Abstract
In order to investigate the underlying genetic structure and genomic ancestry proportions of Peruvian subpopulations, we analyzed 551 human samples of 25 localities from the Andean, Amazonian, and Coastal regions of Peru with a set of 40 ancestry informative insertion–deletion polymorphisms. Using genotypes of reference populations from different continents for comparison, our analysis indicated that populations from all 25 Peruvian locations had predominantly Amerindian genetic ancestry. Among populations from the Titicaca Lake islands of Taquile, Amantani, Anapia, and Uros, and the Yanque locality from the southern Peruvian Andes, there was no significant proportion of non-autochthonous genomes, indicating that their genetic background is effectively derived from the first settlers of South America. However, the Andean populations from San Marcos, Cajamarca, Characato and Chogo, and coastal populations from Lambayeque and Lima displayed a low but significant European ancestry proportion. Furthermore, Amazonian localities of Pucallpa, Lamas, Chachapoyas, and Andean localities of Ayacucho and Huancayo displayed intermediate levels of non-autochthonous ancestry, mostly from Europe. These results are in close agreement with the documented history of post-Columbian immigrations in Peru and with several reports suggesting a larger effective size of indigenous inhabitants during the formation of the current country’s population.
Similar content being viewed by others
Introduction
In recent years, some ancestry studies performed with Central and South American populations showed that ancestry proportions (in relation to aboriginal populations of continents) vary depending on their particular demographic dynamics and colonization history.1, 2, 3, 4
Like most Latin American populations,2, 4 current Peruvians were mainly formed during colonial times by three ancestral components: autochthonous Americans, Eurasians (mostly from Europe) and Africans. However, Peru is also known by the large numbers of indigenous populations reported when the Spaniards arrived in the region, particularly represented by the largest cities found in America and the vast Inca Empire at the time of European contact.5
In order to investigate the past dynamics of gene flow and continental roots of Latin American populations, ancestry components and admixture levels can be estimated with informative autosomal markers. A previous study with 642 690 randomly chosen autosomal single-nucleotide polymorphisms6 genotyped in reference continental populations from the HGDP-CEPH panel observed a remarkable structure according with their geographical distribution. Also, another preliminary study with a selected set of 40 autosomal insertion-deletion (INDELs) polymorphisms7 in the same panel of continental populations obtained virtually the same result, showing to be sufficiently informative for an adequate characterization of human population structure at the global level. Thus, a relatively increased resolution can be obtained with informative INDELs selected on the basis of alleles with divergent frequencies among continental populations, commonly known as ancestry informative markers. Recently, these 40 INDELs were successfully used to discriminate among autochthonous American, European and African ancestry in the admixture analysis of populations from different regions of Brazil.3
On the pre-Columbian settlement of Peru
The archeological, paleontological and human skeletal remains indicate that the first hunter-gatherers appeared in Peru at about 12 000 years ago (late Pleistocene), inhabiting Andean areas around the Guitarrero Cave, Ancash8 and Ayacucho complex.9 Along the Pacific coast, some traces of the earliest human groups were dated to about 7000 years ago,10 which later may have originated some ancient civilizations like Caral (north of Lima). The earliest evidences revealed the formation of emergent societies around the Titicaca Lake, dating about 4000 years ago.11 By about 3000 years ago,12, 13 other civilizations appeared, Chavín in the north and Paracas in the south, and soon after 2 100 years ago, others emerged such as Moche (north coast), Nazca (central coast), Wari (south-central Andes), Chimú (north coast) and Chachapoyas (current Amazonas Department). Finally, the Tawantin Suyu Empire (1432–1532), which was dominated by the Incas, controlled the Andean regions of Peru, Ecuador and Bolivia, and part of Argentina, Chile and Colombia.5
During the Tawantin Suyu, the Quechua language expanded throughout most of the Andes by means of the Inca road (Qhapaq Ñan), leading also to a concurrent admixture between northern and southern subpopulations, including Inca and non-Inca groups.14, 15, 16, 17
The migration flows and admixture in the post-Columbian Peru
The first Europeans that arrived in Peru in the XVI century were mainly from Spain, who also brought some Africans as slaves.5 In 1849 started an immigration from China to all regions of Peru to work in plantations and guano exploitation,18 and since 1899 there were also some Japanese immigrants. In 1853, some German families immigrated with the goal to colonize the Amazon region, but large numbers of Europeans from Italy and other countries came in the beginning of the XX century,5 particularly during the first world war and beginning of the second (1918–1938). During the first decade of the XX century, an important internal migration flow happened in the Peruvian Amazon, when many urban and indigenous communities were displaced from their homelands to profit or run away from the rubber industry boom.5 However, since 1940 a large migration movement took place inside of Peru, mainly to Lima coming from Junín, Ayacucho, La Libertad, Ica, Lambayeque, Cajamarca, Piura and in a lesser degree from other places.5 This late XX century internal migration was mostly composed by rural and indigenous people that moved to urbanized cities, thus we would expect a large impact on the genomic ancestry of the inhabitants of large urban centers like Lima.
Our present study is focused on uncovering the population structure due to possibly different ancestral backgrounds in the human genomes of contemporaneous subpopulations of Peru, based on the detailed analyses of 40 INDELs.7 We calculated the genomic ancestry proportions of 25 subpopulations from all major regions of Peru and inferred the admixture level to ascertain pre- and post-Columbian genetic influences in a historical perspective. We identified a predominant Amerindian genomic ancestry in all regions of Peru and a pattern of non-indigenous admixture that is concordant with the known post-Columbian history of immigration.
Materials and methods
Subjects
The samples (blood or buccal swabs) were collected between 1998 and 2010 from unrelated volunteers, inhabitants from different regions of Peru. These participants were recruited with written informed consents approved by the USMP, Lima, Peru. For this study, 551 samples from 25 Peruvian localities were analyzed (Figure 1).
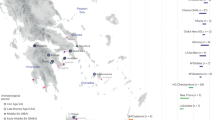
Map of Peru with sampling locations of 25 cities or districts from 13 Departments (right map): Loreto (LO), Ucayali (UC), Amazonas (AM), San Martin (SM), Cajamarca (CA), Ancash (AN), Junín (JU), Ayacucho (AY), Apurimac (AP), Arequipa (AR), Puno (PU), Lambayeque (LA) and Lima (LI). The sample composes of 122 individuals from the Amazon region (Andoas: And_LO=71, Iquitos: Iq_LO=8, Pucallpa: Puc_UC=10, Chachapoyas: Chp_AM=15, Lamas: SMla_SM=18); 355 individuals from the Andean region (Cajamarca: CA_CA=34, San Marcos: CAsm_CA=19, Ocopon: Oco_AN=11, Chogo: Ch_AN=14, Huarochiri: LIhr_LI=15, Huancayo: Hyo_JU=29, Ayacucho: AY_AY=31, Andahuaylas: Ahy_AP=19, Kaquiabamba: Kaq_AP=9, Cabanaconde: Cb_AR=20, Chivay: Cy_AR=25, Yanque: Yke_AR=10, Characato: Char_AR=8, Mollebaya: Mll_AR=8, Taquile: Taq_PU=23, Amantani: Amt_PU=31, Uros: Ur_PU=25, Anapia: Ap_PU=24) and 74 individuals from the Pacific coast (Lambayeque: LA_LA=31, Lima: LI_LI=43).
PCR and genotyping analysis
DNA was extracted and quantified according to the standard protocols19 in the laboratories from Peru (CGBM-USMP) and Brazil (LBEM-UFMG). The multiplex PCR reactions for 40 INDELs were performed following the previously standardized protocols.7, 20 Two microlitres of PCR products were added to 8 μl of Hi-Di formamide/GeneScan-500-LIZ solutions and subjected to capillary electrophoresis using the ABI 3130xl Genetic Analyzer (Life Technologies, Carlsbad, CA, USA). For allelic size scoring and visualization we used the GeneMapper ID v3.2 software (Life Technologies).
We used the same set of 40 INDELs (called MID#) that has been listed by Pena et al.,3 and the reference sequences (rs#) for each polymorphism are available in the NCBI Nucleotide Sequence Variation database (dbSNP) (http://www.ncbi.nlm.nih.gov/snp) and Marshfield Clinic Research Foundation (http://research.marshfieldclinic.org/genetics/home).
For comparisons we used published genotypic data7 for the 40 INDELs typed in 1064 individuals from 52 different worldwide populations (reference continental populations) of the HGDP-CEPH panel, which are distributed in seven geographical regions in all continents (http://www.cephb.fr/HGDP-CEPH-Panel).
Statistical analysis
General population genetic tests and analysis of molecular variance (AMOVA) were performed using ARLEQUIN v3.5.1.2 (Bern, Switzerland)21 and GENEPOP 4.0 (Montpelier, France)22 packages. To estimate the genomic proportions of American, Eurasian and African ancestry in Peruvian subpopulations, we applied Bayesian MCMC clustering analyses using the software STRUCTURE v2.3 (Chicago, IL, USA).23, 24 We have processed and visualized STRUCTURE outputs in the software STRUCTURE HARVESTER,25 DISTRUCT,26 CLUMPP,27 and R project (http://www.r-project.org/main.shtml) packages SimCo and ade4. We also used a weighted-least-square method implemented in the ADMIX program (http://www.genetica.fmed.edu.uy/software.htm) to estimate admixture. Additionally, we analyzed the INDEL genotypic data with different methods based on the Bayesian admixture analysis available in BAPS (Bayesian Analysis of Population Structure)28 and clustering based on the principal component analysis available in PCAGEN (http://www2.unil.ch/popgen/softwares/pcagen.htm).
Results
Ancestry proportions among peruvian subpopulations using structure
Population clustering and estimation of individual ancestry proportions were obtained with a model-based MCMC Bayesian algorithm implemented in the STRUCTURE software, which uses allelic frequencies for estimating a posterior distribution of the probability of membership to the predefined clusters (K), assuming that multiple loci are independent and are in Hardy-Weinberg equilibrium, as previously tested among 360 unrelated Brazilians.20
On the basis of historical records of Peru about the post-Columbian immigrations, we considered the admixture model and correlated allelic frequencies with parameters MCMC=200 000, burn-in=50 000 and MCMC=2 000 000, burn-in=100 000. The first analyses included only Peruvian samples, and they were performed without any information about the continental origin of Peruvians (PopFlag=0), and runs were made in 10 replicates with each K-value, ranging from K=1 to K=10. To identify substructuring among Peruvian subpopulations, the Q-membership (coefficient of ancestry proportion of membership to one of the K groups) results of STRUCTURE were processed by the Evanno method with STRUCTURE HARVESTER,25 which indicated ΔK=2 as the modal value and the best number of clusters fitting the data (Supplementary Figure 1). The second step of analyses (10 runs for each K, ranging from K=1 to K=10) were performed together with a selected set of data with 161 reference samples from Europe, 251 from East Asia and 105 from America, using the genetic data published in Bastos-Rodrigues et al.7 Reference continental samples were considered as known ‘parental’ populations (PopFlag=1), and Peruvian samples were labeled as unknown origin (PopFlag=0) in STRUCTURE input format. These two independent STRUCTURE analyses used the same parameters and showed the same partition of Peruvian subpopulations in two clusters (ΔK=2) by an Evanno method in STRUCTURE HARVESTER. The scores of averaged coefficients of Q-membership for partition K=2 generated by CLUMPP27 were plotted by a method of correspondence analysis implemented in ade4 package for visualization of the asymmetric clustering of Peruvian subpopulations (showed only without reference populations), indicating that there is a population substructure in Peru (Supplementary Figure 2).
The SimCo statistical package was used for comparisons of 10 STRUCTURE runs defining the clustering solution (ΔK=2) by similarity coefficients (SimCoef). Thus, in a population level, without reference populations (only 25 Peruvian subpopulations), the mean SimCoef was 0.978 (s.e.=0.002). When including selected reference populations (25 Peruvian and 3 reference populations), the mean SimCoef was 0.995 (s.e.=0.0004). In both cases, the SimCoef values indicate that the clustering performed (ΔK=2) using STRUCTURE was highly similar among the 10 compared runs (98% and 99% respectively). Using the same procedure, we determined the SimCoef value in an individual level. On the other hand, the average Q-membership values were obtained by using CLUMPP, and graphics were drawn with DISTRUCT to visualize the clustering pattern of individuals and populations. The results of analysis including the selected reference populations from Europe, East Asia and America, and using partition K=2, are displayed in Figure 2, where a clear admixture pattern can be also observed among Peruvian subpopulations.
Clustering result (K=2) obtained using STRUCTURE and plotted with DISTRUCT on 25 Peruvian subpopulations (see legend of Figure 1), using reference samples of the HGDP-CEPH. The populations are represented in delimited bar segments and the individuals by colored vertical lines showing their ancestry membership proportions. The number codes for subpopulations are: 1=AY, 2=Hyo, 3=Cb, 4=Cy, 5=Yke, 6=Char, 7=Mll, 8=Oco, 9=Ch, 10=CA, 11=CAsm, 12=Ahy, 13=Kaq, 14=LIhr, 15=Ur, 16=Ap, 17=Amt, 18=Taq, 19=And, 20=Iq, 21=Puc, 22=Chp, 23=SMla, 24=LA, 25=LI . A full color version of this figure is available at the Journal of Human Genetics journal online.
We also used data of 1064 individuals from 52 reference HGDP-CEPH populations divided a priori in four regions (Africa, Eurasia (Europe, Middle East, Central Asia, East Asia), Oceania and America) to estimate the genomic ancestry proportions of Peruvians. The STRUCTURE results (only from K=3 to K=6) are shown in Supplementary Figure 3. For illustrative purposes, we also included a 3D plot with partition K=3 (Supplementary Figure 4), where the Peruvian subpopulations are distributed in a gradient pattern, dependent on the admixture level. The best overall partition obtained was K=5 (Figure 3), which fits to five geographic regions (East Asia, Eurasia – Europe, Middle East, Central Asia – Oceania, Africa and America), although Eurasia appears not to be regionally structured and highly intermixed with Oceania. Indeed, clustering in more than two population conglomerates is dependent on the level of relatedness, for example, the partitions K=3 or K=4 seem to be as valid as K=5, due often to the observation that Eurasians and Oceanians show similar genetic profiles in the admixture model (see Supplementary Figure 3). A likely explanation for this result is due to the fact that these 40 INDELs were initially selected as ancestry informative markers to discriminate among native Africans, Europeans and Americans, but not Oceanians.3, 7 However, when we used only Europeans (n=161, from eight subpopulations) and Native Americans (n=108, from five subpopulations) as ‘parental’ populations, we obtained very similar values of admixture proportions (Supplementary Table 1) using the weighted-least-square method implemented in the ADMIX program, which is based on gene identity probability.29
DISTRUCT barplot of estimated Q-coefficients for 52 reference populations of HGDP-CEPH panel and 25 Peruvian subpopulations (see number codes in Figure 2 legend) calculated using STRUCTURE with partition K=5. The populations are represented in delimited bar segments and the individuals by colored vertical lines showing their ancestry membership proportions of the individuals. The bottom graphic is a zoom view of the 25 Peruvian subpopulations data shown above.
The analyses in STRUCTURE have also generated equivalent results with a model based on ‘no-admixture’ and ‘allele frequencies not-correlated’, using a priori K=5 (Supplementary Figure 5). It means that independently of assumed population model (admixture or not admixture), the clustering of samples by the Bayesian MCMC algorithm revealed a similar admixture profile of individuals or populations. However, the ‘admixture’ model explains better the results according to the scenario described by the known history of pre- and post-Columbian colonization of Peru.
The STRUCTURE results concerning the averaged proportions of membership (Q) are shown in Table 1. They were obtained in relation to the predefined reference populations of the HGDP-CEPH panel, and partitioned in K=2 (America and not-America) and in K=5 (Africa, Europe-Middle East (ME)-Central Asia (CA), East Asia, Oceania and America). In general, Peruvian subpopulations present a high proportion of autochthonous American ancestry (Q=0.538–0.965) and heterogeneous levels of non-autochthonous admixture. The values of Eurasian (Europe, ME, CA) ancestry proportions are displayed in Figure 4. In Table 1, the localities of San Marcos, Characato, Cajamarca, Chogo, Lambayeque and Lima presented the highest average proportions of membership (31.2%, 24.4%, 20.5%, 14.6%, 14.5% and 14.3%, respectively) to the Eurasian region (Europe, ME, CA). Intermediate levels of Eurasian ancestry were associated with the localities of Lamas, Ayacucho and Huancayo (8.7%, 8.1% and 6.1%, respectively). These Peruvian localities associated with a partial genomic ancestry derived from Europe, Middle East and Central Asia also present small genomic ancestry proportions from Africa (<3.4%). It is interesting to note that some ancestry proportions (Table 1) are more related to the East Asia region in Chachapoyas (8.2%), Mollebaya (8%) and Iquitos (6%), but Pucallpa presents 5.2% East Asian ancestry together with 9% from Oceania and 8% from Europe, Middle East, Central Asia. A large East Asian contribution is likely associated with recent post-Columbian migration into these areas, as indicated by recent historical reports.18 Because our sampling was anonymous and collected without genealogical information, East Asian ancestry for individual samples cannot be verified.
Bayesian clustering of peruvian subpopulations using BAPS
To compare with the results of the STRUCTURE clustering, we also performed a genetic admixture analysis using the BAPS28 with the 40 INDELs data of 25 Peruvian subpopulations at the individual and group levels. We used a priori upper bound value K=25, and 10 replicate runs of the stochastic estimation algorithm model using 10 000 iterations, which yielded the optimal posterior partition of the number of clusters of K=2 (Cluster 1: Cabanaconde, Chivay, Yanque, Mollebaya, Ocopon, Andahuaylas, Kaquiabamba, Huarochiri, Uros, Anapia, Amantani, Taquile, Andoas, Iquitos and Cluster 2: Ayacucho, Huancayo, Characato, Chogo, Cajamarca, San Marcos, Pucallpa, Chachapoyas, Lamas, Lambayeque, Lima), with a Log (marginal likelihood)=−24 471.48 and probability=0.986.
This independent analysis converged into the same partition for Peruvian subpopulations indicated using the STRUCTURE software (ΔK=2). Furthermore, a BAPS clustering approach using the HGDP-CEPH reference samples with a priori K=4 also showed a similar admixture pattern (Supplementary Figure 6) to the one obtained with STRUCTURE.
Clustering of peruvian subpopulations by principal components analysis
We performed a principal components analysis on 40 INDELs data to correlate allele frequencies and genotypes among all sampled individuals or populations. The computer package PCAGEN was used to estimate the percentage inertia of each PC axis and its associated P-value by 10 000 randomizations of genotypes. Next, two dimensional scatter plots of the first two principal components were produced. The analysis showed that the eigen values for the first two components (PC1=23.8%; PC2=11.3%) were highly significant (P-value=0), and also the PC1-Fst’s (1%) show a larger differentiation among populations than the PC2-Fst’s (0.5%). The global observed Fst was 4.4%, and the total heterozygosity was 37.3%.
In Supplementary Figure 7, the populations from CAsm (San Marcos), CA (Cajamarca), Char (Characato), LA (Lambayeque), LI (Lima) and Ch (Chogo) were distantly placed in comparison with populations from Taq (Taquile), Amt (Amantani), Ap (Anapia), Ur (Uros) and Yke (Yanque).
Genetic diversity and interpopulation relationships
Several analyses were performed to characterize genetic diversity within and among Peruvian subpopulations from 25 localities. The AMOVA and Fst analyses done in ARLEQUIN followed three hierarchical groupings: (i) the localities were distributed in three different geographical regions (Amazon, Andes and Coast). In this case, the difference among groups was 0.33%, among populations within groups 2.18%. (ii) The localities from each region were analyzed independently. In this situation, the difference among subpopulations from the Amazon (n=122) was 0.99% (P-value =0.88), from the Andes (n=355) was 2.74% (P-value=0), and from the Coast (n=74) was 0.37% (P-value=0.86). (iii) The localities were considered belonging to a single macro region (Peru), without internal geographical division. Despite the difference among populations (Fst) was also relatively low (2.37%), it was significant (P-value=0). In the three AMOVA approaches, the results indicate little genetic differentiation among populations (< 2.74%), and most of differences were detected between individuals (>96%) (Supplementary Table 2). This low but significant interpopulation differentiation agrees with the known history of pre- and post-Columbian settlement of the country, composed by recurrent interchange of migrants from north to south, and between Coast, Andes and Amazon. The exact test of population differentiation was performed using the program GENEPOP with 100 000 permutations across all 40 INDELs, and it showed highly significant P-values (result not shown), particularly between subpopulations from the Titicaca Lake (Taquile, Amantani, Anapia, Uros) and subpopulations from San Marcos, Cajamarca, Characato, Lambayeque and Lima. Furthermore, MDS graphics generated using GenAlEx30 from pairwise Fst distances of Peruvian subpopulations only (figure not shown), or including also reference populations from continents (Figure 5), reveal a clustering pattern that is congruent with the analyses performed using STRUCTURE, BAPS and principal components analysis.
MDS plot (pairwise Fst distances) of 18 selected reference populations (n=386) of the HGDP-CEPH panel (Africa (n=55), Oceania (n=17), Middle East (n=29), Europe (n=43), Central Asia (n=25), East Asia (n=109), America (n=108)) and 25 Peruvian subpopulations (see number codes Figure 2 legend). A full color version of this figure is available at the Journal of Human Genetics journal online.
Among the 25 Peruvian subpopulations, the average observed heterozygosity (Ho=36%) for all 40 loci was similar to the expected heterozygosity (He=37%). Considering all subpopulations as one population group, the values were the same (Ho=36%; He=36%) and similar values were obtained using the PCAGEN package (Htotal=37.3%). However, the average heterozygosity over all loci among the 25 subpopulations showed a direct relationship with the admixture levels of subpopulations (Supplementary Figure 8). The highest expected heterozygosity was identified in Characato, followed by San Marcos, Cajamarca, Lambayeque, Lima and Chogo, and the lowest heterozygosity was found in Taquile, Anapia, Amantani, Yanque and Uros. Indeed, a correlation analysis (Supplementary Figure 8) between expected heterozygosities (He) and the gradient of admixture degree observed among Peruvian subpopulations in relation to non-autochthonous Americans (not-America) (Table 1) has shown a high and significant Pearson’s correlation index (r=0.975; P-value=2.20E−16).
Discussion
Recently, different sets of AIMs were used to estimate ancestry proportions of Latin American populations in comparison with ‘parental’ or reference continental populations.2, 3, 31 The AIM set of 40 INDELs7 was used in this study to estimate the impact of pre- and post-Columbian colonization in the current Peruvian population.
Previously, the study of these 40 INDELs among Brazilian subpopulations from the political regions North, Northeast, Southeast and South (n=934) revealed that the level of admixture proportions is relatively uniform, with a predominant European ancestry (ranging 60.6–77.7%) in all regions.3 In contrast, our results on Peruvians using the same INDELs identified a predominant autochthonous American ancestry (ranging 53.8–96.5%). Those discrepant results agree with the known histories of colonization of Brazil and Peru, which indicate a much larger effective population size of indigenous Peruvians at the time of contact and during European colonization, particularly along the Andes.32
Some previous studies have indicated some level of structuration between regions of Peru. A study of dermatoglyphic patterns33 detected similar features between northern and central Peruvian subpopulations, but they were both differentiated from highlanders of the Puno region (Titicaca Lake). In a recent study using STR markers,34 Peruvians were suggested to be clustered in three main subgroups according to their geographical locations (north, central and south of Peru) and reported about 30% of admixture with non-autochthonous populations, a similar overall value compared with our results using K=2 (Table 1).
It is worth noting that for the inference of the genetic relationships of current worldwide populations, most studies assume that ‘native’ continental populations are geographically structured and not-admixed, which could result in a bias in recent admixture analyses. For example, the ‘native’ subpopulations of Middle East, Europe and north of Africa appear to be relatively admixed (Figure 3), as well as other regions of the world, which is fairly supported by a recent study using autosomal single-nucleotide polymorphisms arrays.35 In our STRUCTURE results, using an admixture model and partition K=2, the populations from Europe and East Asia were clustered in one macrogroup, whereas autochthonous Americans were found in another cluster (Figure 2). This evidence is in close agreement with the results obtained by Wang et al.,36 but in contrast with some previous studies where the East Asian populations were clustered with Native Americans.6, 7, 37 Nevertheless, our clustering results with partitions K=3 to K=6 including all 52 reference HGDP-CEPH subpopulations (Supplementary Figure 3) are similar to those obtained by all previous studies. In any case, the Q-membership proportions for each individual/population should be seen with caution, as ‘ad hoc approximations’, as they may change depending on number and type of markers, number of samples, the reference populations used and also the demographic history or degree of inter-population differentiation in the studied area.38
The population structure analyses of 25 Peruvian locations, using partition K=5, showed that in San Marcos, Cajamarca, Characato, Lima, Lambayeque, Chogo, Lamas, Huancayo and Ayacucho there is a high level of post-Columbian admixture (mainly with Europeans).5 It is in close agreement with the European colonization history of Peru. Also, we identified a significant admixture with East Asians in the localities of Mollebaya (Andes), Pucallpa, Chachapoyas and Iquitos (Amazon region), which is consistent with the historical records that report an immigration wave of Chinese, who occupied several parts of Peru since 1870, particularly the sampled Amazon locations.18
Among the inhabitants of the Titicaca Lake region (Taquile, Amantani, Uros, and Anapia) and Yanque (from Colca Canyon in Arequipa), there was no significant admixture with non-autochthonous Americans (in K=5). This could be explained by the relative isolation of this harsh Andean area, but could be also related with local mating practices and historical peculiarities. For example, in the Taquile Island there is an endogamous mating practice that excludes foreigners marrying local inhabitants to reside in the community (unpublished observations). Besides, the association between subpopulations from Titicaca Lake and the Yanque locality in Arequipa can be further supported by historical records before the Inca Empire, when the same Amerindian groups were occupying this southwestern region of Peru.39
The AMOVA results on Peruvians showed a low, but slightly higher level of genetic differentiation among Andean subpopulations when compared with Amazonian or Coastal. Indeed, previous studies of South American indigenous subpopulations14, 15, 16, 17 through the use of autosomal and uniparental markers (excluding admixture) have suggested a lower degree of genetic differentiation among the Andean communities than among the eastern Amerindian communities (Amazon and Brazilian plateau). Part of this pre-Columbian homogenization in the Andes was suggested to be due to the Mit’a system of forced movement of populations, which was a labor draft system used by the Inca Empire.32 These differences observed between genetic data on non-admixed indigenous populations, and INDEL markers on Peruvian subpopulations clearly support the impact of post-Columbian admixture in the population dynamics owing to the promotion of gene flow. Indeed, during the Spanish colonization a new reformulated Mit’a system was empowered by encomenderos,32 leading to relocation of many populations in the area as forced labor, which was especially documented for the colonial establishment of the silver mines in Potosí (in present-day Bolivia).32
Peruvian subpopulations from Taquile, Anapia and Amantani presented lower heterozygosity (<31.8%) and also an insignificant admixture, in contrast to the high heterozygosity and admixture observed in Cajamarca and Characato (Arequipa). The two latter populations form a compact cluster in MDS analyses with populations from San Marcos, Lima, Lambayeque and Chogo, suggesting a higher impact of post-Columbian admixture events, which is coherent with historical records.5, 32 Subpopulations from Ayacucho, Huancayo, Lamas, Chachapoyas and Pucallpa appeared closely related to each other, which could be explained by their similar level of admixture with Eurasians as well as by their shared autochthonous ancestry.40 Other associations found in the bidimensional analyses (MDS and principal components analysis) between subpopulations may have alternative explanations. Taquile and Amantani subpopulations appear very closely related in the MDS graphic, which agrees with their proximity (they are neighbors living in close islands in the Titicaca Lake) and also fit to local reports about their common origin, from Capachica peninsula (www.ogdpuno.org). Besides, these two subpopulations are near to Anapia (Aymara-speaking subpopulation), another island of the Titicaca Lake, but are located in the frontier between Peru and Bolivia. However, the Uros subpopulation (inhabitants of the floating islands of Titicaca) was shown to be more related with other far located subpopulations (Andoas, Chivay, and Cabanaconde), even though previous mitochondrial DNA analyses41 have shown a shared ancestry with their neighbors from Titicaca Lake region. The subpopulations from Cabanaconde and Chivay appear closely related in the MDS graphic, and both are located in the Colca Canyon as well as Yanque, but this latter subpopulation is separated from those by the observed pattern of non-admixture. Interestingly, Anapia, Taquile, Amantani and Yanque subpopulations present a closer affinity to an autochthonous group from the Brazilian Amazon (Karitiana), which also presents low levels of heterozygosity and admixture, thus it is likely due to their shared Amerindian ancestry. It is also interesting to observe that Peruvian Amazon subpopulations (Pucallpa, Iquitos, and Chachapoyas) appear closely related to other autochthonous American groups from the Amazon, such as Surui (Brazil) and Piapoco (Colombia). Although there is a long geographic distance between Mollebaya (Arequipa) and Ocopon (Ancash), they appear to be genetically close to each other in the bidimensional analyses. Their similarity is probably due to an equivalent admixture proportion estimated using STRUCTURE, like it also happens between Characato (south of Peru) and Cajamarca (north of Peru).
In summary, when clustering analyses are done with K=5 (coincident with five continental regions), the total genomic ancestry proportions in Peruvians are 83% for America and 17% are non-autochthonous, mainly from Europe. Using a partition in two main subgroups (K=2), the total average of non-autochthonous proportion among Peruvian genomes rises to about 20%, mainly due to European admixture, and autochthonous genomic heritage in Peru is about 80%, corresponding to a very high prevalence of pre-Columbian genes in the current population. These results indicate a clear effect of post-Columbian admixture in the population structure of Peru, portraying a gradient of autochthonous/non-autochthonous genomic background due to different degrees of admixture and shared ancestry among Peruvian subpopulations.
References
Avena, S., Via, M., Ziv, E., Pérez-Stable, E. J., Gignoux, C. R., Dejean, C. et al. Heterogeneity in genetic admixture across different regions of Argentina. PLoS One 7, e34695 (2012).
Galanter, J. M., Fernandez-Lopez, J. C., Gignoux, C. R., Barnholtz-Sloan, J., Fernandez-Rozadilla, C., Via, M. et al. Development of a panel of genome-wide ancestry informative markers to study admixture throughout the Americas. PLoS Genet. 8, e1002554 (2012).
Pena, S. D. J., Di Pietro, G., Fuchshuber-Moraes, M., Genro, J. P., Hutz, M. H., Kehdy, F. D. S. G. et al. The genomic ancestry of individuals from different geographical regions of Brazil is more uniform than expected. PLoS One 6, e17063 (2011).
Wang, S., Ray, N., Rojas, W., Parra, M. V., Bedoya, G., Gallo, C. et al. Geographic patterns of genome admixture in Latin American Mestizos. PLoS Genet. 4, e1000037 (2008).
Lexus Edit. Gran Enciclopedia del Perú (Barcelona, España, 1998).
Li, J. Z., Absher, D. M., Tang, H., Southwick, A. M., Casto, A. M., Ramachandran, S. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
Bastos-Rodrigues, L., Pimenta, J. R. & Pena, S. D. J. The genetic structure of human populations studied through short insertion-deletion polymorphisms. Ann. Hum. Genet. 70, 658–665 (2006).
Lynch, T. A. & Kennedy, K. A. R. Early human cultural and skeletal remains from Guitarrero Cave, Northern Peru. Science 169, 1307–1309 (1970).
Macneish, R. S., Berger, R. & Protscha, R. Megafauna and man from ayacucho, highland peru.from Ayacucho, highland Peru. Science 168, 975–977 (1970).
Leon, C. E. Origenes humanos en los Andes del Per ú Ed. USMP, (Lima: Perú, 2007).
Aldenderfer, M., Craig, N. M., Speakman, R. J. & Popelka-Filcoff, R. Four-thousand-year-old gold artifacts from the Lake Titicaca basin, southern Peru. Proc. Natl Acad. Sci. USA 105, 5002–5005 (2008).
Caceres, M. J. Las culturas pre-hispanicas del Perú (3rd Edn CONCYTEC (Lima: Perú, 1989).
Lumbreras, L. Los origenes de la civilizacion en el Perú (5ta Ed. Milla Batres (Lima: Perú, 1981).
Cavalli-Sforza, L. L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes, (Princeton University Press: NJ, USA, 1994).
Luiselli, D., Simoni, L., Tarazona-Santos, E., Pastor, S. & Pettener, D. Genetic structure of Quechua-speakers of the Central Andes and geographic patterns of gene frequencies in South Amerindian populations. Am. J. Phys. Anthropol. 113, 5–17 (2000).
Rodriguez-Delfin, L. A., Rubin-de-Celis, V. E. & Zago, M. A. Genetic diversity in an Andean population from Peru and regional migration patterns of Amerindians in South America: data from Y chromosome and mitochondrial DNA. Hum. Hered. 51, 97–106 (2001).
Tarazona-Santos, E., Carvalho-Silva, D. R., Pettener, D., Luiselli, D., De Stefano, G. F., Labarga, C. M. et al. Genetic differentiation in South Amerindians is related to environmental and cultural diversity: evidence from the Y chromosome. Am. J. Hum. Genet. 68, 1485–1496 (2001).
Herrera, I. L. Los Inmigrantes Chinos en la Amazonia Peruana. Bull. Inst. Fr. Et. And. XV N3-4, 49–60 (1986).
Jota, M. S., Lacerda, D. R., Sandoval, J. R., Vieira, P. P. R., Santos-Lopes, S. S., Bisso-Machado, R. et al. A new subhaplogroup of native American Y-Chromosomes from the Andes. Am. J. Phys. Anthropol. 146, 553–559 (2011).
Pimenta, J. R. & Pena, S. D. J. Efficient human paternity testing with a panel of 40 short insertion-deletion polymorphisms. Genet. Mol. Res. 9, 601–607 (2010).
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Rousset, F. GENEPOP’007: a complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106 (2008).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Earl, D. A. & vonHoldt, B. M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Cons. Genet. Resour. 4, 359–361 (2012).
Rosenberg, N. A. Distruct: a program for the graphical display of population structure. Mol. Ecol. 4, 137–138 (2003).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007).
Corander, J., Marttinen, P., Sirén, J. & Tang, J. Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinformatics 9, 539 (2008).
Chakraborty, R., Kamboh, M. I. & Nwankwo.M., F. e. r. r. e. l. l. R. E. Caucasian genes in American blacks: new data. Am. J. Hum. Genet. 50, 145–155 (1992).
Peakall, R. & Smouse, P. E. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. 6, 288–295 (2006).
Santos, N. P. C., Ribeiro-Rodrigues, E. M., Ribeiro-Dos-Santos, A. K. C., Pereira, R., Gusmão, L., Amorim, A. et al. Assessing individual interethnic admixture and population substructure using a 48-insertion-deletion (INSEL) ancestry-informative marker (AIM) panel. Hum. Mut. 31, 184–190 (2010).
Hunefeldt, C. A brief history of Peru, (Lexington Associates: NY, USA, 2004).
Ramirez, B. O., Del Valle, M. L. & Arzola, G. N. Dermatoglyphics of a high altitude Peruvian population and interpopulation comparisons. High Alt. Med. Biol. 2, 31–40 (2001).
Iannacone, G. C., Parra, R., Bermejo, M., Rojas, Y., Valencia, C. & Portugues, L. Peruvian genetic structure and their impact in the identification of Andean missing persons: A perspective from Ayacucho. Forensic Sci. Int.: Genet. Suppl. Series 3, e291–e292 (2011).
Reich, D., Patterson, N., Campbell, D., Tandon, A., Mazieres, S., Ray, N. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012).
Wang, S., Lewis, C. M., Jakobsson, M., Ramachandran, S., Ray, N., Bedoya, G. et al. Genetic variation and population structure in native Americans. PLoS Genet. 3, e185 (2007).
Rosenberg, N. A., Pritchard, J. K., Weber, J. L., Cann, H. M., Kidd, K. K., Zhivotovsky, L. A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002).
Latch, E. K., Dharmarajan, G., Glaubitz, J. C. & Rhodes, O. E. Relative performance of Bayesian clustering software for inferring population substructure and individual assignment at low levels of population differentiation. Conserv. Genet. 7, 295–302 (2006).
Bouysse-Cassagne, T. Poblaciones humanas antiguas y actuales, (ORSTOM HISBL: Bolivia, 1991) 481–498.
Pardo, C. M., Doherty, V. J. & Sangama, S. I. Los Kechuas Lamistas y la Educacion Bilingue Intercultural: Historia y Razon de un compromiso Ed. PT S. A. C., Tarapoto S (Martin: Perú, 2001).
Sandoval, J., Delgado, B., Rivas, L., Bonilla, B., Nugent, D. y. & Fujita, R. Variantes del ADNmt en isleños del lago Titicaca: máxima frecuencia del haplotipo B1 y evidencia de efecto fundador. Rev. Peru. Biol. 11, 161–168 (2004).
Acknowledgements
We thank all volunteers who donated their samples to this project, M.Sc. Jaime Descailleaux and Margarita Velasquez (UNMSM, Lima, Peru), Cesar Ñique (USAT, Lambayeque, Peru) for providing storage of some samples, Dr. Daniela R. Lacerda (LBEM, Brazil) and Heloisa B. Pena (GENE-MG, Brazil) for technical support, PEC-PG CAPES/Brazil for the PhD scholarship to JRS, and FAPEMIG and CNPq from Brazil for financial support to the laboratory work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Sandoval, J., Salazar-Granara, A., Acosta, O. et al. Tracing the genomic ancestry of Peruvians reveals a major legacy of pre-Columbian ancestors. J Hum Genet 58, 627–634 (2013). https://doi.org/10.1038/jhg.2013.73
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.73
Keywords
This article is cited by
-
Allelic frequencies with 23 autosomic STRS in the Aymara population of Peru
International Journal of Legal Medicine (2021)
-
ATXN10 Microsatellite Distribution in a Peruvian Amerindian Population
The Cerebellum (2019)
-
Admixture, Genetics and Complex Diseases in Latin Americans and US Hispanics
Current Genetic Medicine Reports (2018)
-
Genetic ancestry of families of putative Inka descent
Molecular Genetics and Genomics (2018)
-
Latin Americans show wide-spread Converso ancestry and imprint of local Native ancestry on physical appearance
Nature Communications (2018)