Abstract
Adams–Oliver syndrome (AOS, OMIM; 100300) is a rare genetic disease characterized by aplasia cutis congenita, terminal transverse limb defects and cutis marmorata with vascular anomalies such as congenital heart defects. The etiology of this syndrome has remained largely unknown but defective Notch signaling during vascular formation has been suggested. Here we describe a sporadic Japanese newborn case with clinically diagnosed AOS. Trio whole-exome sequencing identified a de novo, novel, heterozygous missense mutation in the Delta-like 4 ligand gene (DLL4 c.572G>A, p.Arg191His) in the patient. DLL4 functions as a requisite ligand for NOTCH1 receptor, which is essential for vascular formation. Amino acid substitution of Arg191 to His was predicted by molecular models to interfere with direct binding between DLL4 and NOTCH1. DLL4 has recently been identified as a causative gene of an autosomal dominant type of AOS with milder symptoms. The case described here showed gradual recovery from skull defects after birth and no psychomotor developmental delay has been observed. This is the second report of an AOS case with DLL4 mutation, and the phenotypic characteristics between the two cases are compared and discussed.
Similar content being viewed by others
Introduction
Adams–Oliver syndrome (AOS, OMIM; 100300) was first reported by Adams and Oliver in 1945.1 AOS is an extremely rare disease, with an estimated incidence of 1/225 000. Various clinical manifestations have been reported, including aplasia cutis congenita of the scalp, terminal transverse limb defects, and cutis marmorata with vascular anomalies such as congenital heart defects.2 The most common defects are terminal limb malformations (84% of cases), including osseous syndactyly, rudimentary bones, or completely absent digits. Congenital cutis aplasia, the second-most-common defect (75% of cases), usually occurs over the posterior parietal region. Underlying bone defects can be present, and tortuous veins can occur on the posterior scalp.3, 4
Six causative genes for AOS have been identified, namely ARHGAP31 (AOS type 1),5 RBPJ (AOS type 3),6 NOTCH1 (AOS type 5),7, 8 DOCK6 (AOS type 2),9 EOGT (AOS type 4)10 and DLL4.11 Other genes are still to be identified because many patients with AOS do not exhibit mutations in any of the known causative genes. RBPJ, NOTCH1 and EOGT are involved in the Notch signaling pathway, which has an important role in stabilization of arterial endothelial fate and angiogenesis.12 Consistent with this, malformation of vasculature and defects in the remodeling of vessels in the early fetal phase are considered common features of this syndrome. Of the six known causative genes, mutations in ARHGAP31, RBPJ, NOTCH1 and DLL4 are inherited in an autosomal dominant manner, and those in DOCK6 and EOGT are inherited in an autosomal recessive manner. Patients carrying DOCK6 mutations show the most severe phenotype. Mutations in DLL4 have been recently identified in several families,11 with these patients showing relatively mild phenotypes compared with autosomal recessive cases. Here we report the first Japanese case of AOS with a novel missense variant in DLL4. All procedures were reviewed and approved by the Institutional Review Board of Kobe University School of Medicine and were performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki. We performed the analysis after obtaining written informed consent from the parents of the patients.
Clinical report
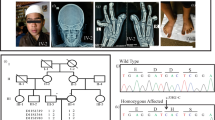
A Japanese family, including the proband, was investigated in this study (Figure 1a). The proband, the first child of non-consanguineous, healthy parents, was born at 34 weeks, 4 days gestation with a birth weight of 1334 g (−2.7 s.d.), height of 39.6 cm (−2.2 s.d.), and occipitofrontal circumstance of 26 cm (−2.5 s.d.). During pregnancy, intrauterine growth restriction and single umbilical cord artery were present. The patient was transferred to the newborn intensive care unit (NICU) soon after birth, presenting with bleeding around aplasia cutis congenita of the skull (Figure 1b). Limb abnormalities, such as bilateral syndactyly of the lower extremities and brachydactyly and cutis marmorata were also observed (Figure 1b). Clinical presentation suggested a diagnosis of typical AOS. No congenital heart defect was detected by echocardiogram. Cranial computed tomography indicated a wide range of bone defects around the anterior fontanelle (Figure 1c, upper), but brain MRI was normal (not shown).
Family pedigree and the clinical features of the case. (a) Family pedigree. Squares, males; circles, females; open shape, unaffected; filled shape, affected; P, proband. (b) Patient’s clinical features. Left, Aplasia cutis congenita; middle upper, symbrachydactyly of right toe; right upper, X-ray of the right toe; middle lower, hypoplastic nails; right lower, X-ray of the left toe. (c) Three dimensional representation of the patient’s scalp by computed tomography. Left, front view; right, upper views. Wide area of bone defect around frontal fontanelle (day 7 after birth) and gradual ossification were seen at 10 and 18 months after birth. (d) Aplasia cutis congenita. Left, day 0; middle, day 31; right, 10 months after birth. Skin bleeding diminished gradually and crust formation was complete around 10 months after birth.
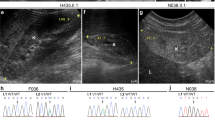
In the first 10 months after birth, bleeding symptoms around aplasia cutis congenita of the skull gradually improved with treatment, including petroleum jelly application and gauze protection (Figure 1d). Genetic analysis was conducted for further diagnosis. Firstly, chromosomal analysis showed a normal karyotype. Next, genomic DNA was obtained from peripheral blood leukocytes. Comparative genomic hybridization and single-nucleotide polymorphism array analysis (180 K, SurePrintG3, Agilent Technologies, Santa Clara, CA, USA) were performed on DNA from the proband using an Agilent DNA Microarray Scanner BA (Agilent Technologies). This analysis discovered no pathogenic copy-number variations. For next-generation sequencing (NGS) analysis of the patient, we first used an Illumina TruSight One sequencing panel on a MiSeq platform according to the manufacturer’s instructions (Illumina, San Diego, CA, USA). However, no pathogenic sequence variations were identified. Therefore, we conducted whole-exome sequencing on the proband and his parents using Illumina Nextera Rapid Capture Exome on a Hiseq platform. For data analysis, we used VariantStudio (ver. 2.1.46; Illumina) and Integrative Genomics Viewer software (ver. 2.3.32; Broad Institute, Cambridge, MA, USA). We identified a de novo, novel, heterozygous missense mutation in the Delta-like 4 ligand gene (DLL4 c.572G>A, p.Arg191His) on chromosome 15q15.1 in the patient (Figure 2a). This variant was within a conserved region and had not previously been reported or registered in the human genome Human Genome Variation browser. The in silico mutation analysis tools, SIFT, PolyPhen-2 and Mutation Taster (http://mutationtaster.org) produced scores of 0 (deleterious), 1.0 (damaging) and ‘disease causing’, respectively. This variant was confirmed by direct sequencing and it was determined that the missense mutation was not transmitted from the parents (Figure 2b). At 18 months of age, no intellectual disability or psychomotor developmental delay was displayed. Brain CT showed a wide range of skull defects around the anterior fontanelle, but the extent of ossification gradually increased, as shown by brain CT on day 0 and at 10 months (Figure 1c). The patient still required a custom-made head protector and had difficulty using eating utensils because of his malformed digits.
Genetic analysis of the proband. (a) Integrative Genomics Viewer picture of the father (upper lane), mother (second lane), patient (third lane), and the patient’s umbilical cord (bottom lane). The pathogenic heterozygous DLL4 variant (c.572G>A, p.Arg191His) was detected only in the proband’s genome. (b) Sanger sequencing of the pathogenic DLL4 variant. (c) The domain containing identified variants within DLL4. Orange rectangle shows DSL domain. (d) Structure of human notch ligand delta-like 1 (DLL1) and delta-like 4 (DLL4) showing conserved arginine. Full length sequence of DLL4 (Homo sapiens), which consists of 665 amino acid residues, was taken from the NCBI data bank. DLL4 shows good pairwise sequence similarity (using BLAST) with the crystal structure of DLL1 (Protein Database (PDB) ID 4XBM).18 Notch receptor (PDB ID 4CC0).19 These two PDB structures were chosen as a template to model the structure of DLL4 by MODELLER9v8.20 The structure of the R191H mutant was generated in a similar way to that for wild type. Ten models were generated and the top scoring model (DOPE Score) was analyzed and used for figures. Figures were prepared in Chimera. (a) The overall structure of DLL1 (PDB ID: 4XBM) showing the C2 domain and Arg195. (b) Modeled DLL4 structure showing Arg191 in a similar position to that of Arg195 in DLL1. Close-up view of Arg191 in the DLL4 model (c) and of His191 in the R191H mutant (d). The DLL1 structure is used to build up a model of DLL4.
Discussion
Involvement of the Notch signaling pathway in vascular development has been demonstrated by both gain- and loss-of-function mutations in humans, mice and zebrafish.13 DLL4 is implicated as an essential modulator of arteriogenesis and angiogenesis albeit together with NOTCH1.12 In mice, DLL4 is specifically expressed in arterial endothelial cells, and haploinsufficiency of DLL4 is embryonic lethal due to defective arterial and vascular development.14 DLL4 also regulates the formation of normal vessel sprouting and branching in the mouse retina and DLL4 expression in human is also predominantly observed in vascular endothelium.15 It has also been suggested that DLL4 is essential only at times of active vascularization, but is less important for maintaining normal vessel architecture.16 DLL4 contains five domains, an extracellular N-terminal domain, a Delta/Serrate/Lag-2 (DSL) domain, EGF-like domains, a membrane penetrating domain and an intracellular domain (Figure 2c). Structural analysis has shown that the DSL domain in DLL4 binds to the EGF-like domains in NOTCH1, the interaction being controlled by the mutual interaction between the glycosylated EGF-like domains in NOTCH1 and the N-terminal domain in DLL4.17 The patient described in this paper carries a heterozygous missense mutation in DLL4, resulting in an Arg to His substitution at codon 191 within the DSL domain. This arginine is located at the surface of the DSL domain in DLL4. Moreover, Arg191 is well conserved in various species, suggesting these sites have important roles in binding the EGF-like domain 11 in NOTCH1.17 Although DLL4 has not yet been crystalized, BLAST analysis showed that human DLL4 has good pairwise sequence similarity with Notch ligand Delta-like 1 (DLL1)18 (Figures 2d(a) and d(b)). Arg195 of DLL1 is located within the DSL domain and is also conserved, and is crucial for the interaction with the receptor (Figure 2d(a)).18 It is thus expected that Arg191 in DLL4 and Arg195 have similar roles in ligand binding. Therefore, Arg191 in DLL4 is suggested to have a key role in binding the Notch receptor.19 By modeling the DLL4 structure and the R191H mutant using MODELLER9v8,20 the side chain of histidine was shown to be shorter compared to that of arginine (Figures 2d(c) and d(d)). Therefore, the Arg191His mutation might lead to weakening of the electrostatic interaction between DLL4 and the receptor and it may have reduced access to receptor binding site residues, resulting in the AOS phenotype. The present case is likely to be caused by a loss-of-function mutation in the DSL domain.
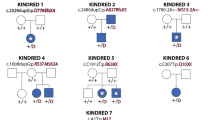
In AOS, although rare, complications such as cardiac anomaly, and neurological symptoms such as epilepsy and cortical dysplasia have been reported.11, 16 We compared the phenotypes of our case and nine previously observed families11 as shown in Table 1. We could not find any distinct phenotype–genotype correlations with the mutated domains of the DLL4 protein in these cases because of limited numbers. However, four cases, including our case, who carry mutations in the DLS domain seemed to have milder symptoms compared with patients with mutations in other domains. This was not conclusive, owing to limited sample size. Therefore, further analysis will be necessary to confirm genotype–phenotype correlations in AOS with DLL4 mutations. In our case, ossification of the skull was remarkable, suggesting that mutation in DLL4 has less effect after birth, leading to the milder phenotype of the case.
We report here a novel de novo pathogenic mutation in DLL4 in a sporadic AOS case. This genetic analysis was conducted because a precise diagnosis was desired by the parents, to obtain information on the patient’s natural history and prognosis, and for their future family planning. NGS was used to provide precise AOS information and the genetic information required for informed genetic counseling.
Change history
25 August 2017
This article has been corrected since Advance Online Publication, and a corrigendum is also printed in this issue.
References
Adams, F. H. & Oliver, C. P. Hereditary deformities in man-due to arrested development. J. Hered. 36, 2–7 (1945).
Hoyme, H. E., Jones, K. L., Van Allen, M. I., Saunders, B. S. & Benirschke, K. Vascular pathogenesis of transverse limb reduction defects. J. Pediatr. 101, 839–843 (1982).
Whitley, C. B. & Gorlin, R. J. Adams-Oliver syndrome revisited. Am. J. Med. Genet. 40, 319–326 (1991).
Bork, K. & Pfeifle, J. Multifocal aplasia cutis congenita, distal limb hemimelia, and cutis marmorata telangiectatica in a patient with Adams-Oliver syndrome. Br. J. Dermatol. 127, 160–163 (1992).
Southgate, L., Machado, R. D., Snape, K. M., Primeau, M., Dafou, D., Ruddy, D. M. et al. Gain-of-function mutations of ARHGAP31, a Cdc42/Rac1 GTPase regulator, cause syndromic cutis aplasia and limb anomalies. Am. J. Hum. Genet. 88, 574–585 (2011).
Hassed, S. J., Wiley, G. B., Wang, S., Lee, J. Y., Li, S., Xu, W. et al. RBPJ mutations identified in two families affected by Adams-Oliver syndrome. Am. J. Hum. Genet. 91, 391–395 (2012).
Stittrich, A. B., Lehman, A., Bodian, D. L., Ashworth, J., Zong, Z., Li, H. et al. Mutations in NOTCH1 cause Adams-Oliver syndrome. Am. J. Hum. Genet. 95, 275–284 (2014).
Southgate, L., Sukalo, M., Karountzos, A. S., Taylor, E. J., Collinson, C. S., Ruddy, D. et al. Haploinsufficiency of the NOTCH1 receptor as a cause of Adams-Oliver syndrome with variable cardiac anomalies. Circ. Cardiovasc. Genet. 8, 572–581 (2015).
Shaheen, R., Faqeih, E., Sunker, A., Morsy, H., Al-Sheddi, T., Shamseldin, H. E. et al. Recessive mutations in DOCK6, encoding the guanidine nucleotide exchange factor DOCK6, lead to abnormal actin cytoskeleton organization and Adams-Oliver syndrome. Am. J. Hum. Genet. 89, 328–333 (2011).
Shaheen, R., Aglan, M., Keppler-Noreuil, K., Faqeih, E., Ansari, S., Horton, K. et al. Mutations in EOGT confirm the genetic heterogeneity of autosomal-recessive Adams-Oliver syndrome. Am. J. Hum. Genet. 92, 598–604 (2013).
Meester, J. A., Southgate, L., Stittrich, A. B., Venselaar, H., Beekmans, S. J., den Hollander, N. et al. Heterozygous loss-of-function mutations in DLL4 cause Adams-Oliver syndrome. Am. J. Hum. Genet. 97, 475–482 (2015).
Liu, Z. J., Shirakawa, T., Li, Y., Soma, A., Oka, M., Dotto, G. P. et al. Regulation of Notch1 and Dll4 by vascular endothelial growth factor in arterial endothelial cells: implications for modulating arteriogenesis and angiogenesis. Mol. Cell. Biol. 23, 14–25 (2003).
Shawber, C. J. & Kitajewski, J. Notch function in the vasculature: insights from zebrafish, mouse and man. Bioessays 26, 225–234 (2004).
Gale, N. W., Dominguez, M. G., Noguera, I., Pan, L., Hughes, V., Valenzuela, D. M. et al. Haploinsufficiency of delta-like 4 ligand results in embryonic lethality due to major defects in arterial and vascular development. Proc. Natl. Acad. Sci. USA 101, 15949–15954 (2004).
Shutter, J. R., Scully, S., Fan, W., Richards, W. G., Kitajewski, J., Deblandre, G. A. et al. Dll4, a novel Notch ligand expressed in arterial endothelium. Genes Dev. 14, 1313–1318 (2000).
Ridgway, J., Zhang, G., Wu, Y., Stawicki, S., Liang, W. C., Chanthery, Y. et al. Inhibition of Dll4 signalling inhibits tumour growth by deregulating angiogenesis. Nature 444, 1083–1087 (2006).
Luca, V. C., Jude, K. M., Pierce, N. W., Nachury, M. V., Fischer, S. & Garcia, K. C. Structural biology. Structural basis for Notch1 engagement of Delta-like 4. Science 347, 847–853 (2015).
Kershaw, N. J., Church, N. L., Griffin, M. D. W., Luo, C. S., Adams, T. E. & Burgess, A. W. Notch ligand delta-like1: X-ray crystal structure and binding affinity. Biochem. J. 468, 159–166 (2015).
Chillakuri, C. R., Sheppard, D., Ilagan, M. X. G., Holt, L. R., Abbott, F., Liang, S. et al. Structural analysis uncovers lipid-binding properties of Notch ligands. Cell Rep. 5, 861–867 (2013).
Eswar, N., Webb, B., Marti-Renom, M. A., Madhusudhan, M. S., Eramian, D. & Shen, M.-Y. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. Chapter 2 (9), 1–31 (2007).
Acknowledgements
We greatly appreciate the patient and his family members’ involvement in this study. We also thank Asami Kita and Michiko Ujihara for technical assistance. We greatly appreciate critical comments on structural analysis to Professor Naoyuki Taniguchi. This work was supported by a donation from Kobe City to the Department of General Pediatrics, Kobe University Graduate School of Medicine (K550003302).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Nagasaka, M., Taniguchi-Ikeda, M., Inagaki, H. et al. Novel missense mutation in DLL4 in a Japanese sporadic case of Adams–Oliver syndrome. J Hum Genet 62, 851–855 (2017). https://doi.org/10.1038/jhg.2017.48
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2017.48