Abstract
Waardenburg syndrome (WS) is an autosomal dominant inherited neurogenic disorder with the combination of various degrees of sensorineural deafness and pigmentary abnormalities affecting the skin, hair and eye. The four subtypes of WS were defined on the basis of the presence or absence of additional symptoms. Mutation of human microphthalmia-associated transcription factor (MITF) gene gives rise to WS2. Here, we identified a novel WS-associated mutation at the stop codon of MITF (p.X420Y) in a Chinese WS2 patient. This mutation resulted in an extension of extra 33 amino-acid residues in MITF. The mutant MITF appeared in both the nucleus and the cytoplasm, whereas the wild-type MITF was localized in the nucleus exclusively. The mutation led to a reduction in the transcriptional activities, whereas the DNA-binding activity was not altered. We show that the foremost mechanism was haploinsufficiency for the mild phenotypes of WS2 induced in X420Y MITF.
Similar content being viewed by others
Introduction
Waardenburg syndrome (WS) is a disorder with the combination of various degrees of sensorineural deafness and pigmentary abnormalities affecting the skin, hair and eye. The four subtypes of WS were defined based on the presence or absence of additional symptoms1, 2 including developmental anomalies of the eyelids, eyebrows and nose root with pigmentary defects of the iris, head hair and with congenital deafness. This syndrome is found in ~2–5% of all congenitally deaf persons, and in 0.9–2.8% of the deaf population,3 which equally affects both sexes and all races. Several genes have been associated with WS, including PAX3, MITF, SNAI2, EDNRB, EDN3 and SOX10.4, 5, 6, 7, 8, 9, 10, 11, 12
The microphthalmia-associated transcription factor (MITF) encodes a 419-amino-acid residue, which contains an acid region (AA117–184) as the activation domain, a basic domain (AA199–217) for DNA binding, the basic helix–loop–helix domain (218–261) for dimerization and the Zip domain (262–290).13 MITF plays an important role in melanocyte development through regulating the expression of tyrosinase (TYR) and tyrosinase-related protein 1 (TYRP1), by binding to symmetrical DNA sequences (5ʹ-CACGTG-3ʹ, that is, E-boxes) in their promoters.14
Most WS-associated MITF mutations are located in exons 7, 8 and 9 that correspond to the b-helix–loop–helix-Zip motifs.1, 6, 15, 16, 17, 18, 19 The mutants may impair the transcription activities, phosphorylation, the capacity of DNA binding and nuclear localization.20, 21, 22 It seems more likely to result in haploinsufficiency of MITF function, with consequent downregulation of TYR gene expression.17
Here, we identified a novel WS-associated mutation at the stop codon of MITF (p.X420Y) in a Chinese WS2 patient. This mutation resulted in an extension of extra 33 amino-acid residues in MITF and led to a reduction in the transcriptional activities.
Materials and methods
The patient and control subjects
The patient was diagnosed as WS2 at ShanXi Special Education Center. Two hundred randomly selected normal individuals were also included in this study. This study was approved by committee of Xiangya Hospital of Central South University (equivalent to an Institutional Review Board).
Sanger sequencing
Genomic DNA was extracted from the peripheral blood cells of subjects and healthy controls using the phenol/chloroform method. We performed PCR Sanger sequencing on all coding exons and flanking intronic sequences of WS2-associated MITF (OMIM #156845) and SNAI2 (OMIM #602150). PCR was carried out using the standard protocol (Qiagen, Germantown, MD, USA). The PCR products were treated with the shrimp alkaline phosphatase and exonuclease-I to degrade deoxynucleotide triphosphates and unincorporated PCR primers. The purified amplicons were mixed with 10 picomoles of the forward and reverse PCR primers for bidirectional sequencing on an ABI-Prism 3100 DNA sequencer via dye termination chemistry (Applied Biosystems, Foster City, CA, USA). SeqMan II program (DNA-STAR, Madison, WI, USA) was used to compare results. DNA samples from the controls were further screened for the same mutation.
Reporter and expression constructs
The luciferase reporter containing the human TYR promoter (pGL3-Tyr-Luc) and the pCMV-3 × Flag-MITF plasmid had been described previously and were kindly provided by Vachtenheim et al.23, 24 The nonstop mutation (p.X420Y) introduced 33 extra amino-acid residues (YRILPALHSHKLLPFLIRRFNNLPEGVFLIIFL) to the C terminus of MITF, which was introduced by PCR and cloned into pCMV-3 × Flag.24 The sequences were reviewed by direct nucleotide sequencing.
Cell culture, transfections and luciferase reporter assays
The following cell lines were used in this study: NIH3T3 fibroblast cells and human melanoma UACC903 cells. All cells were grown at 37°C and 5% CO2 in Dulbecco’s modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (Invitrogen, Carlsbad, CA, USA) and 100 U ml−1 of penicillin/streptomycin (Invitrogen). The cells were prepared in 24-well plates at approximately 50% confluence a day in advance, and then the UACC903 cells were co-transfected transiently with 5 ng of reporter plasmid, 20 ng of expression plasmid and 5 ng of pCMV-β-gal (BD Biosciences/Clontech, Palo Alto, CA, USA), according to the manufacturer’s protocol in advance using Lipofectamine 2000 reagent (Invitrogen). Empty vectors were used to adjust the amount of DNA to 200 ng in each well. The cells were harvested 48 h later; after washing once with phosphate-buffered saline (PBS), the transfected UACC903 cells were incubated on ice with Reporter Lysis Buffer (Promega, Madison, WI, USA). The received samples were subsequently analyzed for luciferase activities and β-galactosidase by the SIRIUS luminometer (Berthold Detection System GmbH, Prorzheim, Germany). The test was repeated at least three times and performed in triplicate using different batches of cells. The data were analyzed and histograms were obtained using Prism 5 software (GraphPad Software, San Diego, CA, USA).
Western blot analysis
The UACC903 cells were cultured in 12-well plates and transfected with 400 ng of 3 × Flag-MITF or the 3 × Flag-X420Y-MITF expression plasmids dealing with Lipofectamine 2000 reagent. The cells were harvested 48 h later, and were incubated on ice with 2 × SDS loading buffer, which was composed of 0.2 mm β-mercaptoethanol supplemented with protease inhibitor cocktail (Sigma, St Louis, MO, USA) and 1 mm phenylmethane sulfonyl fluoride (Sigma). Proteins were separated on 12% SDS–polyacrylamide gel electrophoresis and transferred onto a polyvinylidene fluoride membrane (Millipore, Billerica, MA, USA). Blocking was performed at room temperature for 2.5 h with 5% nonfat milk dissolved in 0.2% Triton X-100 in PBS and then stained with mouse monoclonal anti-Flag M2 antibody (1:1000 dilution, Sigma) at 4 °C overnight. About 0.1% Tween 20 (Sigma) supplemented in Tris-buffered saline was used to wash the membrane three times, and then the membrane was stained for 1.5 h with a horseradish-peroxidase-conjugated anti-mouse IgG antibody (1:10000 dilution, Sigma) at room temperature. In the light of the manufacturer’s instructions, detection was performed using the ECL plus Western blotting detection system (GE Healthcare, Buckinghamshire, UK)
Biotinylated DNA affinity precipitation
The MITF-binding oligonucleotide 5′-GAAAAGTCAGTCATGTGCTTTTCAG-3′ and mutant DNA-binding probe 5′-GAAAAGTCAGTGTGCATCTTTTCAG-3′ were derived from human TYR gene’s promoter, were biotinylated at the 5’ terminus and then wild-type (WT) and mutant double-stranded oligonucleotides were generated by annealing with its complementary strand, respectively. The transfected melanoma UACC903 cells by Flag-MITF or Flag-X420Y-MITF expression plasmids were cultured in 6-well plates. Forty-eight hours later, cell lysates were obtained by sonication in HKMG buffer (10 mm HEPES, pH 7.9, 100 mm KCl, 5 mm MgCl2, 10% glycerol, 1 mm DTT and 0.5% of Nonidet-P40) supplemented with protease inhibitors. The cell lysates were centrifuged for 5 min at 4 °C at 13000 r.p.m., and cellular debris was removed. Again they are centrifuged for 30 min, and everted the supernatant at 4 °C for 3 h. Cell lysates of 10% were extracted and supplemented with 2 × SDS/0.2 mm β-mercaptoethanol for input immunoblotting. The remaining cell lysates were divided into two parts and supplemented with HKMG buffer to a final volume of 1 ml, and then everted it at 4 °C with or without 8 μg of biotinylated double-stranded oligonucleotide and 10 μg of poly(dI-dC).poly (dI-dC) (Sigma). About 20 h later, samples were incubated by rolling over again with 40 μl of streptavidin agarose beads (Sigma) for 3 h at 4 °C.
After washing the beads/DNA/protein or beads/protein complexes three times with ice-cold HKMG buffer, the samples were resuspended in 2 × SDS/0.2 mm β-mercaptoethanol. The samples were boiled for 10 min at 95 °C and subsequently separated by 12% SDS–polyacrylamide gel electrophoresis. In the subsequent immunoblotting, anti-Flag M2 antibody was added.
Immunofluorescence assays
The acid and alkali solutions were prepared in advance in a 15- mm microscope glass; the NIH3T3 cells or UACC903 cells were placed in 24-well plates. The cells were transfected with 200 ng of each plasmid construct. After transfection for 48 h, the cells were fixed with 4% paraformaldehyde at room temperature for 30 min. Then they were permeabilized for 1 h in PBS/0.2% Triton X-100 (Bio Basic, Amherst, NY, USA), blocked with solution (PBS, 3% bovine serum albumin, 5% goat serum) for 1 h at room temperature and incubated with primary anti-Flag M2 (1:600 dilution) at 4 °C overnight. The cells were washed with PBS/0.2% Triton X-100 for 5 min three times, and incubated for 2 h by avoiding light for preparing fluorescence-labeled secondary anti-mouse antibodies (1:300 dilution; Invitrogen). Then they were incubated with 4,6-diamino-2-phenylindole (DAPI; Invitrogen) for 3 min. Cells were detected by fluorescence images with a laser scanning confocal system installed on a Carl Zeiss microscope (Zeiss, GÖttingen, Germany) with a × 100 oil-immersion objective. Images were analyzed using the Metaphor software package (Zeiss).
Results
Identification of a nonstop mutation in MITF gene in a WS2 patient
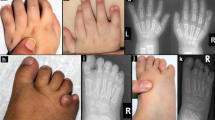
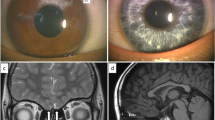
The subject was a 19-year-old girl from ShanXi Special Education Center. She was an orphan and grew up in a welfare center. She showed severe-to-profound bilateral sensorineural prelingual hearing loss without vestibular impairment. She had mild hypoplastic blue irises and a hand of white forelock, but lacked pigmentary anomalies in the skin or dystopia canthorum and Hirschsprung disease. Thus, she was diagnosed with WS2. Large vestibule duct and deformity was not found in inner by temporal bone CT.
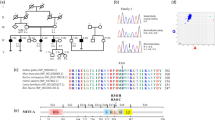
PCR Sanger sequencing of all coding exons and flanking intronic sequences of MITF and SNAI2 were performed. In consequence, we determined a novel heterozygous MITF mutation, c.1260G>C (p.X420Y), which was not detected in 200 normal controls (Figure 1). The MITF gene is 1260 bp in length and encodes 419 amino acids. This mutation occurred at the stop codon (TAG was changed to TAC), resulting in an extension of extra 33 amino-acid residues (YRILPALHSHKLLPFLIRRFNNLPEGVFLIIFL); the molecular weight of the mutant protein was predicted to be 50.9 kDa, whereas the WT one is 46.9 kDa.
Schematic representation of a novel mutation and function in the MITF gene. (a) Identification of a novel mutation in the MITF gene in a sporadic WS2. Sequence chromatograms of the wild-type control (top) and MITF X420Y mutation (below). The arrow above the chromatogram highlights the base affected by the mutation. (b) In humans, the MITF locus is mapped to chromosome 3 and spans 229 kbp. MITF is characterized by a b-HLH-Zip (basic helix–loop–helix leucine zipper) DNA binding and dimerization, which targets (T)CATGTGC(A) motifs and amino-terminal transactivation domain (AD). Proteins containing b-HLH-Zip regulate gene expression by binding to the E-box (CANNTG) as heterodimers or homodimers. Ten point mutations pathogenic to WS and eight among them located precisely in exon. They centralized in the b-HLH-Zip domain. X420Y is a missense mutant and the site in the stop codon, last exon, induced to extend the protein, which disrupts dimer formation, and as a result causes insufficient development of melanocytes. MITF, microphthalmia-associated transcription factor; WS, Waardenburg syndrome.
Functional analysis of mutant mitf
Initially, to determine whether X420Y mutation affects the transcription activity of MITF, we transfected the WT and X420Y MITF expression plasmids into UACC903 cells, respectively, with the luciferase reporter under control of TYR promoter (pGL3-Tyr-Luc) (Figure 2b). The results show that (Figure 2a) WT MITF improved the TYR promoter activity up to 17-fold, consistent with previous reports;23, 24 however, X420Y MITF only induced threefold increase.
(a) Transcriptional activities of MITF and X420Y determined by luciferase activity assays. The luciferase reporter plasmid Tyr-Luc was transiently transfected into melanoma UACC903 cells in combination with 20 ng of WT or X420Y expression plasmids. (b) Effect of X420Y on WT MITF transactivity. Increasing amounts of mutant X420Y expression plasmids were co-transfected with a fixed amount of WT MITF expression plasmid (20 ng of the MITF plasmids constantly, 20, 40 and 80 ng of the X420Y plasmids, respectively) and the luciferase reporter plasmid Tyr-Luc. For a, b, the basal level of luciferase was set as 1. Data from all other transfections are presented as fold induction above this level. Luciferase activity was normalized by measuring β-galactosidase activity. Each value was shown as mean±s.e.m. of three replicates from a single assay. The results shown were representative of at least three independent experiments. (c) Melanoma UACC903 cells were transiently transfected with 200 ng of the plasmids expression either the WT or X420Y in 12-well plates. Proteins from whole-cell lysates were analyzed by SDS–PAGE (12%), and the expression of MITF was visualized using monoclonal antibody against Flag epitope (#P<0.05, ##P<0.01, as compared with the basal activity; P<0.01, as compared with the WT; unpaired Student’s t-test). (d) The graph described the difference in the expression levels of wild-type and mutant MITF proteins by densitometry with the same quantifications of WT and X420Y plasmids transfected in melanoma UACC903. The mutant protein has low density compared with WT. MITF, microphthalmia-associated transcription factor; ns, not significant; WT, wild type.
To confirm whether the p.X420Y MITF expresses normally, western blot experiments were performed next using the mouse monoclonal anti-Flag M2 antibody. It showed that the protein extracting from melanoma UACC903 cells transfected with WT or X420Y MITF expression plasmids expressed light differences in molecular weight (Figures 2c and d).
To ascertain whether X420Y was capable of binding the DNA like the WT MITF, we precipitated the protein abstracts from the melanoma UACC903 cells transfected with WT or X420Y MITF expression plasmids using a biotinylated double-stranded oligonucleotide containing MITF-binding sequence as our previous work.24 The X420Y MITF showed similar DNA-binding activity as WT MITF (Figure 3).
DNA-binding capacity of WS-associated MITF. (a) The melanoma UACC903 cells transfected with 1µg WT, X420Y plasmids were incubated with or without biotinylated double-stranded oligonucleotides of the MITF-binding region at the TYR promoter. The DNA/protein complex was pulled down with streptavidin agarose beads. The precipitated proteins were separated on SDS–PAGE in equal amounts of whole-cell lysates and analyzed by immunoblotting using anti-Flag M2 antibody. Actin was used as an internal control. DNA precipitation demonstrates that X420Y mutant retains the DNA-binding activity. As a negative control, WT and X420Y MITF protein did not bind to mutated double-stranded oligonucleotides. There were significant differences in the expression of WT and X420Y MITF protein binding with the TYR promoter. (b) The graph illustrates the quantification of WT and X420Y MITF by densitometry of experiments. (***P<0.05, by unpaired t-test). MITF, microphthalmia-associated transcription factor; SDS–PAGE, SDS–polyacrylamide gel electrophoresis; TYR, tyrosinase; WS, Waardenburg syndrome.
We then performed immunofluorescence to examine the subcellular distribution of WT and mutant MITF. As expected, WT MITF was localized in the nucleus; however, the X420Y-MITF was detected in both nucleus and cytoplasm (Figures 4 and 5).
Subcellular localization of WS-associated MITF proteins and X420Y-NLS in NIH3T3 cells. WT MITF and X420Y proteins (a, d) are shown in blue; DAPI (b, e) revealing nucleus is shown in green; and the merged images (c, f) are shown. Scale bar: 25 µm. MITF, microphthalmia-associated transcription factor; WS, Waardenburg syndrome.
Subcellular localization of WS-associated MITF proteins and X420Y-NLS in UACC903 cells. WT MITF and X420Y proteins (a, d) are shown in blue; DAPI (b, e) revealing nucleus is shown in red; and the merged images (c, f) are shown. Scale bar: 25 µm. MITF, microphthalmia-associated transcription factor; WS, Waardenburg syndrome.
Discussion
There are close to 60 000 missense/nonsense mutation reports in 4000 different genes in the Human Gene Mutation Database (HGMD), all of which are linked to human genetic disorders.25 Among them, a special mutation termed ‘nonstop’, ‘nostop’ or ‘readthrough’ mutations, which occurred within stop codons, estimated only 119 mutations (in 87 different genes).25
These mutations caused termination failure in normal translation stop codons and were likely to result in a persistent translation of messenger RNA (mRNA) downstream into the 3'-untranslated region. The ribosome could continue translation in the same reading frame until a subsequent stop codon, leading to proteins including additional peptides.26, 27 The distance from the occurrence of the next stop codon is a determining factor for whether the clinical disease phenotype is caused. Some of the stop codons were replaced by base-pair mutations, but another stop codon appeared shortly afterward, and thus the translated amino-acid lengths and biological functions approached normal proteins.28 On the other hand, no alternative stop codons appeared within 49 nucleotides; the nonstop mutations is worthy of clinical attention because of the potential for disease. It probably explicated that the decay of the nonstop mRNA was triggered by the amino-acid residues at some distance from the mutated stop codon. Suitably, the amount of protein synthesis was distinctly decreasing and giving rise to a clinical phenotype enough to cause clinical concern.25 For instance, the nonstop mutant mRNA transcripts in RPS19 (Diamond–Blackfan anemia),29 F10 (factor X deficiency)30 and FOXE3 (anterior segment dysgenesis)31 were found to be severely lower than those of their WT counterparts. X420Y mutation in MITF introduces 99 nucleotide acids before the alternative in-frame stop codon; it is likely to trigger nonstop mRNA decay.32, 33
MITF regulates the transcription of TYR gene, which encodes TYR, a key enzyme involved in melanin synthesis. Ten point mutations have been reported in WS, which were located at exon 1, 4, 5, 7, 8 and 9, respectively (Figure 2b), and one mutation was located at the stop codon (c.1258 T>C).34 In this study, we identified a nonstop mutation (X420Y) in MITF in a WS patient with mild hypoplastic blue irises, hearing loss and white forelock in one hand. X420Y mutation introduced 99 nucleotide acids before the alterative in-frame stop codon, resulting in a mutant MITF protein with 33 extra amino acids at the C terminus and reduced transactivity on TYR promoter.
The basic domain of MITF may contain an NLS (210NLIERRRRFNIN221) at the C-terminal basic domain (http://cubic.bioc.columbia.edu/predictNLS/), which had been determined in our previous study.24 Furthermore, the nonstop mutation will introduce a number of amino acids with novel properties to a protein’s C terminus.35 Extremely hydrophobic residues might have a greater impact. The extended amino acids may alter the secondary, tertiary and quaternary structures of the protein, and influence the subcellular distribution and/or biological functions.36 In this study, X420Y mutation introduces an extra 33 amino acids to the C terminus of MITF. About 22 of these 33 amino acids are hydrophobic residues (alanine, isoleucine, leucine, proline, valine, phenylalanine or tyrosine). As a result, this nonstop mutation affected the subcellular localization and transcriptional activity, but did not influence the DNA-binding activity.
The missense and frameshift mutations in MITF seemed more likely to result in haploinsufficiency of MITF function, with subsequent downregulation of TYR gene expression.16, 17 However, there is no evidence showing that the mutated MITF proteins predicted by mutant MITF allele from patients with WS2 directly affect the function of the in silico WT MITF protein. The haploinsufficiency could be caused by the dimerization between the mutant and the WT protein, which may interfere with the access of the WT dimers to the nucleus.19
In summary, we have set forth one unusual mutation-associated WS2 identified in our previous study, which weakened the transcriptional activity of MITF. Haploinsufficiency is the most suitable mechanism rather than the dominant-negative effect. The mutation at the stop codon was in the heterozygous form where the wild MITF protein resulted from one copy of the normal allele and does not meet the criterion needed for full function of the protein.
References
Read, A. P. & Newton, V. E. Waardenburg syndrome. J. Med. Genet. 34, 656–665 (1997).
Solia-Nasser, L., de Aquino, S. N., Paranaiba, L. M., Gomes, A., Dos-Santos-Neto, P., Coletta, R. D. et al. Waardenburg syndrome type I: dental phenotypes and genetic analysis of an extended family. Med. Oral. Patol. Oral. Cir. Bucal. 21, e321–e327 (2016).
Song, J., Feng, Y., Acke, F. R., Coucke, P., Vleminckx, K. & Dhooge, I. J. Hearing loss in Waardenburg syndrome: a systematic review. Clin. Genet. 89, 416–425 (2016).
Baldwin, C. T., Hoth, C. F., Amos, J. A., da-Silva, E. O. & Milunsky, A. An exonic mutation in the HuP2 paired domain gene causes Waardenburg's syndrome. Nature 355, 637–638 (1992).
Hoth, C. F., Milunsky, A., Lipsky, N., Sheffer, R., Clarren, S. K. & Baldwin, C. T. Mutations in the paired domain of the human PAX3 gene cause Klein-Waardenburg syndrome (WS-III) as well as Waardenburg syndrome type I (WS-I). Am. J. Hum. Genet. 52, 455–462 (1993).
Tassabehji, M., Newton, V. E. & Read, A. P. Waardenburg syndrome type 2 caused by mutations in the human microphthalmia (MITF gene. Nat. Genet. 8, 251–255 (1994).
Sanchez-Martin, M., Rodriguez-Garcia, A., Perez-Losada, J., Sagrera, A., Read, A. P. & Sanchez-Garcia, I. SLUG (SNAI2) deletions in patients with Waardenburg disease. Hum. Mol. Genet. 11, 3231–3236 (2002).
Edery, P., Attie, T., Amiel, J., Pelet, A., Eng, C., Hofstra, R. M. et al. Mutation of the endothelin-3 gene in the Waardenburg-Hirschsprung disease (Shah-Waardenburg syndrome). Nat. Genet. 12, 442–444 (1996).
Hofstra, R. M., Osinga, J., Tan-Sindhunata, G., Wu, Y., Kamsteeg, E. J., Stulp, R. P. et al. A homozygous mutation in the endothelin-3 gene associated with a combined Waardenburg type 2 and Hirschsprung phenotype (Shah-Waardenburg syndrome). Nat. Genet. 12, 445–447 (1996).
Inoue, K., Khajavi, M., Ohyama, T., Hirabayashi, S., Wilson, J., Reggin, J. D. et al. Molecular mechanism for distinct neurological phenotypes conveyed by allelic truncating mutations. Nat. Genet. 36, 361–369 (2004).
Bondurand, N., Dastot-Le Moal, F., Stanchina, L., Collot, N., Baral, V., Marlin, S. et al. Deletions at the SOX10 gene locus cause Waardenburg syndrome types 2 and 4. Am. J. Hum. Genet. 81, 1169–1185 (2007).
Shi, Y., Li, X., Ju, D., Li, Y., Zhang, X. & Zhang, Y. A novel mutation of the MITF gene in a family with Waardenburg syndrome type 2: a case report. Exp. Ther. Med. 11, 1516–1518 (2016).
Hou, L. & Pavan, W. J. Transcriptional and signaling regulation in neural crest stem cell-derived melanocyte development: do all roads lead to Mitf? Cell Res. 18, 1163–1176 (2008).
Chen, L., Guo, W., Ren, L., Yang, M., Zhao, Y., Guo, Z. et al. A de novo silencer causes elimination of MITF-M expression and profound hearing loss in pigs. BMC Biol. 14, 52 (2016).
Pingault, V., Ente, D., Dastot-Le Moal, F., Goossens, M., Marlin, S. & Bondurand, N. Review and update of mutations causing Waardenburg syndrome. Hum. Mutat. 31, 391–406 (2010).
Nobukuni, Y., Watanabe, A., Takeda, K., Skarka, H. & Tachibana, M. Analyses of loss-of-function mutations of the MITF gene suggest that haploinsufficiency is a cause of Waardenburg syndrome type 2A. Am. J. Hum. Genet. 59, 76–83 (1996).
Morell, R., Spritz, R. A., Ho, L., Pierpont, J., Guo, W., Friedman, T. B. et al. Apparent digenic inheritance of Waardenburg syndrome type 2 (WS2) and autosomal recessive ocular albinism (AROA). Hum. Mol. Genet. 6, 659–664 (1997).
Lalwani, A. K., Attaie, A., Randolph, F. T., Deshmukh, D., Wang, C., Mhatre, A. et al. Point mutation in the MITF gene causing Waardenburg syndrome type II in a three-generation Indian family. Am. J. Med. Genet. 80, 406–409 (1998).
Izumi, K., Kohta, T., Kimura, Y., Ishida, S., Takahashi, T., Ishiko, A. et al. Tietz syndrome: unique phenotype specific to mutations of MITF nuclear localization signal. Clin. Genet. 74, 93–95 (2008).
Smith, S. D., Kelley, P. M., Kenyon, J. B. & Hoover, D. Tietz syndrome (hypopigmentation/deafness) caused by mutation of MITF. J. Med. Genet. 37, 446–448 (2000).
Wilkie, A. O. The molecular basis of genetic dominance. J. Med. Genet. 31, 89–98 (1994).
Takeda, K., Takemoto, C., Kobayashi, I., Watanabe, A., Nobukuni, Y., Fisher, D. E. et al. Ser298 of MITF, a mutation site in Waardenburg syndrome type 2, is a phosphorylation site with functional significance. Hum. Mol. Genet. 9, 125–132 (2000).
Vachtenheim, J., Novotna, H. & Ghanem, G. Transcriptional repression of the microphthalmia gene in melanoma cells correlates with the unresponsiveness of target genes to ectopic microphthalmia-associated transcription factor. J. Invest. Dermatol. 117, 1505–1511 (2001).
Zhang, H., Luo, H., Chen, H., Mei, L., He, C., Jiang, L. et al. Functional analysis of MITF gene mutations associated with Waardenburg syndrome type 2. FEBS Lett. 586, 4126–4131 (2012).
Hamby, S. E., Thomas, N. S., Cooper, D. N. & Chuzhanova, N. A meta-analysis of single base-pair substitutions in translational termination codons ('nonstop' mutations) that cause human inherited disease. Hum. Genomics 5, 241–264 (2011).
Bruno, C., Martinuzzi, A., Tang, Y., Andreu, A. L., Pallotti, F., Bonilla, E. et al. A stop-codon mutation in the human mtDNA cytochrome c oxidase I gene disrupts the functional structure of complex IV. Am. J. Hum. Genet. 65, 611–620 (1999).
Bjornsson, A., Mottagui-Tabar, S. & Isaksson, L. A. Structure of the C-terminal end of the nascent peptide influences translation termination. EMBO J. 15, 1696–1704 (1996).
Ikeuchi, K. & Inada, T. Ribosome-associated Asc1/RACK1 is required for endonucleolytic cleavage induced by stalled ribosome at the 3' end of nonstop mRNA. Sci. Rep. 6, 28234 (2016).
Chatr-Aryamontri, A., Angelini, M., Garelli, E., Tchernia, G., Ramenghi, U., Dianzani, I. et al. Nonsense-mediated and nonstop decay of ribosomal protein S19 mRNA in Diamond-Blackfan anemia. Hum. Mutat. 24, 526–533 (2004).
Ameri, A., Machiah, D. K., Tran, T. T., Channell, C., Crenshaw, V., Fernstrom, K. et al. A nonstop mutation in the factor (F)X gene of a severely haemorrhagic patient with complete absence of coagulation FX. Thromb. Haemost. 98, 1165–1169 (2007).
Doucette, L., Green, J., Fernandez, B., Johnson, G. J., Parfrey, P. & Young, T. L. A novel, non-stop mutation in FOXE3 causes an autosomal dominant form of variable anterior segment dysgenesis including Peters anomaly. Eur. J. Hum. Genet 19, 293–299 (2011).
Simms, C. L., Thomas, E. N. & Zaher, H. S. Ribosome-based quality control of mRNA and nascent peptides. Wiley Interdiscip. Rev. RNA 8, e1366 (2016).
Ikeuchi, K., Yazaki, E., Kudo, K. & Inada, T. Conserved functions of human Pelota in mRNA quality control of nonstop mRNA. FEBS Lett. 590, 3254–3263 (2016).
Sun, L., Li, X., Shi, J., Pang, X., Hu, Y., Wang, X. et al. Molecular etiology and genotype-phenotype correlation of Chinese Han deaf patients with type I and type II Waardenburg Syndrome. Sci. Rep. 6, 35498 (2016).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Doamekpor, S. K., Lee, J. W., Hepowit, N. L., Wu, C., Charenton, C., Leonard, M. et al. Structure and function of the yeast listerin (Ltn1) conserved N-terminal domain in binding to stalled 60S ribosomal subunits. Proc. Natl Acad. Sci. USA 113, E4151–E4160 (2016).
Acknowledgements
We thank the patient for her participation in the study and the ShanXI Special Education, especially the collaboration D Hao at the Department of Center Laboratory in Taiyuan Central Hospital of China. This study was funded by the National Basic Research Program of China (973 Program) (Grant No 2014CB541702, 2014CB943003 and 2014CB541701), Key Projects of Subordinate Hospital of Chinese Ministry of Health for Clinical Subjects and by grants from National Nature Science Foundation of China (81470705, 81500803, 81301172) and the Natural Science Foundation of Hunan Province, China (14JJ7009). J-DL is a recipient of Lotus Scholar Professorship from Hunan Province, China.
Ethical standards
All experiments performed in this study comply with the current laws of People’s Republic of China.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Sun, J., Hao, Z., Luo, H. et al. Functional analysis of a nonstop mutation in MITF gene identified in a patient with Waardenburg syndrome type 2. J Hum Genet 62, 703–709 (2017). https://doi.org/10.1038/jhg.2017.30
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2017.30
This article is cited by
-
Identification and in vivo functional investigation of a HOMER2 nonstop variant causing hearing loss
European Journal of Human Genetics (2023)
-
Waardenburg syndrome type 4 coexisting with open-angle glaucoma: a case report
Journal of Medical Case Reports (2022)
-
The Regulation of MiTF/TFE Transcription Factors Across Model Organisms: from Brain Physiology to Implication for Neurodegeneration
Molecular Neurobiology (2022)
-
Four mutations in MITF, SOX10 and PAX3 genes were identified as genetic causes of waardenburg syndrome in four unrelated Iranian patients: case report
BMC Pediatrics (2021)
-
A pan-cancer analysis reveals nonstop extension mutations causing SMAD4 tumour suppressor degradation
Nature Cell Biology (2020)