Abstract
The most important modality of treatment in the two-thirds of patients with an estrogen receptor (ER)-positive early breast cancer is endocrine therapy. In postmenopausal women, options include the selective ER modulators (SERMs), tamoxifen and raloxifene, and the ‘third-generation’ aromatase inhibitors (AIs), anastrozole, exemestane and letrozole. Under the auspices of the National Institutes of Health Global Alliance for Pharmacogenomics, Japan, the Mayo Clinic Pharmacogenomics Research Network Center and the RIKEN Center for Genomic Medicine have worked collaboratively to perform genome-wide association studies (GWAS) in women treated with both SERMs and AIs. On the basis of the results of the GWAS, scientists at the Mayo Clinic have proceeded with functional genomic laboratory studies. As will be seen in this review, this has led to new knowledge relating to endocrine biology that has provided a clear focus for further research to move toward truly personalized medicine for women with breast cancer.
Similar content being viewed by others
Introduction
Breast cancer is the most common form of cancer in women both in the United States1 and Japan.2 Endocrine therapy is the most important modality in the two-thirds of patients with an estrogen receptor (ER)-positive early breast cancer. There are two classes of drugs that are the mainstay of endocrine therapy in postmenopausal women. These are the selective ER modulators (SERMs), tamoxifen and raloxifene, and the ‘third-generation’ aromatase inhibitors (AIs), anastrozole, exemestane and letrozole. A recent update of the worldwide experience3 revealed tamoxifen to have substantial value in reducing the risk of disease recurrence. Numerous clinical trials in the adjuvant setting have also been performed utilizing the third-generation AIs, anastrozole, exemestane and letrozole versus tamoxifen,4 and a recent meta-analysis revealed that the AIs were superior in that they produced significantly lower recurrence rates than tamoxifen, either as initial monotherapy or after 2 to 3 years of tamoxifen5. A recent American Society of Clinical Oncology practice guideline recommended AI use at some point during adjuvant endocrine therapy.6
SERMs have also been found to be of value in women at high risk of developing breast cancer7 and the US Food and Drug Administration (FDA) has approved both tamoxifen and raloxifene for treatment of these women. The basis for the FDA approval were two studies conducted by the National Surgical Adjuvant Breast and Bowel Project (NSABP) that showed 5 years of treatment with either tamoxifen or raloxifene can reduce the occurrence of breast cancer in these high-risk women by one-half. These large and influential breast cancer prevention trials were the double-blind, placebo-controlled NSABP P-1 trial of tamoxifen8, and the double-blind NSABP P-2 trial that compared raloxifene with tamoxifen.9, 10 Combined, these two studies involved over 33 000 women, which constituted about 59% of the world’s experience with patients entered on prospective trials of tamoxifen or raloxifene for breast cancer prevention in high-risk women.
It is because of the high level of importance of endocrine therapy to women with breast cancer and the marked variability that is observed clinically that our group at Mayo Clinic has focused on the AIs and SERMs. That is, clinical observations reveal a marked variability between patients in terms of response to treatment. Two identical patients can have markedly different outcomes, with one patient never having any disease recurrence whereas the other will have a recurrence and progression of disease. In addition, there is marked variability in adverse events (AEs). A striking example is the variability seen in terms of the musculoskeletal AEs that can occur with AI therapy. Some patients have absolutely no musculoskeletal symptoms whereas others can become disabled from them. Although some AEs, such as musculoskeletal and vasomotor AEs, are not in themselves life threatening, they represent a potential serious threat to a patient’s outcome because of an adverse effect on compliance. Likely related to the variability in patient outcomes and AEs is the variability we have identified with the AI anastrozole in terms of its metabolism and pharmacodynamic effect.11 That is, in a study of 191 women with early-stage breast cancer, we obtained blood for DNA extraction and plasma for the determination of estrone, estradiol, estrone conjugates, androstenedione and testosterone before and after therapy with anastrozole. In addition, after achievement of steady-state levels of anastrozole, we determined plasma anastrozole and anastrozole metabolite concentrations. There were large inter-individual variations in pretreatment and post drug plasma hormone levels, as well as plasma anastrozole and anastrozole metabolite concentrations. This large degree of variability has potentially important implications with regard to efficacy and AEs with anastrozole and suggests that the approved anastrozole dose of 1 mg per day may not be optimal for all patients.
In this review, the current results of our pharmacogenomic studies in patients receiving AIs or SERMs will be reviewed. As will be seen, the approach taken is the performance of a genome-wide association study (GWAS) as the initial step in a process that goes beyond the identification of associations to study the relationship of the single-nucleotide polymorphisms (SNPs) to genes and the relationships of these SNPs and genes to the drug effect and the phenotype under study (see Figure 1). This approach was considered a ‘new pharmacogenomic paradigm’ in an editorial12 that accompanied the manuscript reporting our initial GWAS and functional genomics study13 that will be discussed subsequently.
Pharmacogenomics of AIs in the adjuvant setting
MA.27 is the largest adjuvant endocrine therapy trial conducted to date that has exclusively studied AIs and, importantly, prospectively collected blood for DNA extraction and patient consent for its use in genetic studies. This study will be briefly described because it is the source of patients for multiple GWAS that have been or are currently underway with different phenotypes that will be discussed. This trial was conducted under the auspices of the North America Breast Cancer Groups and coordinated by the NCIC Clinical Trials Group in Canada. The results of this trial have recently been published14. Briefly, postmenopausal women who had adequately excised, histologically or cytologically confirmed primary breast cancer that was hormone receptor positive were eligible for this trial. Women were randomized to an AI, either the steroidal AI exemestane or the non-steroidal AI anastrozole. A total of 7576 women were randomized on MA.27 between 2003 and 2008. The primary end point was event-free survival, defined as the time from randomization to the time of documented locoregional or distant recurrence, new primary breast cancer, or death from any cause. Secondary end points included overall survival, time to distant recurrence, incidence of contralateral breast cancer, and long-term clinical and laboratory safety.
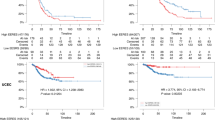
The final results from this study14 revealed no difference in efficacy between anastrozole and exemestane. Specifically, at median follow-up of 4.1 years, 4-year event-free survival was 91.0% for exemestane and 91.2% for anastrozole (stratified hazard ratio 1.02, 95% confidence interval 0.87–1.18, P=0.85). Overall, distant disease-free survival and disease-specific survival were similar for anastrozole and exemestane.
GWAS with phenotype of musculoskeletal AEs
It is well established that a substantial proportion of women are suboptimally adherent to anastrozole therapy15, and that about half of patients treated with AIs have joint-related complaints,16, 17 which likely contributes to decreased compliance. A review of the patients who discontinued anastrozole on MA.27 revealed that the major reason for discontinuation was musculoskeletal AEs. We hypothesized that the variability seen with respect to these musculoskeletal complaints in women treated with AIs could be related to genetic variability of the patients, and we proceeded to perform a GWAS with the goal of identifying SNPs associated with this variability. A nested, matched, case–control design was used, with matching on the following factors: age, treatment with exemestane or anastrozole, presence or absence of prior adjuvant chemotherapy, whether or not the patient had received celecoxib (the first 1662 patients entered had been randomized to celecoxib or placebo but this was stopped after reports of cardiotoxicity with celecoxib) and time on study. To minimize population stratification, the GWAS was restricted to white patients, as 94% of the patient’s entered on MA.27 were self-reported to be white. Additional covariates evaluated were body mass index, presence or absence of bisphosphonate use, whether or not the patient had had a fracture in the previous decade, baseline performance status (using Eastern Cooperative Oncology Group criteria), whether the patient had received prior hormone replacement therapy, prior adjuvant radiotherapy and prior taxane therapy.
To be classified as a case, a patient must have had one of the following six musculoskeletal complaints: joint pain, muscle pain, bone pain, arthritis, diminished joint function or other musculoskeletal problems. Cases were required to either have at least grade 3 toxicity, which is defined as severe pain and limiting self-care activities of daily living, according to the National Cancer Institute’s Common Terminology Criteria for Adverse Events v3.0, or go off protocol treatment for any grade of musculoskeletal complaint within the first 2 years of therapy with the AI. Controls were those women who did not experience any of the musculoskeletal complaints, were followed for at least 2 years and had at least 6 months longer follow-up than a case to which they were matched.
The genotyping for this study was performed at the RIKEN Center for Genomic Medicine and was of outstanding quality. Only 1.9% of the SNPs were considered failures and, after exclusion of SNPs with a minor allele frequency of <0.01 because of limited power for association analyses and exclusion of 82 SNPs with P-value <1E−06, 551,395 SNPs were used in the GWAS.
The GWAS identified three SNPs on chromosome 14 with P-values <1E−06, and an additional SNP with a low P-value was identified by imputation using HapMap 2 as a reference and then verified through additional genotyping. This imputed SNP (rs11849538) was associated with the musculoskeletal AEs with an odds ratio of 2.21 (MAF cases/controls: 0.172/0.091; P=6.67E−07). Upon identification of the SNPs, their location on chromosome 14 was examined and they were found to be near the T-cell leukemia 1A (TCL1A) gene.
Through our GWAS, we had observed promising—but not genome-wide significant—associations, but it was because of the availability of a panel of well-characterized, genomic data rich, lymphoblastoid cell lines (LCLs) that we were able to explore hypotheses relating to these findings. Our panel of LCLs, developed and characterized by Liewei Wang, MD, PhD at Mayo, has dense SNP and mRNA expression data that has been used for generating and testing pharmacogenomic hypotheses.18, 19 Using this LCL model system, we demonstrated that TCL1A was variably expressed in these cell lines. The TRANSFAC database suggested that the rs11849538 SNP would create an estrogen response element (ERE), and this was demonstrated to be the case through a chromatin immunoprecipitation (ChIP) assay in which LCLs with known genotypes for the rs11849538 SNP were transfected with ERα. As the effect of AIs is to perturb the level of estrogens, we determined whether TCL1A expression was estrogen inducible by utilizing U2OS cells stably transfected with either ERα or ERβ and found this to be the case with substantial, six- to eight-fold, increases in TCL1A expression.
The next steps were to determine the effect of different genotypes of the four SNPs on the estrogen-dependent TCL1A expression. Again, the LCLs were utilized in these experiments as the genotype of the LCLs with respect to the four SNPs was already known. After transiently transfecting LCLs of known genotype with ERα, the cells were exposed to varying concentrations of estradiol and the relationship between TCL1A expression and the SNP genotypes was determined. TCL1A expression was significantly greater in cells with variant SNP sequences than in those with the wild-type sequences in all three ethnic groups. It is important to remember that the variant sequence at rs11849538 that created an ERE.
The next steps in the functional genomics studies were influenced by the clinical impression that the musculoskeletal complaints seen in patients treated with AIs appeared consistent with an inflammatory response.20 Once again, using the LCLs, we determined that the expression of TCL1A was highly correlated with the expression of a series of genes encoding cytokines and cytokine receptors including the IL17 receptor A (IL17RA). The expression of TCL1A and IL17RA was highly correlated, P<1.9E−10. Additional studies in U2OS cells revealed that knockdown of TCL1A resulted in decreased expression of IL17RA but increased expression of IL17. Conversely, overexpression of TCL1A was associated with increased expression of IL17RA but decreased expression of IL17.
The studies relating TCL1A expression to cytokines were subsequently expanded by Liu et al.21 Again, extensive use was made of the LCLs to determine whether variation in TCL1A mRNA expression was associated with cytokine or cytokine receptor expression in these cells. A significant correlation was identified between TCL1A expression and a number of cytokine receptor genes. These five genes and the corresponding P-values for correlation with TCL1A expression were: IL13RA1 (interleukin 13 receptor, α1; P=3.16E−14), IL18R1 (interleukin 18 receptor 1; P=2.27E−13), IL1R2 (interleukin 1 receptor, type 2; P=1.73E−11), IL17RA (interleukin receptor A; P=1.92E−10) and IL12RB2 (interleukin 12 receptor, β2; P=4.84E−9). The effect of estrogen-dependent TCL1A expression in LCLs with known variant or wild-type SNP sequences on the expression of these receptors and their ligands was then determined. With increasing concentrations of estradiol, the expression of TCL1A and all of these interleukin receptors was all altered in a SNP-dependent manner. Additionally, a series of experiments was conducted that showed that TCL1A is ‘upstream’ of IL17RA, IL12RB2 and IL1R2.
As the main goal of this research was to determine how a reduction in estrogen concentrations, as caused by AI administration, might be related to the apparent clinical picture of inflammation in women who experience musculoskeletal complaints, this led us to focus on nuclear factor-κB (NF-κB), which is known to mediate joint inflammation.22 Again, using the LCLs with known variant and wild-type SNP genotypes, a series of experiments was performed with increasing concentrations of estradiol, both in the absence and the presence of a blocker of ERα (ICI 182,780). With increasing concentrations of estradiol, average TCL1A expression increased by about fivefold in the LCLs with the variant genotypes, but only about 40% in the LCLs with the wild-type genotype. Remarkably, with blockade of ERα, TCL1A expression dropped dramatically in the LCLs with the variant genotype to levels substantially below baseline, while in the LCLs with the wild-type genotype TCL1A expression increased ∼3.5-fold.
After the identification of these SNP-dependent effects, experiments were done to determine the impact of blockade of ERα on NF-κB transcriptional activity. This was done by utilizing NF-κB reporter gene assays in the same LCLs noted above. There was little change in NF-κB transcriptional activity with increasing doses of estradiol. However, again remarkably, the addition of an ERα blocker demonstrated a marked difference between the NF-κB transcriptional activity for the LCLs with the variant and the wild-type genotypes. That is, with the addition of ICI 182 780, NF-κB transcriptional activity increased by over threefold, whereas LCLs with the wild-type genotype showed a slight decrease in NF-κB transcriptional activity. This marked increase in NF-κB transcriptional activity after blockade of ERα seen with the variant genotypes may offer an explanation for the development of musculoskeletal complaints in women who have decreased estrogen levels following AI therapy.
Additional phenotypes being studied with patients from the MA.27 clinical trial
It is clear that the large MA.27 trial offers a unique opportunity to study the pharmacogenomics of AIs in postmenopausal women with resected early-stage breast cancer. It is highly unlikely that another clinical trial of this magnitude will be conducted in patients who receive monotherapy with an AI. Thus, it is crucial that as much knowledge as possible be obtained. Because of this, our group is focused on identifying the most important phenotypes to examine in collaboration with the RIKEN Center for Genomic Medicine. At present, there are two specific projects that are being conducted. The rationale for these projects is described in subsequent paragraphs.
A GWAS in patients experiencing bone fractures while receiving AIs on the MA.27 trial
Bone mass declines and fracture risk increases with advancing age, particularly in women as they enter the postmenopausal years.23 Osteoporotic fractures are known to be a major cause of morbidity and mortality, especially in developed countries,24 including Japan.25 Genetic factors clearly have a role in bone mineral density and osteoporosis risk,26 and GWAS have identified many statistically significant SNPs.27 As the mechanism of action of AIs involves a substantial reduction in estrogens, a major concern is an accelerated adverse impact on bone health in women already at an age when they are at an increased risk for bone loss and bone fragility fractures. This adverse impact on bone health appears to be the case for all the third-generation AIs and, in clinical trials comparing them to either tamoxifen or placebo, it has been estimated that fracture risk difference may be as high as up to 60% when AIs are employed.28, 29
On the basis of the high-quality data available in the MA.27 trial and the importance of fractures to women receiving AIs, we examined the fracture experience in this trial. We carefully selected sites of fractures that would be expected to be related to AI-associated bone loss, specifically those in the spine, forearm, humerus and proximal femur/hip, which would be considered fragility fractures. All reports of new fractures were reviewed by a team of investigators that included a recognized authority on bone health, Dr Khosla30 from Mayo clinic. We identified patients in these categories who had banked DNA and consented to genetic testing and, after strict quality control, we utilized 231 patients in our analyses. Thus, the trial had sufficient patients who experienced a relevant clinical fracture to allow for a GWAS study powered to detect SNPs associated with a large risk for bone fractures and a case–cohort study was performed. The genotyping for this study has been completed by the RIKEN Center for Genomic Medicine, the analysis is completed and the manuscript is in preparation.
A GWAS in patients experiencing breast events while receiving AIs on the MA.27 trial
The phenotype being studied in the ‘breast events GWAS’ is the STEEP31 end point, an acronym for ‘Standardized Definitions for Efficacy End Points in Adjuvant Breast Cancer Trials’, of breast cancer-free interval (BCFI). A BCFI event is defined as time from randomization to the first locoregional breast cancer recurrence, distant breast cancer recurrence, contralateral breast cancer or death with or from breast cancer without prior recurrence date. Follow-up is censored at non-breast cancer death. Although BCFI is the primary phenotype for this study, we recognize that there could be genetic differences that influence risk of recurrence versus risk of new breast cancers. For this reason, we will perform sensitivity analyses by repeating our planned analyses with contralateral breast cancers censored, to exclude them from the BCFI determination.
This study will focus on the efficacy of AIs when administered as monotherapy in women with resected early-stage breast cancer to prevent recurrence of the cancer. As noted in the Introduction, the worldwide experience with tamoxifen was utilized in a meta-analysis by the Early Breast Cancer Trialists' Collaborative Group (EBCTCG) and this revealed that 5 years of tamoxifen therapy reduced the breast cancer recurrence rates by about one-half during the first 5 years and by about one-third during the second 5 years, after discontinuation of the drug. The value of the AIs can be seen from the meta-analysis of trials comparing them to tamoxifen in which the AIs were found to be superior. This meta-analysis was performed by the Aromatase Inhibitors Overview Group (AIOG), composed of the leaders of adjuvant trials involving AIs, as a joint effort with the EBCTCG. The first publication5 from the AIOG comparing AIs with tamoxifen involved 9856 patients with a mean follow-up of 5.8 years and revealed at the 5-year time point, an absolute 2.9% reduction in recurrence (2P<0.00001) and a nonsignificant 1.1% reduction in breast cancer mortality (2P=0.1) for those women randomized to an AI vis-à-vis those randomized to tamoxifen. Despite the clear efficacy of the AIs as adjuvant endocrine therapy for early breast cancer, many women will still have a recurrence. For example, in the meta-analysis just described5, 9.6% of women treated with either anastrozole or letrozole experienced a recurrence of their breast cancer and there was no indication of a plateau in the recurrence rates. Given that MA.27 is the largest adjuvant endocrine therapy trial conducted to date that has exclusively studied AIs and, importantly, prospectively collected blood for DNA extraction and patient consent for its use in genetic studies, it represents a unique opportunity to conduct pharmacogenomic studies. The primary hypothesis in our ‘breast events’ GWAS is that there are genes related to hormone-dependent breast cancers that affect breast cancer relapse. The first step in this process is the identification of SNPs associated with BCFI. We will then relate these SNPs to genes and then follow the pharmacogenomic paradigm relating the genes to the drug effect and the clinical phenotype of breast cancer recurrence (Figure 1).
GWAS in postmenopausal women
The main pathway for estrogen synthesis in postmenopausal women is through conversion of androstenedione to estrone, and testosterone to estradiol by aromatase32, an enzyme present in many non-endocrine tissues including muscle, fat, and normal and malignant breast tissue. As noted, there is a remarkable variability in the response of postmenopausal women to AIs in terms of effectiveness of therapy and toxicities. To investigate this variability, Mayo investigators developed a prospective clinical study (MC0532), in collaboration with investigators at M.D. Anderson Cancer Center and Memorial Sloan Kettering Cancer Center, in women with resected early-stage breast cancer who were to undergo therapy with the AI anastrozole. The hypothesis to be tested was that inherited variation in pathways for anastrozole metabolism or transport (pharmacokinetics) and/or steroid hormone biosynthesis, metabolism and effect (pharmacodynamics) might contribute to individual variation in anastrozole efficacy and/or side effects. The Mayo group has extensive experience studying the human aromatase gene (CYP19) having resequenced the gene and performed initial functional genomic studies.33
The blood was collected for DNA extraction, for determination of hormone levels at baseline and while receiving anastrozole, and for determination of blood drug levels of anastrozole and its metabolites. In addition, we collected baseline and on-treatment mammograms and bone mineral density determinations. Thus, we have the ability to perform GWAS with multiple phenotypes including (1) baseline hormones (estradiol, estrone, estrone conjugates, androstenedione and testosterone), (2) change in hormone levels with anastrozole therapy with knowledge of levels of anastrozole and anastrozole metabolites, (3) baseline mammographic breast density, (4) change in mammographic breast density with anastrozole therapy, (5) baseline bone mineral density and (6) change in bone mineral density with anastrozole therapy. This population of almost 900 patients is remarkable because of the wealth of data available on each of the patients. That is, we have the five hormones determined by a very sophisticated validated bioanalytic method using gas chromatography–negative ion tandem mass spectrometry11, both at baseline and while on anastrozole therapy. The utilization of this highly sensitive assay for the hormones was considered essential, given the profound decrease in estrogens that occurs in women while taking anastrozole. In addition, we have mammograms for determination of mammographic breast density and dual-energy X-ray absorptiometry scans for bone mineral density, both at baseline and while on anastrozole therapy. Finally, as mentioned previously, the portfolio of data on each patient includes determination of anastrozole and anastrozole metabolite concentrations.
We have recently published our first report on a GWAS utilizing baseline, that is, before anastrozole, estradiol concentrations as the phenotype34 that involved 772 women. Genotyping was conducted at the RIKEN Center for Genomic Medicine utilizing the Illumina Human610-Quad BeadChip (Illumina, San Diego, CA, USA). After a rigorous quality control process, there were a final total of 563 945 SNPs included in the association analysis. We utilized the genome-wide SNP data obtained by genome-wide genotyping of the LCLs, previously described,18 to classify each specimen into one of the three major racial groups, which were Caucasian, African–American and Han Chinese. To avoid bias that might arise from these different racial groups, an eigen analysis was performed that resulted in the inclusion of six eigenvectors in the final model.
The association analysis involved 772 women who had plasma estradiol results. The factors included in the model were race, eigenvectors, body mass index, age, prior chemotherapy, ER and PgR status, and site at which the patient was entered. A SNP (rs1864729) on chromosome 8 near the TSPYL5 gene had the lowest P-value and achieved genome-wide significance (P=3.49E–08). Imputation, using 1000 Genomes Project data35, within 200 kb of this SNP was performed and revealed 17 additional SNPs that, after genotyping, were found to have P-values even lower than that of the rs1864729 SNP, that is, 1.50E−09 to 2.29E−08.
Examination of plasma estradiol concentrations revealed that patients homozygous for the variant rs1864729 SNP had average concentrations over twice as high as those for patients who were homozygous for the wild-type allele. Of interest is the fact that in a prior study,36 we had identified two SNPs in the aromatase gene (CYP191A) that were associated with elevated plasma estradiol concentrations and were in the CYP19A1 I.1 (placental) promoter. Upon genotyping these two SNPs in our current study population, a similar strong association was also identified.
Proceeding with our pharmacogenomic paradigm approach (Figure 1), we examined whether any of the chromosome 8 SNPs that achieved genome-wide significance (<5E−08) might have functional importance. Examination of the TRANSFAC database revealed that the variant allele for the rs2583506 SNP was predicted to create an ERE. Therefore, a ChIP assay was performed with LCLs that were either heterozygous for the rs2583506 SNP or were homozygous for the wild-type allele. These studies were performed after stably transfecting the LCLs with ERα. The ChIP assays showed no ERα binding for DNA from LCLs with wild-type rs2583506 SNP genotype but did show binding for DNA from cells heterozygous for the rs2583506 SNP variant sequence, thus confirming that this variant SNP created a functional ERE.
Because of the central role performed by CYP19A1 in determining estradiol concentrations in postmenopausal women, the relationship between TSPYL5 and CYP19A1 was examined. This was accomplished by both knockdown and overexpression of TSPYL5 in three different cell lines and examining CYP19A1 expression, taking into account that this gene has 10 different promoters37 that are considered generally tissue specific. These studies revealed that in MCF-7 cells, the expression of the I.4 promoter paralleled that of the TSPYL5 expression whether TSPYL5 was knocked down or overexpressed. Western blot analyses for TSPL5 and CYP19A1 paralleled the results of the expression studies.
The finding of an association between expression of TSPL5 and CYP19A1 was followed by a series of experiments examining the possibility of a TSPYL5 SNP-dependent relationship with the expression of CYP19A1. There was particular interest in these studies as, was noted above, one of the imputed SNPs, rs2583506, that had a genome-wide level of significance, was shown by a ChIP assay to create an ERE. Again, using LCLs stably transfected with ERα with known genotypes, the cells with the heterogeneous genotypes for rs2583506, and thus a functional ERE, showed greater TSPYL5 induction with increasing estradiol concentrations then did the homozygous wild-type cells that did not have the SNP that created the ERE. Of particular importance is that transcripts encoded by three different CYP19A1 promoters (I.1, I.4 and I.3) in cells with the variant genotype also showed a greater CYP191A expression then did the cells with the wild type.
To further examine the relationship between TSPYL5 expression and CYP19A1 expression, human adipocytes were utilized in which TSPYL5 was either knocked down or overexpressed. With TSPYL5 overexpression, there were increases in CYP19A1 expression that was driven by all three promoters.
As TSPYL5 had been shown to influence CYP19A1 expression in MCF-7 cells, LCLs and adipocytes by acting through the CYP19A1 I.4 promoter, a series of experiments was performed to see whether TSPYL5 directly bound to this promoter. Those studies revealed that an ∼120-bp region of DNA from this promoter was shown by ChIP assay to bind TSPYL5. The next step was to take this 120-bp sequence and do a homology search across the entire genome utilizing the Basic Local Alignment Search Tool (BLAST), a search that identified numerous genes that contained a portion of this sequence in their core promotors. After alignment of these sequences, a motif (5′-TCANNGAAGGCAG-3′) was identified that was present in 43 genes, 27 of which were expressed in three cell lines (MCF-7, IMR-90 and HEK293T). Again using knockdown or overexpression of TSPYL5 in these three cell lines, we found a correlation between TSPYL5 and the majority of the genes tested. That is, with TSPYL5 knockdown, the expression of 26 of the 27 genes decreased, and with TSPYL5 overexpression, the expression of 16 of the 27 genes increased.
This series of experiments began with the identification of variant SNPs in or near TSPYL5 that were associated with higher levels of estradiol in postmenopausal women; then showed an association of TSPYL5 expression with increased CYP19A1 expression, resulting in increased estradiol concentrations, which was also associated with increased expression of TSPYL5. The end result is a positive-feedback loop. Importantly, these studies provide a novel SNP-dependent mechanism for the regulation of CYP19A1 expression. These findings may have potential implications for research into individualizing AI therapy in postmenopausal women with breast cancer. They have also identified a novel transcription factor, TSPYL5.
Pharmacogenomics of SERMs in the prevention setting
As noted, breast cancer is the most common form of cancer in women both in the United States1 and Japan2, and prevention of this common disease is an area of high priority. The US FDA approval for tamoxifen and raloxifene was based on the NSABP P-1 and P-2 clinical trials that involved about 33 000 patients. Although tamoxifen and raloxifene have FDA approval, the acceptance of these drugs by American women and their physicians has been poor,38, 39 because of the relatively high number needed to treat to prevent one case of breast cancer (about 51) and the potential for not only serious but rare side effects such as thromboembolic events, but also bothersome side effects such as vasomotor symptoms. The Mayo PGRN established a collaboration with NSABP to perform a nested case–control GWAS with the phenotype being development of breast cancer in these high-risk women who were treated with one of these SERMs. Preliminary results have been presented40 that demonstrated SNPs on chromosome 16 that were associated with the development of breast cancer in these high-risk women. The variant and wild-type SNPs were associated with striking differences in estradiol-induced expression of ZNF423, BRCA1 and BRCA2, the latter two of which are the most important breast cancer predisposition genes. Extensive functional genomic studies were subsequently performed and a manuscript describing these is currently in press.41
A major question that exists with tamoxifen therapy is the role of cytochrome P450 2D6 (CYP2D6) genotype in the efficacy of tamoxifen. Most of the research on this question has been conducted in the adjuvant therapy setting in women with resected invasive breast cancer. However, as the association between CYP2D6 and efficacy of tamoxifen for prevention is unknown, we utilized the 591 cases and 1126 controls in this GWAS to determine the impact of CYP2D6 genotype, CYP2D6 inhibitor use and CYP2D6 metabolizer status, which combines genotype and inhibitor use, to explore this question. Using comprehensive CYP2D6 genotyping, we found that alterations in CYP2D6 metabolism were not associated with either tamoxifen or raloxifene efficacy in women at high risk of developing breast cancer in these prevention trials.42
Conclusion
The studies noted above illustrate the utilization of a pharmacogenomic paradigm that begins with the highest quality genome-wide genotyping of germline DNA of well-defined large cohorts of women with well-defined phenotypes that is then followed by focused functional genomic studies. The SNPs identified in the GWAS are related to genes, which in turn are related to drug effect and clinical phenotype (Figure 1). The findings of SNP-dependent influences on the expression of numerous genes has led to the identification of new biological hypotheses that continue under investigation. We feel that this paradigm has been productive of new knowledge that should bring us closer to true personalized endocrine therapy of breast cancer.
References
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J. Clin. 63, 11–30 (2013).
National Cancer Center.. Cancer Statistics in Japan ’2010, (Foundation for Promotion of Cancer Research: Tokyo, Japan, 2010).
Early Breast Cancer Trialists' Collaborative Group. Relevance of breast cancer hormone receptors and other factors to the efficacy of adjuvant tamoxifen: patient-level meta-analysis of randomised trials. Lancet. 378, 771–784 (2011).
Ingle, J. N. Overview of adjuvant trials of aromatase inhibitors in early breast cancer. Steroids 76, 765–767 (2011).
Dowsett, M., Cuzick, J., Ingle, J., Coates, A., Forbes, J., Bliss, J. et al. Meta-analysis of breast cancer outcomes in adjuvant trials of aromatase inhibitors versus tamoxifen. J. Clin. Oncol. 28, 509–518 (2010).
Burstein, H. J., Prestrud, A. A., Seidenfeld, J., Anderson, H., Buchholz, T. A., Davidson, N. E. et al. American Society of Clinical Oncology Clinical Practice Guideline: update on adjuvant endocrine therapy for women with hormone receptor-positive breast cancer. J. Clin. Oncol. 28, 3784–3796 (2010).
Cuzick, J., Powles, T., Veronesi, U., Forbes, J., Edwards, R., Ashley, S. et al. Overview of the main outcomes in breast-cancer prevention trials. The Lancet 361, 296–300 (2003).
Fisher, B., Costantino, J. P., Wickerham, D. L., Redmond, C. K., Kavanah, M., Cronin, W. M. et al. Tamoxifen for prevention of breast cancer: report of the National Surgical Adjuvant Breast and Bowel Project P-1 Study. J. Natl. Cancer Inst. 90, 1371–1388 (1998).
Vogel, V. G., Costantino, J. P., Wickerham, D., Cronin, W. M., Cecchini, R. S., Atkins, J. N. et al. Effects of tamoxifen vs raloxifene on the risk of developing invasive breast cancer and other disease outcomes: the NSABP Study of Tamoxifen and Raloxifene (STAR) P-2 Trial. JAMA 295, 2727–2741 (2006).
Vogel, V. G., Costantino, J. P., Wickerham, D. L., Cronin, W. M., Cecchini, R. S., Atkins, J. N. et al. Update of the National Surgical Adjuvant Breast and Bowel Project Study of Tamoxifen and Raloxifene (STAR) P-2 Trial: preventing breast cancer. Cancer Prev. Res. 3, 696–706 (2010).
Ingle, J. N., Buzdar, A. U., Schaid, D. J., Goetz, M. P., Batzler, A., Robson, M. E. et al. Variation in anastrozole metabolism and pharmacodynamics in women with early breast cancer. Cancer Res. 70, 3278–3286 (2010).
Offit, K. & Robson, M. E. New pharmacogenomic paradigm in breast cancer treatment. J. Clin. Oncol. 28, 4665–4666 (2010).
Ingle, J. N., Schaid, D. J., Goss, P. E., Liu, M., Mushiroda, T., Chapman, J. A. et al. Genome-wide associations and functional genomic studies of musculoskeletal adverse events in women receiving aromatase inhibitors. J. Clin. Oncol. 28, 4674–4682 (2010).
Goss, P. E., Ingle, J. N., Pritchard, K. I., Ellis, M. J., Sledge, G. W., Budd, G. T. et al. Exemestane versus anastrozole in postmenopausal women with early breast cancer: NCIC CTG MA.27--a randomized controlled phase III trial. J. Clin. Oncol. 31 (11), 1398–1404 (2013).
Partridge, A. H., LaFountain, A., Mayer, E., Taylor, B. S., Winer, E. & Asnis-Alibozek, A. Adherence to initial adjuvant anastrozole therapy among women with early-stage breast cancer. J. Clin. Oncol. 26, 556–562 (2008).
Crew, K. D., Greenlee, H., Capodice, J., Raptis, G., Brafman, L., Fuentes, D. et al. Prevalence of joint symptoms in postmenopausal women taking aromatase inhibitors for early-stage breast cancer. J. Clin. Oncol. 25, 3877–3883 (2007).
Henry, N. L., Giles, J. T., Ang, D., Mohan, M., Dadabhoy, D., Robarge, J. et al. Prospective characterization of musculoskeletal symptoms in early stage breast cancer patients treated with aromatase inhibitors. Breast Cancer Res. Treat. 111, 365–372 (2008).
Niu, N., Qin, Y., Fridley, B. L., Hou, J., Kalari, K. R., Zhu, M. et al. Radiation pharmacogenomics: a genome-wide association approach to identify radiation response biomarkers using human lymphoblastoid cell lines. Genome Res. 20, 1482–1492 (2010).
Li, L., Fridley, B., Kalari, K., Jenkins, G., Batzler, A., Safgren, S. et al. Gemcitabine and cytosine arabinoside cytotoxicity: association with lymphoblastoid cell expression. Cancer Res. 68, 7050–7058 (2008).
Miossec, P., Korn, T. & Kuchroo, V. K. Interleukin-17 and type 17 helper T cells. N. Engl. J. Med. 361, 888–898 (2009).
Liu, M., Wang, L., Bongartz, T., Hawse, J. R., Markovic, S. N., Schaid, D. J. et al. Aromatase inhibitors, estrogens and musculoskeletal pain: estrogen-dependent T-cell leukemia 1A (TCL1A) gene-mediated regulation of cytokine expression. Breast Cancer Res. 14, R41 (2012).
Bokarewa, M., Nagaev, I., Dahlberg, L., Smith, U. & Tarkowski, A. Resistin, an adipokine with potent proinflammatory properties. J. Immunol. 174, 5789–5795 (2005).
Cummings, S. R. & Melton, L. J. III. Epidemiology and outcomes of osteoporotic fractures. Lancet 359, 1761–1767 (2002).
Johnell, O. & Kanis, J. A. An estimate of the worldwide prevalence and disability associated with osteoporotic fractures. Osteoporos. Int. 17, 1726–1733 (2006).
Finkelstein, J. S., Brockwood, S. E., Mehta, V., Greendale, G. A., Sowers, M. R., Ettinger, B. et al. Bone mineral density changes during the menopause transition in a multiethnic cohort of women. J. Clin. Endocrinol. Metab. 93, 861–868 (2008).
Ferrari, S. Human genetics of osteoporosis. Best. Pract. Clin. Endocrinol. Metab. 22, 723–735 (2008).
Estrada, K., Styrkarsdottir, U., Evangelou, E., Hsu, Y.-H., Duncan, E. L., Ntzani, E. E. et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat. Genet. 44, 491–501 (2012).
Eastell, R. & Hannon, R. Long-term effects of aromatase inhibitors on bone. J. Steroid Biochem. Molec. Biol. 95, 151–154 (2005).
Chien, A. J. & Goss, P. E. Aromatase inhibitors and bone health in women with breast cancer. J. Clin. Oncol. 24, 5305–5312 (2006).
Khosla, S. Increasing options for the treatment of osteoporosis. N. Engl. J. Med. 361, 818–820 (2009).
Hudis, C. A., Barlow, W. E., Costantino, J. P., Gray, R. J., Pritchard, K. I., Chapman, J. W. et al. proposal for standardized definitions for efficacy end points in adjuvant breast cancer trials: the STEEP system. J. Clin. Oncol. 25, 2127–2132 (2007).
Simpson, E. R., Mahendroo, M. S., Means, G. D., Kilgore, M. W., Hinshelwood, M. M., Graham-Lorrence, S. et al. Aromatase cytochrome P450, the enzyme responsible for estrogen biosynthesis. Endocr. Rev. 15, 342–355 (1994).
Ma, C. X., Adjei, A. A., Salavaggione, O. E., Coronel, J., Pelleymounter, L., Wang, L. et al. Human aromatase: gene resequencing and functional genomics. Cancer. Res. 65, 11071–11082 (2005).
Liu, M., Ingle, J. N., Fridley, B. L., Buzdar, A. U., Robson, M. E., Kubo, M. et al. TSPYL5 SNPs: Association with plasma estradiol concentrations and aromatase expression. Mol. Endocrinol. 4, 657–670 (2013).
Durbin, R. M., Abecasis, G. R., Altshuler, D. L., Auton, A., Brooks, L. D., Gibbs, R. A. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Wang, L., Ellsworth, K. A., Moon, I., Pelleymounter, L. L., Eckloff, B. W., Martin, Y. N. et al. Functional genetic polymorphisms in the aromatase gene CYP19 vary the response of breast cancer patients to neoadjuvant therapy with aromatase inhibitors. Cancer Res. 70, 319–328 (2010).
Bulun, S. E., Sebastian, S., Takayama, K., Suzuki, T., Sasano, H. & Shozu, M. The human CYP19 (aromatase P450) gene: update on physiologic roles and genomic organization of promoters. J. Steroid. Biochem. Mol. Biol. 86, 219–224 (2003).
Ropka, M. E., Keim, J. & Philbrick, J. T. Patient decisions about breast cancer chemoprevention: a systematic review and meta-analysis. J. Clin. Oncol. 28, 3090–3095 (2010).
Armstrong, K., Quistberg, D., Micco, E., Domchek, S. & Guerra, C. Prescription of tamoxifen for breast cancer prevention by primary care physicians. Archiv. Int. Med. 166, 2260–2265 (2006).
Ingle, J. N., Liu, M., Wickerham, D. L., Schaid, D. J., Mushiroda, T., Kubo, M. et al. Genome-wide associations of breast events and functional genomic studies in high-risk women receiving tamoxifen or raloxifene on NSABP P1 and P2 prevention trials. A Pharmacogenomics Research Network-RIKEN-NSABP Collaboration. Cancer Res. 70 (24 Suppl), 110s (2010).
Ingle, J. N., Liu, M., Wickerham, D. L., Schaid, D. J., Wang, L., Mushiroda, T. et al. Selective estrogen receptor modulators and pharmacogenomic variation in ZNF423 regulation of BRCA1 expression: individualized breast cancer prevention. Cancer Discov. (in press).
Goetz, M. P., Schaid, D. J., Wickerham, D. L., Safgren, S., Mushiroda, T., Kubo, M. et al. Evaluation of CYP2D6 and efficacy of tamoxifen and raloxifene in women treated for breast cancer chemoprevention: results from the NSABP P1 and P2 clinical trials. Clin. Cancer Res. 17, 6944–6951 (2011).
Acknowledgements
Dr Ingle acknowledges the many investigators and scientists who have contributed to this body of work, in particular, Drs Richard Weinshilboum, Michiaki Kubo, Yusuke Nakmura, Daniel Schaid and Mohan Liu.
Funding sources: These studies were supported in part by NIH grants U19 GM61388 (The Pharmacogenomics Research Network), P50 CA116201 (Mayo Clinic Breast Cancer Specialized Program of Research Excellence), U10 CA37377, U10 CA69974, U24 CA114732, U01 GM63173, U10 CA77202, U10 CA32102, R01 CA38461, R01 GM28157, R01 CA113049, R01 CA 138461, U01 HG005137, a gift from Bruce and Martha Atwater, CCS 015469 from the Canadian Cancer Society, and the RIKEN Center for Genomic Medicine and the Biobank Japan Project funded by the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Ingle, J. Pharmacogenomics of endocrine therapy in breast cancer. J Hum Genet 58, 306–312 (2013). https://doi.org/10.1038/jhg.2013.35
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.35
Keywords
This article is cited by
-
Adjuvant endocrine therapy is associated with improved overall survival in elderly hormone receptor-positive breast cancer patients
Breast Cancer Research and Treatment (2020)