of Additional sex combs, and analysis of the Asx-like gene family. Gene 2006; 369: 109–118.
An Q, Wright SL, Konn ZJ, Matheson E, Minto L, Moorman AV et al. Variable breakpoints target PAX5 in patients with dicentric chromosomes: a model for the basis of unbalanced translocations in cancer. Proc Natl Acad Sci USA 2008; 105: 17050–17054.
Foekens JA, Kos J, Peters HA, Krasovec M, Look MP, Cimerman N et al. Prognostic significance of cathepsins B and L in primary human breast cancer. J Clin Oncol 1998; 16: 1013–1021.
Hughes SJ, Glover TW, Zhu XX, Kuick R, Thoraval D, Orringer MB et al. A novel amplicon at 8p22–23 results in overexpression of cathepsin B in esophageal adenocarcinoma. Proc Natl Acad Sci USA 1998; 95: 12410–12415.
Acknowledgements
This work was supported by the Kay Kendall Leukaemia Fund and by Leukaemia Research UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Boultwood, J., Perry, J., Zaman, R. et al. High-density single nucleotide polymorphism array analysis and ASXL1 gene mutation screening in chronic myeloid leukemia during disease progression. Leukemia 24, 1139–1145 (2010). https://doi.org/10.1038/leu.2010.65
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2010.65
Keywords
This article is cited by
-
Low prevalence of the BCR–ABL1 fusion gene in a normal population in southern Sarawak
International Journal of Hematology (2020)
-
RUNX1 mutations promote leukemogenesis of myeloid malignancies in ASXL1-mutated leukemia
Journal of Hematology & Oncology (2019)
-
PRR14L mutations are associated with chromosome 22 acquired uniparental disomy, age-related clonal hematopoiesis and myeloid neoplasia
Leukemia (2019)
-
ASXL1 alteration cooperates with JAK2V617F to accelerate myelofibrosis
Leukemia (2019)
-
Frequent ASXL1 mutations in children and young adults with chronic myeloid leukemia
Leukemia (2018)