Abstract
In clinical practice, it is important to diagnose the carrier state of female patients with X-linked diseases for genetic counseling to calculate the recurrent risk of offspring. Because some X-linked diseases show high rates of gonadal mosaicism, this diagnosis is sometimes difficult, when there are few offspring in a family and no mutation is detected in the maternal genomic DNA. Here, we report two male siblings with ATR-X syndrome carrying an intragenic deletion of 78.6 kb involving exons 2–5 out of the 35 exons in the ATRX, as revealed by PCR amplification of these exons. The mother was expected to be an obligate carrier, but we could not confirm her as a mutation carrier by quantitative PCR (qPCR) for the exons. However, we identified the breakpoint of ATRX, and qPCR with breakpoint-specific primers revealed gonosomal mosaicism, with a relative frequency of the mutation of <1% in genomic DNA of her peripheral blood. For these obligate carriers of X-linked disease, we should aggressively investigate the maternal genomic status, not only because her genetic condition is important for estimating the recurrent risk of her offspring but also because a diagnosis of her gonosomal mosaicism can render negligible the possibility that her female siblings are carriers. We should reconfirm that a female who has a risk of being a carrier has a gonosomal or somatic mutation, even if she is an obligate carrier or apparently harbors a mutation.
Similar content being viewed by others
Introduction
X-linked α-thalassemia mental retardation syndrome (ATR-X syndrome; MIM# 603040) is characterized by severe intellectual disability, dysmorphic facies, hypotonia, genital and skeletal abnormalities and downregulation of the α-globin genes (α-thalassemia).1 More than 80 patients have been molecularly diagnosed in Japan and more than 200 patients worldwide.2 Most of their mutations are clustered in two functionally important regions: the ADD (ATRX-DNMT2-DNMT3) and helicase domains, resulting in loss of function.2
Generally, in sporadic cases of severe X-linked disease with zero-fitness, the probability that the mother is a mutation carrier is 2/3 and the probability that the patient has a de novo mutation is 1/3.3 Thus, it is important to diagnose the mother’s mutation carrier state to calculate the probability of the recurrence risk in the offspring. On the other hand, even if no mutations are detected in genomic DNA extracted from their peripheral blood, we should consider the possibility of germline, somatic or gonosomal mosaicism. Germline and gonosomal mosaicism have been recently reported in several other X-linked diseases,4 and two cases of mosaicism, including gonosomal and germline, have been documented among 20 families of sporadic cases with ATR-X syndrome.5
Here, we report the case of two brothers affected with ATR-X syndrome, having an intragenic deletion involving exons 2–5 of the 35 exons in the ATRX, and their mother exhibiting gonosomal mosaicism, with the mutant allele at <1% incidence in the peripheral blood. This finding renders negligible the probability that his mother’s female siblings are carriers of the mutation.
Case report
Case 1 (Figure 1; III-2) is a 9-year-old boy. He was suspected of having ATR-X syndrome at the age of 2 years on the basis of severe motor and psychiatric developmental delay, characteristic hypotonic facies, spaced teeth and presence of HbH inclusions in his brilliant cresyl blue (BCB)-stained peripheral blood. PCR of all exons and exon–intron boundaries was performed with specific primers, and PCR amplifications were not observed for exons 2–5 out of all 35 exons. His genomic DNA showed a deletion of exons 2–5 in the ATRX, confirming his diagnosis molecular genetically. His mother’s mutation carrier state could not be diagnosed. He had received surgery for undescended testes. He has a 1-week vomiting episode annually. He had gastroesophageal reflux during his infancy but has grown out of it. He was diagnosed as having epilepsy and takes valproic acid. He has recently begun walking with assistance. He cannot speak any meaningful words but can understand simple words and situations around him. His weight, height and head circumference are 18.3 kg (−1.9 s.d.), 116.3 cm (−3.1 s.d.) and 47.2 cm (−2.5 s.d.), respectively.
Case 2 (Figure 1; III-3) is a 5-year-old boy, a younger brother of Case 1. At birth, he presented with multiple congenital anomalies, including tetralogy of Fallot, heart anomaly, complete tracheal ring, bilateral severe deafness, total blindness due to bilateral persistent hyperplastic primary vitreous, hypospadias and bilateral undescended testis. He had operations in infancy for tetralogy of Fallot and gastroesophageal reflux. He had epilepsy at the age of 4 years. He shows very severe psychomotor developmental delay and central hypotonic facies. He can roll over but cannot sit unassisted. He cannot speak any meaningful words. His clinical condition is much severer than that of his brother and initially ATR-X syndrome was not suspected for his diagnosis.
He was referred to our medical center at 5 years. Approximately, 60% of his BCB-stained peripheral blood revealed HbH inclusions, and he was diagnosed to have the same mutation in the ATRX. His weight, height and head circumference are 10.185 kg (−3.0 s.d.), 95.5 cm (−3.4 s.d.) and 39.7 cm (−6.4 s.d.), respectively.
Results
Diagnosis of the patient (III-3) by PCR
We could not amplify exons 2–5 of the 35 exons in ATRX in the genomic DNA extracted from the peripheral blood of the patient (III-3), indicating an intragenic deletion involving exons 2–5 in ATRX as well as in his elder brother (III-2). His cDNA shows that the 3′ end of exon 1 abuts the 5′ end of exon 6, indicating exon skipping of exons 2–5 resulting in a 350-bp deletion (Supplementary Figure S1). This transcript is expected to be translated into a prematurely truncated protein, p.Ser7Argfs*13.
Identification of the breakpoint
We designed PCR primers to amplify the flanking region of the deleted regions of genomic DNA from the patient (Figure 2a). No amplification of amplicons using the primers was observed in a normal control (Figure 2c).
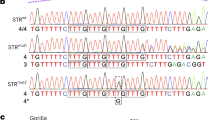
(a) (top) Schematic presentation of the part of X chromosome around ATRX on Xq21; (middle) the direction of the ATRX; and (bottom) a schema of the deleted exons of the ATRX detected in the patient. The deletion spans approximately 78.6 kb (chromosome X, NC_000023.10: 77030517-76951924). (b) Sequence chromatogram showing breakpoint of genomic DNA from patient III-3. (c) PCR using the primers flanking the deletion from intron 1 to intron 5 in the ATRX amplifies 1141-bp products in 40 cycles (top) and 30 cycles (middle). GAPDH is the reference gene (bottom). CTR1 and CTR2 are normal control samples. A full color version of this figure is available at the Journal of Human Genetics journal online.
We sequenced the PCR product and identified the 5′ end at the breakpoint in intron 1 and the 3′ end of intron 5, with primers flanking the deleted region. The deletion spans approximately 78.6 kb (chromosome X, NC_000023.10: 77030517-76951924) (Figure 2b).
Diagnosis of the mother’s carrier state
Quantitative PCR (qPCR) using primers for each exon did not detect a copy number difference between exon 5, which was deleted, and exon 6, which was not deleted (Figure 3).
The result of quantitative PCR using primers for exons 5 and 6. It did not detect a copy number difference between mother and a control for exon 5, which was deleted in the patient. HPRT (X-linked) is the reference gene. A full color version of this figure is available at the Journal of Human Genetics journal online.
Primers flanking the deletion amplified an 1144-bp product in both the patients and the mother at 40 cycles but not in the mother at 30 cycles (Figure 2c). These findings suggest that the mother carried the same deleted mutation by gonosomal mosaicism, considering that she has two sons with ATR-X.
Estimation of the relative frequency of the mutant allele
Using qPCR with primers flanking the deletion, we estimated at <1% the relative frequency of the mutant allele in genomic DNA from peripheral blood of the mother. A dilution series of known template concentrations of the patient’s genomic DNA was used to establish a standard curve. The standard curve showed a linear form (R2=0.9996) (Supplementary Figure S2).
Discussion
We report two affected brothers of ATR-X syndrome with an intragenic deletion of 78.6 kb involving exons 2–5 in the ATRX, resulting from their mother’s gonosomal mosaicism. It was technically difficult to confirm her mosaicism by qPCR for each exon of the ATRX, because the relative frequency of the mutant allele was low, at <1%. After we had identified the breakpoint in the patients’ genomic DNA and designed primers straddling the breakpoint, we could easily detect the mutant allele in her genomic DNA.
When we diagnosed the first patient by the usual PCR, we did not know the mother’s carrier state. The second patient was diagnosed as having ATR-X syndrome with the same mutation, and their mother was expected to be an obligate mutation carrier. There were two problems in diagnosing her carrier state, which are as follows: first, the mutation was a deletion that was not detected by conventional PCR and sequencing, and second, the relative frequency of the mutant allele in the peripheral blood was too low for the copy number to be determined by qPCR.
In fact, the information about the mother’s mosaicism does not aid in genetic counseling to calculate the recurrent risk of subsequent offspring, because she is an obligate mutation carrier. But this information is important for genetic counseling of her two sisters (Figure 1, II-3 and II-4), because her mutation must have occurred de novo after fertilization, and the probability that her sisters carry the same mutation is expected to be negligible. Therefore, their genetic counseling was not required.
In clinical practice, somatic, gonadal or gonosomal mosaicism presents several problems, particularly if a patient’s mother does not have the mutant allele in her blood; isolated gonadal mosaicism can be suspected, if the same mutant allele is transmitted to a second offspring and he is found to be affected. The relative frequency of the mutant allele in the mosaicism of her peripheral blood is not helpful for estimating the frequency in her ova. In our case, the relative frequency of the mutant allele in the ova could be <1% as in the peripheral blood or could be much higher, approximately 50%.
It is often difficult to contact the families of maternal female siblings and provide them with clinical information. The present study highlights the importance of thoroughly diagnosing the mother’s mutation carrier status, even if only one offspring is affected or she does not exhibit the mutation. These findings allow us to decide whether to inform her female siblings and their families.
In the next-generation sequencing era, we will encounter more patients or carriers with mosaicism of mutant alleles.6 We should reconsider mosaicism from the standpoint of genetic counseling.
References
Gibbons, R. J., Brueton, L., Buckle, V. J., Burn, J., Clayton-Smith, J., Davison, B. C. et al. Clinical and hematologic aspects of the X-linked alpha-thalassemia/mental retardation syndrome (ATR-X). Am. J. Med. Genet. 55, 288–299 (1995).
Gibbons, R. J., Wada, T., Fisher, C. A., Malik, N., Mitson, M. J., Steensma, D. P. et al. Mutations in the chromatin-associated protein ATRX. Hum. Mutat. 29, 796–802 (2008).
Young, I. D. Introduction to Risk Calculation in Genetic Counseling 2nd edn. (Oxford: New York, NY, USA, 1999).
Wang, Y., Busin, R., Reeves, C., Bezman, L., Raymond, G., Toomer, C. J. et al. X-linked adrenoleukodystrophy: ABCD1 de novo mutations and mosaicism. Mol. Genet. Metab. 104, 160–166 (2011).
Bachoo, S. & Gibbons, R. J. Germline and gonosomal mosaicism in the ATR-X syndrome. Eur. J. Hum. Genet. 7, 933–936 (1999).
Pagnamenta, A. T., Lise, S., Harrison, V., Stewart, H., Jayawant, S., Quaghebeur, G. et al. Exome sequencing can detect pathogenic mosaic mutations present at low allele frequencies. J. Hum. Genet. 57, 70–72 (2012).
Acknowledgements
We thank the patients and their families for their clinical information. This study was supported by research grants from the Ministry of Health, Labour and Welfare (H25-nanchi-ippan-114; to TW).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Shimbo, H., Ninomiya, S., Kurosawa, K. et al. A case report of two brothers with ATR-X syndrome due to low maternal frequency of somatic mosaicism for an intragenic deletion in the ATRX. J Hum Genet 59, 408–410 (2014). https://doi.org/10.1038/jhg.2014.45
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.45