Abstract
Hereditary spastic paraplegia (HSP) is one of the most genetically heterogeneous neurodegenerative disorders characterized by progressive spasticity and pyramidal weakness of lower limbs. Because >30 causative genes have been identified, screening of multiple genes is required for establishing molecular diagnosis of individual patients with HSP. To elucidate molecular epidemiology of HSP in the Japanese population, we have conducted mutational analyses of 16 causative genes of HSP (L1CAM, PLP1, ATL1, SPAST, CYP7B1, NIPA1, SPG7, KIAA0196, KIF5A, HSPD1, BSCL2, SPG11, SPG20, SPG21, REEP1 and ZFYVE27) using resequencing microarrays, array-based comparative genomic hybridization and Sanger sequencing. The mutational analysis of 129 Japanese patients revealed 49 mutations in 46 patients, 32 of which were novel. Molecular diagnosis was accomplished for 67.3% (33/49) of autosomal dominant HSP patients. Even among sporadic HSP patients, mutations were identified in 11.1% (7/63) of them. The present study elucidated the molecular epidemiology of HSP in the Japanese population and further broadened the mutational and clinical spectra of HSP.
Similar content being viewed by others
Introduction
Hereditary spastic paraplegia (HSP) is a neurodegenerative disorder characterized by progressive lower limb spasticity and pyramidal weakness, which is one of the most genetically and clinically heterogeneous disorders.1, 2 HSP is clinically divided into two forms, pure and complicated forms, depending on whether the neurological symptoms are basically confined to spasticity and pyramidal weakness of the lower limbs or accompanied by additional neurological symptoms such as cognitive dysfunction, cerebellar signs, optic atrophy, retinitis pigmentosa, amyotrophy and peripheral neuropathy. HSP is characterized by enormous genetic heterogeneity; to date, more than 50 genetic loci (SPG1-57) and 37 causative genes have been identified: L1CAM (SPG1), PLP1 (SPG2), ATL1 (SPG3A), SPAST (SPG4), CYP7B1 (SPG5A), NIPA1 (SPG6), SPG7 (SPG7), KIAA0196 (SPG8), KIF5A (SPG10), SPG11 (SPG11), RTN2 (SPG12), HSPD1 (SPG13), SPG15/ZFYVE26 (SPG15), BSCL2 (SPG17), ERLIN2 (SPG18), SPG20 (SPG20), SPG21 (SPG21), DDHD1 (SPG28), KIF1A (SPG30), REEP1 (SPG31), ZFYVE27 (SPG33), FA2H (SPG35), PNPLA6 (SPG39), SLC33A1 (SPG42), GJC2 (SPG44), GBA2 (SPG46), AP4B1 (SPG47), KIAA0415 (SPG48), TECPR2 (SPG49), AP4M1 (SPG50), AP4E1 (SPG51), AP4S1 (SPG52), VPS37A (SPG53), DDHD2 (SPG54), C12ORF65 (SPG55), CYP2U1 (SPG56), and TFG (SPG57).
Because of the limited availability of information on genotype–phenotype correlations and locus heterogeneity, it is often difficult to prioritize genes for the mutational analysis of HSP. Therefore, it is essential to incorporate knowledge of the molecular epidemiology of HSP and relative frequencies of the types of mutations (substitution, insertion/deletion or rearrangement) in each gene into the algorithm of molecular diagnosis of HSP. We also need to be aware that different methodologies are required to detect each type of mutations with high sensitivities. Although there have been studies on molecular epidemiology focusing on selected causative genes in large case series,3, 4, 5, 6, 7, 8, 9, 10 there have been only few studies based on comprehensive mutational analyses focusing on multiple genes as well as various types of mutation. Thus, the comprehensive molecular epidemiology of HSP is largely unestablished.
For these reasons, a comprehensive mutational analysis of multiple genes is necessary to efficiently provide molecular diagnosis for individual HSP patients and, furthermore, to clarify the molecular epidemiology of HSP. To accomplish high sensitivities for detection of various kinds of mutations, we have conducted comprehensive mutational analyses incorporating custom-made resequencing microarrays,11 which enable comprehensive detection of single-nucleotide variations, custom-made comparative genomic hybridization (CGH) microarrays,12 which enable efficient detection of large insertion/deletion variants, and Sanger sequencing, which enables detection of small insertions/deletions in addition to single base substitutions. We herein describe molecular epidemiology and the clinical spectrum of HSP based on a large-scale comprehensive mutational analysis of 129 Japanese patients with various forms of HSP.
Subjects and methods
Patients
One hundred and twenty-nine Japanese patients (75 male and 54 female) with a clinical diagnosis of HSP were enrolled in the study, including 45 patients who visited the University of Tokyo Hospital and 89 patients referred to our Department of Neurology, the University of Tokyo Tokyo, Japan for the molecular diagnosis of HSP from various regions in Japan between 1994 and July 2007. Genomic DNAs were prepared from peripheral blood leukocytes or an autopsied brain (1 patient) using a standard procedure. Written informed consent was obtained from all the participants or their family members. The study was approved by the institutional review board of the University of Tokyo.
Outline of mutational analysis system
The outline of the comprehensive mutational analysis is shown in Figure 1. All the samples were first subjected to resequencing microarray analysis for analyzing 13 causative genes of HSP. Among the patients in whom mutations were not detected by resequencing microarray analysis, direct nucleotide sequence analysis of SPAST and REEP1 was carried out in patients with autosomal dominant HSP (AD-HSP) and sporadic pure-form HSP; in patients with a thin corpus callosum and cognitive impairment, direct nucleotide sequence analysis of SPG11 was carried out. For those in which mutations were not detected by any of these methods, array-based CGH (aCGH) analysis was carried out.
Flowchart of mutational analysis of HSP genes. We first performed resequencing microarray analysis, which could analyze 13 causative genes of HSP. Then, samples of mutation-negative AD-HSP and 10 sporadic pure HSP patients were subjected to direct nucleotide sequence analysis of SPAST and REEP1, because small insertion/deletion mutations are relatively frequent in these genes. Samples of mutation-negative HSP patients with a thin corpus callosum and cognitive impairment were subjected to direct nucleotide sequence analysis of SPG11. Finally, aCGH analysis was performed in the 92 mutation-negative patients.
Mutational analysis using custom-made resequencing microarrays
We developed resequencing microarrays using GeneChip CustomSeq (Affymetrix, Santa Clara, CA, USA).11 We utilized custom-designed microarrays of the 30-kb format that contain tiled sequences for SPAST (NM_014946.3) and ATL1 (NM_015915) (TKYPD01), those for SPG7 (NM_003119) (TKYALS01), those for L1CAM (NM_000425) and PLP1 (NM_000533) (TKYAD01) and those for NIPA1 (NM_144599), KIF5A (NM_004984) and SPG20 (NM_015087) (TKYPD02), as previously described.13, 14, 15 In this study, we additionally designed two 50-kb-format microarrays. One was TKYPD03, which contained tiled sequences for SPAST, ATL1 and REEP1, and the other was TKYALS02, which contained tiled sequences for 10 causative genes (L1CAM (NM_000425), PLP1 (NM_000533), NIPA1 (NM_144599), SPG7 (NM_003119), KIAA0196 (NM_014846), KIF5A (NM_004984), HSPD1 (NM_002156), BSCL2 (NM_001122955), SPG20 (NM_015087) and SPG21 (NM_016630)), enabling mutational analysis of 13 causative genes of HSP. Experiments were performed following the manufacturer’s instructions (Supplementary Information S1). Data were analyzed using GeneChip DNA Analysis Software version 2.0 (GDAS2.0) for 30-kb-format microarrays,16 or updated GeneChip Sequence Analysis Software version 4.0 (GSEQ4.0) adjunctively with a custom-designed program (Supplementary Information S2). All the called mutations were verified by direct nucleotide sequence analysis. The frequencies of detected nonsynonymous variations in the populations were checked using dbSNP (http://www.ncbi.nlm.nih.gov/snp/), the 1000 genomes database (http://www.1000genomes.org/) and by screening of >150 control chromosomes by direct nucleotide sequence analysis or restriction fragment length polymorphism analysis.
Mutational analysis using aCGH
We custom-designed a CGH microarray (Agilent Technology, Santa Clara, CA, USA) in the 8 × 15 K format.12 The genomic sequences of 16 causative genes of HSP and the flanking regions (L1CAM, PLP1, ATL1, SPAST, CYP7B1, NIPA1, SPG7, KIAA0196, KIF5A, SPG11, HSPD1, BSCL2, SPG20, SPG21, REEP1 and ZFYVE27) were tiled on the array (Supplementary Information S1). The average distance between probes was 200 bp. When insertion/deletion mutations were detected by the aCGH analysis, breakpoints were determined by PCR analysis using primer pairs flanking the breakpoints and direct nucleotide sequence analysis.17, 18
Direct nucleotide sequence analysis
Direct nucleotide sequence analysis was performed using ExoSAP-IT (USB, Cleveland, OH, USA), a BigDye Terminator v3.1 kit, and XTerminator using an ABI PRISM3100 sequencer (Applied Biosystems, Foster City, CA, USA). Primers and the amplification condition are described in Supplementary Information S1.
Results
Demographic characteristics of patients
The demographic characteristics of the 129 HSP patients enrolled in this study are summarized in Table 1. The ages at onset of the patients classified on the basis of the clinical form of HSP (Supplementary Figure S1A) revealed a bimodal distribution in patients with pure-form HSP, whereas one large peak for juvenile onset and one small peak of adult to late onset were observed in patients with complicated-form HSP. Focusing on the mode of inheritance (Supplementary Figure S1B), the ages at onset of AD-HSP patients and sporadic HSP patients showed a similar bimodal distribution, whereas those of autosomal recessive HSP (AR-HSP) patients showed a skewed distribution.
We found 14 patients with complicated-form HSP with a thin corpus callosum and cognitive impairment. There were no patients with AD-HSP with motor neuropathy clinically diagnosed as Silver syndrome.19
Mutational analysis by resequencing microarray analysis
All the samples were first subjected to resequencing microarray analysis (Figure 1). The analysis detected 22 mutations, all of which were nucleotide substitutions (Table 2). Representative resequencing microarray data on a heterozygous mutation in SPAST (c.1493+2 T>C) are shown (Figures 2a–c). Using GDAS2.0 or GSEQ4.0, the overall call rate was about 90%. Except for the nucleotides for which base calling was difficult because of high GC contents, G/C stretches or locally repetitive polymorphic sequences (such as a GCG stretch in exon 1 of NIPA1), the overall rate of base calling using GDAS/GSEQ4.0 in combination with visual inspection was as high as 99.9%.
Mutational analysis using resequencing microarrays and comparative genomic hybridization microarrays. (a) This figure shows a scan image obtained by resequencing microarray analysis (TKYPD03) of a sporadic HSP patient. Each tile in a square indicates one of the four nucleotides. Depending on the nucleotide of each allele, each quadrant provides a fluorescent signal. As shown in a square that corresponds to the position of c.1493+2 of SPAST, the upper left tile and the upper right tile, which correspond to T and C, respectively, provided similarly intense hybridization signals. The signal pattern indicates the existence of the T allele (wild type) and the C allele (variant) in that position. (b) Scan image of the same positions of the resequencing microarray as those in panel (a) obtained from a mutation-negative patient, where only the upper left tile corresponding to ‘T’ gives an intense fluorescent signal. (c) Heterozygous c.1493+2 T>C mutation confirmed by direct nucleotide sequence analysis, which is expected to disrupt the consensus splice donor site. (d) Example of comparative genomic hybridization analysis. The vertical axis indicates the log2 ratio of hybridization signal intensities obtained from a patient with SPG4 and a male control subject. The horizontal axis indicates the physical position of oligonucleotide probes. If copy number variations do not exist, the log2 ratios of the hybridization signal intensities are expected to be near 0. In the region indicated by an orange bar, the log2 ratio of hybridization signal intensities is approximately −1, which indicates a heterozygous deletion (halved gene dosage) in SPAST. (e) PCR analysis using primers flanking the deletions revealed that the truncated band corresponding to 1.8 kb was detected only in the patient. No PCR product was detected in a control, because the distance between the primer pair was too long to amplify (∼11 kb). (f) Direct nucleotide sequence analysis determines breakpoints with a deletion size of 9565 bp.
The custom-designed program detected one mutation (c.1741C>T, p.R581* in SPAST), which was not detected by GSEQ4.0, increasing the sensitivity of mutation detection (Supplementary Information S2). No additional base substitutions were detected by the subsequent direct nucleotide sequence analysis of SPAST and REEP1 in AD-HSP patients.
Mutational analysis by Sanger sequencing
We applied Sanger sequencing of SPAST and REEP1 in patients with AD-HSP and sporadic pure-form HSP, in which mutations were not detected by the resequencing microarray analysis, mainly to detect insertion/deletion mutations that are common in these diseases (Figure 1). We detected 10 small insertion/deletion mutations (1–41 bp) in SPAST and 1 insertion in REEP1 (Table 2).
In patients with a thin corpus callosum and cognitive impairment among the cases in which mutations were not detected by the resequencing microarray analysis, we then applied Sanger sequencing of SPG11 (Figure 1). We found homozygous or compound heterozygous mutations of SPG11 in five patients with family histories consistent with the autosomal recessive mode of inheritance. In a sporadic HSP patient, we found compound heterozygous mutations of c.1735+2 delT and c.6999+5 delG, both of which were considered pathogenic because no other pathogenic alleles were detected, and in silico analysis of splicing scores of c.6999+5 delG showed scores that decreased from 10.1 to 7.1 (http://rulai.cshl.edu/new_alt_exon_db2/HTML/score.html) and from 1.0 to 0.62 (http://www.fruitfly.org/seq_tools/splice.html). Another patient had only one null allele (p.R2031*) in SPG11 and no other pathogenic alleles were found.
Detection of large deletions and duplications by aCGH analysis
We applied aCGH analysis in patients in whom mutations were not detected by resequencing microarray analysis and direct nucleotide sequencing analysis (Figure 1). Representative results for a heterozygous large deletion in KIAA0196 are shown in Figure 2d, in which the breakpoint sequence was clearly determined using an appropriate primer pair (Figure 2e).
In total, we identified 7 large deletions (4 in SPAST, 1 in REEP1, 1 in KIAA0196, and 1 in SPG11) and 1 duplication in SPAST in 92 patients examined (Table 2). All the breakpoints but one (in which the deletion was beyond the tiled sequences on the array) were unequivocally determined at the nucleotide level. The sizes of the deletions/duplication ranged from 4634 bp to >170 kb.
Five (four SPAST deletions and one SPG11 deletion) of the seven deletion breakpoints were inside Alu sequences. Among the breakpoint sequences of the remaining deletions, the deletion in REEP1 and the deletion in KIAA0196 showed microhomology of 3 bp. The breakpoint sequence of the SPAST duplication showed no homologous sequences. In total, five of the eight breakpoints (62.5%) were inside Alu sequences.
Molecular epidemiology of HSP in Japanese population
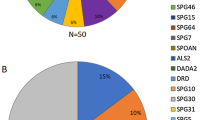
In summary, we found 49 mutations in 46 patients (Table 2). The relative frequencies of individual HSP types classified on the basis of the clinical presentations and the mode of inheritance are summarized in Figures 3a and b.
Relative frequencies of individual HSP types in groups classified on the basis of the clinical presentations and mode of inheritance. The figure shows the relative frequencies of individual HSP types in our cohort. (a) Pure form and (b) complicated form. The family history of each subgroup is indicated above the figures. Mutations were detected in a total of 67.3% of all the AD-HSP patients or 72.7% of the patients with pure-form AD-HSP. Focusing on sporadic HSP patients, six patients (four SPG4, one SPG3A and one SPG11) were identified, which accounted for 9.8% (6/61). Of note, SPAST mutations were present in 6.6% of all sporadic HSP patients, and particularly in 12.9% (4/31) of sporadic pure-form HSP patients, suggesting the usefulness of mutational analysis of SPAST in sporadic cases, particularly in patients with the pure form. Others, patients with unidentified mutation.
Focusing on all AD-HSP patients, SPG4 (55.1%) was the most frequent. SPG3A (2.0%), SPG8 (4.1%), and SPG31 (4.1%) were relatively rare in Japanese HSP patients. The frequency of SPG3A is lower than that in the Caucasian populations.5 The frequencies of SPG8 (4.1%) and SPG31 (4.1%) in AD-HSP are comparable to those reported in Caucasian populations.20, 21
In the AR-HSP group, we found five families with SPG11 and one family with SPG21. Among the 14 patients with a thin corpus callosum and cognitive impairment, 35.7% (5/14) carried SPG11 mutations.
Molecular and clinical spectra of individual HSP types
SPG3A
We found two patients with SPG3A carrying previously reported mutations (Table 2). Although both patients with SPG3A showed basically pure-form HSP with juvenile onset, one patient showed hypesthesia and hypalgesia in the distal lower limbs accompanied by decreased vibratory sensation in all extremities.
SPG4
Of the 32 patients with SPG4, 24 (75%) had nonsense, frameshift or large deletion/duplication mutations leading to truncated proteins, which were distributed throughout the genes (Supplementary Figure S2). On the other hand, seven out of the eight missense mutations were located in the AAA domain (ATPase associated with various cellular activities). We found a novel mutation (p.Y52C) outside the AAA domain. Note that large deletions/duplications in SPAST were detected by aCGH analysis, and small deletion mutations were detected by Sanger sequencing analysis in 22.7% (5/22)22 and 45.5% (10/22) of AD-HSP patients, respectively, in whom no mutations were detected by the resequencing microarray analysis. The ages at onset of patients with SPG4 showed two peaks, in the teens and in 40 s (Supplementary Figure S3A). The types of the mutation in SPAST and age at onset did not correlate (Supplementary Figure S3B).
SPG8
We found a large deletion in KIAA0196, which has not been described to date. The breakpoints of the large deletion in KIAA0196 are located in intron 10 and exon 15 (Figures 4a–c). RT-PCR and direct nucleotide sequence analyses revealed that exons 10–15 were deleted in cDNA, predicting a premature termination codon (Figures 4d and e). There are only three missense mutations reported to date, and in a previous paper, it was proposed that haploinsufficiency is the disease-causing mechanism of SPG8 on the basis of experiments using zebrafish.20 The large deletion in KIAA0196 detected in the present study further supported a disease mechanism of haploinsufficiency and indicate a necessity of screening for rearrangements of KIAA0196 in AD-HSP. SPG8 has been reported to be an ‘aggressive’ subtype of HSP and the disease onset is in the 20s or 30s.20 In contrast, two patients with SPG8 found in the study had adult-onset or late-onset HSP.
Mutations in KIAA0196. (a) Result of CGH array showing a heterozygous deletion in KIAA0196. An orange bar shows heterozygous deletion. (b) Direct nucleotide sequence analysis of the breakpoint in KIAA0196, which shows 4634 bp deletion. (c) Schematic presentation of the exon–intron structure of KIAA0196. The deletion detected by the array CGH analysis is shown. (d) RT-PCR analysis of species of RNAs extracted from the patient with the KIAA0196 deletion and a control. In the control, only a single band with the expected size corresponding to 1422 bp was observed, while a truncated band with the size corresponding to 814 bp in addition to PCR products corresponding to 1422 bp was observed in the patient. (e) Direct nucleotide sequence analysis of the truncated PCR products revealed that exons 11–15 were absent in the KIAA0196 mRNA as a result of a deletion in KIAA0196. (f) Schematic representation of strumpellin, the protein product of KIAA0196, and the mutations identified in patients with SPG8. The position of the large deletion (deletion of exons 11–14 and a part of exon 15) and the novel mutation found in the present study are shown (red). Previously reported mutations in KIAA0196 (p.N471D, p.L619F and p.V626F) are located in the spectrin-repeat-containing domain (amino acids 434–518) or the conserved domain with unknown function (amino acids 576–649). The novel mutation (p.R583S) found in the present study is also located in the conserved domain with unknown function. (g) Pedigree charts of the Japanese SPG8 families. Age at onset and age at examination are indicated.
SPG11
The five patients with SPG11 showed complicated-form HSP with cognitive impairment and a thin corpus callosum. Notably, rearrangement in SPG11 was found in a patient, and aCGH analysis was helpful for accurate diagnosis of the patient. The age at onset ranged from 2 to 25 years. Although SPG11 is allelic to juvenile amyotrophic lateral sclerosis (ALS5),23 none of the patients showed the ALS phenotype.
SPG17
A novel BSCL2 (NM_032667) p.C36Y substitution (which can also be called p.C100Y in NM_001122955 because there are two known start codons) was found in one AD-HSP patient. He suffered from early-onset spastic paraparesis with mild mental retardation and did not show amyotrophy. Clinical and genetic data of other family members were not available. C36 is conserved among species and is located in the first transmembrane domain,24 raising a possibility that p.C36Y can change the function of seipin, the protein product of BSCL2. Because only p.N88S and p.S90L of seipin have been described in Silver syndrome/SPG17, we still need to be cautious about the pathogenicity of p.C36Y substitution.
SPG21
We found a novel homozygous amino-acid substitution (p.A108P) in SPG21 encoding maspardin in a family with late-onset complicated-form HSP (Figure 5, Supplementary Tables S1 and S2). The two patients managed to walk with a cart or a cane in their 70s and 60s. In addition to cognitive decline, callosal disconnection syndrome, such as ideomotor apraxia predominantly of the left hand, agraphia of the left hand and constructional impairment predominantly on the right side, was observed, which was mild but progressed over 5 years in the index patient. There were no extrapyramidal signs, cerebellar signs or bulbar symptoms, as reported in the original family with an SPG21 mutation.25 Magnaetic resonance imaging of the index patient showed progressive thinning of the corpus callosum and predominantly frontotemporal atrophy (Figure 5e–i). 123I-N-isopropyl-p-iodoamphetamine single-photon emission computed tomography revealed decreased blood flow in the frontal and temporal cortices (Figure 5j).
A family with SPG21 and molecular genetic analysis. (a) Pedigree tree of the family. Squares indicate males and circles indicate females. Black squares are affected members and the index patient (II-2) is indicated by an arrow. Symbols with a diagonal line indicate deceased members. Members with dots allowed us neurological and genetic examinations. (b) Electropherograms of the family members carrying homozygous c.322G>C mutation (II-2), heterozygous c.322G>C mutation (II-4) and wild-type allele (II-5). (c) PCR-restriction fragment length polymorphism (RFLP) analysis of family members. The uncut PCR fragment length is 344 bp. With HaeIII digestion, the wild-type allele shows fragment sizes of 149 and 195 bp, whereas the mutant allele shows fragment sizes of 127, 22 and 195 bp. (d) Comparison of amino-acid sequence of ACP33/maspardin among species. A108 is located in the α/β-fold bacterial hydrolase domain, which is highly evolutionally conserved. The G-X-S-X-G motif at the nucleophile elbow is also shown. (e and f) A sagittal T1-weighted image (e) and a transverse fluid-attenuated inversion recovery (FLAIR) image (f) of patient 1 at the age of 70 years show a thin corpus callosum and mildly atrophic cerebrum. Atrophy in the brainstem and cerebellum is not observed. (g–i) A sagittal T1-weighted image (g), a transverse FLAIR image (h) and a coronal FLAIR image (i) of patient 1 at the age of 75 years shows progressive thinning of the corpus callosum mainly in the trunk and progressive atrophy of the cerebrum, which is marked in the frontal and temporal lobes. Slight white matter changes are observed around the lateral ventricles. Atrophy in the brainstem and cerebellum is not observed. (j) 123I-N-isopropyl-p-iodoamphetamine single-photon emission computed tomography (SPECT) at the age of 75 years shows decreased blood flow in the frontal and temporal cortices. Wt, wild type; mut, mutant; Het, heterozygote; Homo, homozygote.
This family is the first family with SPG21 identified outside the Amish population.25 Intriguingly, compared with the original Mast syndrome family with an SPG21 mutation, the ages at onset of HSP symptoms in the patients in the new SPG21 family were strikingly late. Although characteristics such as cognitive decline and a thin corpus callosum were shared in common, characteristic clinical signs in Mast syndrome such as bulbar, extrapyramidal and cerebellar signs were not found in the new family (Supplementary Table S2), thus presenting dissimilar phenotypes. Because the mutation detected in the new family is a missense mutation (p.A108P) next to the active site of the alpha/beta-hydrolase domain (S109), dysfunction of alpha/beta-hydrolase activity of maspardin seems to be related to pathogenicity.
SPG31
The two novel mutations in REEP1 were a frameshift (insertion of A) and a large deletion (Table 2), suggesting haploinsufficiency as the disease-causing mechanism. A large deletion detected in the study demanded a screening of rearrangement of REEP1 in the diagnosis of SPG31. These two patients with SPG31 had pure-form HSP and their disease started in their early teens, compatible with previous reports.9, 21
Sporadic HSP
As much as 11.1% (7/63) of the patients with sporadic HSP were revealed to have mutations in the genes for monogenic diseases. Among sporadic pure-form HSP patients, 12.9% (4/31) had SPAST mutations, and 6.3% (2/32) of sporadic complicated-form HSP patients had SPG11 mutations.
Discussion
We herein described a comprehensive mutational analysis of as many as 16 causative genes of HSP and applied it to the mutational analysis of 129 Japanese HSP patients. An epidemiological study26 based on the Registry of the Ministry of Health, Labour and Welfare, Japan in 2002 reported about 500 HSP patients. Although there remains a possibility that some patients may have not been registered for various reasons, the collection of 129 patients should represent a substantial proportion of Japanese HSP patients. In the 129 HSP patients, we identified 49 mutations, 32 of which were novel. Resequencing microarray and aCGH analyses were proved to be efficacious methods to detect nucleotide substitutions and large duplications/deletions, respectively. Indeed, the fact that we did not find additional base substitution mutations of SPAST and REEP1 in AD-HSP patients by direct sequence analysis, for whom mutations were not detected by resequencing microarrays, indicates a false-negative rate of resequencing microarray analysis was low, if any, by tuning up by our algorithm (a computer program). However, note also that both resequencing microarray and aCGH analyses did not detect small insertion/deletion mutations, and direct nucleotide sequence analysis was needed to detect them. Our results revealed that the combination of these technologies, including resequencing microarray, aCGH, and direct nucleotide sequence analyses, are essential to detect various kinds of mutations, including base substitutions, and insertions/deletions of various sizes with high sensitivities.
Given the results of this study, we propose an algorithm for a comprehensive mutational analysis for HSP. To analyze genes that have relatively frequent small insertion/deletion mutations (for example, SPAST, REEP1, SPG11 and SPG15), direct nucleotide sequence analysis is the first priority. To analyze genes in which most of the mutations are nucleotide substitutions (for example, ATL1, NIPA1, KIF5A, KIAA0196, HSPD1 and BSCL2), resequencing microarray analysis is highly suitable. Considering the throughput, direct nucleotide sequence analysis becomes more laborious as the number of exons to be sequenced increases. In contrast, it is not the case for resequencing microarray and CGH array analyses. That is, the time required for analysis remains constant with increasing number of genes or exons to be sequenced until a limit determined by the structure of arrays. We propose a strategy of utilizing high-throughput microarray techniques and minimizing the use of time-consuming direct nucleotide sequence analysis considering the molecular epidemiology and the mutation types in individual genes (Figure 6). Although there remains a possibility that uncommon mutations (for example, insertions/deletions of intermediate length) or uncommon presentation (for example, SPAST mutation in a family having apparently autosomal recessive mode of inheritance or SPG11 mutations in a pseudoautosomal dominant family) are missed and it might introduce some bias, the algorithm should be highly useful for the efficient identification of the majority, if not all, of the mutations responsible for HSP.
Proposed algorithm for comprehensive mutational analysis of HSP genes. Considering the types and frequencies of mutations in individual SPG genes, we propose an efficaceous strategy for a large-scale mutational analysis of HSP at the time of the study. In patients with ADHSP patients and in familial pure HSP patients with an unknown mode of inheritance, direct nucleotide sequence analysis of spastin and REEP1 followed by CGH analysis is recommended, considering the relatively high frequency of small insertions/deletions in spastin and REEP1 and large deletions/duplications in spastin. In patients with thin corpus callosum and/or cognitive dysfunction, SPG11 and SPG15 should be analyzed first. Next step is CGH analysis followed by resequencing microarray analysis, because throughput of CGH analysis is higher than that of resequencing microarray analysis. *In these days, these stages can be replaced by whole genome or exome sequencing. Direct seq., direct nucleotide sequence analysis.
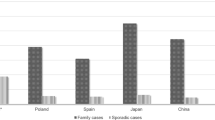
Utilizing the technologies, we elucidated molecular epidemiology of HSP in the Japanese population. Interestingly, the study revealed that the overall trend of molecular epidemiology of AD-HSP/AR-HSP in the Japanese population is similar to those in the Caucasian populations reported previously.3, 5, 6, 20, 21, 27 In contrast, considerable differences in the epidemiology of spinocerebellar ataxias26 or amyotrophic lateral sclerosis (especially in those who have hexanucleotide repeat expansion mutation in C9ORF72)28, 29 have been demonstrated, which presumably reflect founder effects.29, 30 Thus, the similarity in the molecular epidemiology of HSP irrespective of ethnicity suggests that contribution of founder effects is limited in HSP.
We did not find causative mutations in 16 AD-HSP, 8 AR-HSP and 5 familial HSP patients. Although we cannot completely exclude the possibility of false-negative results in our analyses, we assume that these undiagnosed patients would have mutations in causative genes that have recently been identified after the study (RTN2 or GBA2, for example) or mutations in as yet unidentified causative genes.
The extent to which mutations of causative genes account for apparently sporadic HSP is an important but unsolved issue. We found that 7 out of the 62 sporadic HSP patients (11.1%) had mutations of genes for HSP. In particular, we found that SPG4 and SPG11 are relatively frequent in sporadic pure-form HSP and complicated-form HSP patients, respectively. The findings indicate that careful genetic counseling of such patients and families will be required.
With recent progresses in massively parallel sequencing technologies, exome and targeted sequencing are now becoming a robust method for high-throughput resequencing analysis at a relatively reasonable cost.31, 32, 33, 34 Detection of large insertion/deletion mutations based on the short reads generated by next-generation sequencers, however, is still a challenging task. It is of note that a substantial proportion (7/49, 14.3%) of mutations found in the study were insertions/deletions detected by aCGH analysis. Thus, combining multiple technologies, as we did in the study, is indispensable to detect as many mutations as possible even in the next-generation sequencer era. In addition, information on the relative frequencies of HSP types and on the distribution of various types of mutations in each HSP gene as shown in the study is helpful for making strategies for mutational analyses.
In summary, we elucidated the molecular epidemiology of HSP in the Japanese population combining multiple technologies of resequencing microarray, aCGH and Sanger sequencing. The study contributed to further broadening the clinical and mutational spectra of HSP.
References
Fink, J. K. The hereditary spastic paraplegias: nine genes and counting. Arch. Neurol. 60, 1045–1049 (2003).
Finsterer, J., Löscher, W., Quasthoff, S., Wanschitz, J., Auer-Grumbach, M. & Stevanin, G. Hereditary spastic paraplegias with autosomal dominant, recessive, X-linked, or maternal trait of inheritance. J. Neurol. Sci. 318, 1–18 (2012).
Fonknechten, N., Mavel, D., Byrne, P., Davoine, C. S., Cruaud, C., Bönsch, D. et al. Spectrum of SPG4 mutations in autosomal dominant spastic paraplegia. Hum. Mol. Genet. 9, 637–644 (2000).
McDermott, C. J., Burness, C. E., Kirby, J., Cox, L. E., Rao, D. G., Hewamadduma, C. et al. Clinical features of hereditary spastic paraplegia due to spastin mutation. Neurology 67, 45–51 (2006).
Namekawa, M., Ribai, P., Nelson, I., Forlani, S., Fellmann, F., Goizet, C. et al. SPG3A is the most frequent cause of hereditary spastic paraplegia with onset before age 10 years. Neurology 66, 112–114 (2006).
Klebe, S., Lacour, A., Dürr, A., Stojkovic, T., Depienne, C., Forlani, S. et al. NIPA1 (SPG6) mutations are a rare cause of autosomal dominant spastic paraplegia in Europe. Neurogenetics 8, 155–157 (2007).
Elleuch, N., Depienne, C., Benomar, A., Hernandez, A. M., Ferrer, X., Fontaine, B. et al. Mutation analysis of the paraplegin gene (SPG7) in patients with hereditary spastic paraplegia. Neurology 66, 654–659 (2006).
Arnoldi, A., Tonelli, A., Crippa, F., Villani, G., Pacelli, C., Sironi, M. et al. A clinical, genetic, and biochemical characterization of SPG7 mutations in a large cohort of patients with hereditary spastic paraplegia. Hum. Mutat. 29, 522–531 (2008).
Beetz, C., Schüle, R., Deconinck, T., Tran-Viet, K. N., Zhu, H., Kremer, B. P. et al. REEP1 mutation spectrum and genotype/phenotype correlation in hereditary spastic paraplegia type 31. Brain 131, 1078–1086 (2008).
Goizet, C., Boukhris, A., Mundwiller, E., Tallaksen, C., Forlani, S., Toutain, A. et al. Complicated forms of autosomal dominant hereditary spastic paraplegia are frequent in SPG10. Hum. Mutat. 30, E376–E385 (2008).
Warrington, J. A., Shah, N. A., Chen, X., Janis, M., Liu, C., Kondapalli, S. et al. New developments in high-throughput resequencing and variation detection using high density microarrays. Hum. Mutat. 19, 402–409 (2002).
Barrett, M. T., Scheffer, A., Ben-Dor, A., Sampas, N., Lipson, D., Kincaid, R. et al. Comparative genomic hybridization using oligonucleotide microarrays and total genomic DNA. Proc. Natl Acad. Sci. USA 101, 17765–17770 (2004).
Arai, N., Kishino, A., Takahashi, Y., Morita, D., Nakamura, K., Yokoyama, T. et al. Familial cases presenting very early onset autosomal dominant Alzheimer's disease with I143T in presenilin-1 gene: implication for genotype-phenotype correlation. Neurogenetics 9, 65–67 (2008).
Takahashi, Y., Seki, N., Ishiura, H., Mitsui, J., Matsukawa, T., Kishino, A. et al. Development of a high-throughput microarray-based resequencing system for neurological disorders and its application to molecular genetics of amyotrophic lateral sclerosis. Arch. Neurol. 65, 1326–1332 (2008).
Seki, N., Takahashi, Y., Tomiyama, H., Rogaeva, E., Murayama, S., Mizuno, Y. et al. Comprehensive mutational analysis of LRRK2 reveals variants supporting association with autosomal dominant Parkinson’s disease. J. Hum. Genet. 56, 671–675 (2011).
Cutler, D. J., Zwick, M. E., Carrasquillo, M. M., Yohn, C. T., Tobin, K. P., Kashuk, C. et al. High-throughput variation detection and genotyping using microarrays. Genome Res. 11, 1913–1925 (2001).
Mitsui, J., Takahashi, Y., Goto, J., Tomiyama, H., Ishikawa, S., Yoshino, H. et al. Mechanisms of genomic instabilities underlying two common fragile-site-associated loci, PARK2 and DMD, in germ cell and cancer cell lines. Am. J. Hum. Genet. 87, 75–89 (2010).
Maeda-Hashimoto, M., Mitsui, J., Soong, B. W., Takahashi, Y., Ishiura, H., Hayashi, S. et al. Increased gene dosage of myelin protein zero causes Charcot-Marie-Tooth disease. Ann. Neurol. 71, 84–92 (2012).
Silver, J. R. Familial spastic paraplegia with amyotrophy of the hands. Ann. Hum. Genet. 30, 69–75 (1966).
Valdmanis, P. N., Meijer, I. A., Reynolds, A., Lei, A., MacLeod, P., Schlesinger, D. et al. Mutations in the KIAA0196 gene at the SPG8 locus cause hereditary spastic paraplegia. Am. J. Hum. Genet. 80, 152–161 (2007).
Züchner, S., Wang, G., Tran-Viet, K. N., Nance, M. A., Gaskell, P. C., Vance, J. M. et al. Mutations in the novel mitochondrial protein REEP1 cause hereditary spastic paraplegia type 31. Am. J. Hum. Genet. 79, 365–369 (2006).
Beetz, C., Nygren, A. O., Schickel, J., Auer-Grumbach, M., Bürk, K., Heide, G. et al. High frequency of partial SPAST deletions in autosomal dominant hereditary spastic paraplegia. Neurology 67, 1926–1930 (2006).
Orlacchio, A., Babalini, C., Borreca, A., Patrono, C., Massa, R., Basaran, S. et al. SPATACSIN mutations cause autosomal recessive juvenile amyotrophic lateral sclerosis. Brain 133, 591–598 (2012).
Ito, D. & Suzuki, N. Seipinopathy: a novel endoplasmic reticulum stress-associated disease. Brain 132, 8–15 (2009).
Simpson, M. A., Cross, H., Proukakis, C., Pryde, A., Hershberger, R., Chatonnet, A. et al. Maspardin is mutated in mast syndrome, a complicated form of hereditary spastic paraplegia associated with dementia. Am. J. Hum. Genet. 73, 1147–1156 (2003).
Tsuji, S., Onodera, O., Goto, J. & Nishizawa, M., Study Group on Ataxic Diseases. Sporadic ataxias in Japan—a population-based epidemiological study. Cerebellum 7, 189–197 (2008).
Stevanin, G., Santorelli, F. M., Azzedine, H., Cutinho, P., Chomilier, J., Denora, P. S. et al. Mutations in SPG11, encoding spatacsin, are a major cause of spastic paraplegia with thin corpus callosum. Nat. Genet. 39, 366–372 (2007).
Ishiura, H., Takahashi, Y., Mitsui, J., Yoshida, S., Kihira, T., Kokubo, Y. et al. C9ORF72 repeat expansion in amyotrophic lateral sclerosis in the Kii peninsula of Japan. Arch. Neurol. 69, 1154–1158 (2012).
Majounie, E., Renton, A. E., Mok, K., Dopper, E. G., Waite, A., Rollinson, S. et al. Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol.. 11, 323–330 (2012).
Cossée, M., Schmitt, M., Campuzano, V., Reutenauer, L., Moutou, C., Mandel, J. L. et al. Evolution of the Friedreich's ataxia trinucleotide repeat expansion: founder effect and premutations. Proc. Natl Acad. Sci. USA 94, 7452–7457 (1997).
Choi, M., Scholl, U. I., Ji, W., Liu, T., Tikhonova, I. R., Zumbo, P. et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc. Natl Acad. Sci. USA 106, 19096–19101 (2009).
Ku, C. S., Cooper, D. N., Polychronakos, C., Naidoo, N., Wu, M. & Soong, R. Exome sequencing: dual role as a discovery and diagnostic tool. Ann. Neurol. 71, 5–14 (2012).
Ishiura, H., Sako, W., Yoshida, M., Kawarai, T., Tanabe, O., Goto, J. et al. The TRK-fused gene is mutated in hereditary motor and sensory neuropathy with proximal dominant involvement. Am. J. Hum. Genet. 91, 320–329 (2012).
Mitsui, J., Matsukawa, T., Ishiura, H., Higasa, K., Yoshimura, J., Saito, T. L. et al. CSF1R mutations identified in three families with autosomal dominantly inherited leukoencephalopathy. Am. J. Med. Genet. B Neuropsychiatr. Genet. 159B, 951–957 (2012).
Lindsey, J. C., Lusher, M. E., McDermott, C. J., White, K. D., Reid, E., Rubinsztein, D. C. et al. Mutation analysis of the spastin gene (SPG4) in patients with hereditary spastic paraparesis. J. Med. Genet. 37, 759–765 (2000).
Falco, M., Scuderi, C., Musumeci, S., Sturnio, M., Neri, M., Bigoni, S. et al. Two novel mutations in the spastin gene (SPG4) found by DHPLC mutation analysis. Neuromuscul. Disord. 14, 750–753 (2004).
Depienne, C., Tallaksen, C., Lephay, J. Y., Bricka, B., Poea-Guyon, S., Fontaine, B. et al. Spastin mutations are frequent in sporadic spastic paraparesis and their spectrum is different from that observed in familial cases. J. Med. Genet. 43, 259–265 (2006).
Meijer, I. A., Hand, C. K., Cossette, P., Figlewicz, D. A. & Rouleau, G. A. Spectrum of SPG4 mutations in a large collection of North American families with hereditary spastic paraplegia. Arch. Neurol. 59, 281–286 (2002).
Patrono, C., Scarano, V., Cricchi, F., Melone, M. A., Chiriaco, M., Napolitano, A. et al. Autosomal dominant hereditary spastic paraplegia: DHPLC-based mutation analysis of SPG4 revealed eleven novel mutations. Hum. Mutat. 25, 506 (2005).
D’Amico, A., Tessa, A., Sabino, A., Bertini, E., Santorelli, F. M. & Servidei, S. Incomplete penetrance in an SPG3A-linked family with a new mutation in the atlastin gene. Neurology 62, 2138–2139 (2004).
Dürr, A., Camuzat, A., Colin, E., Tallaksen, C., Hannequin, D., Coutinho, P. et al. Atlastin1 mutations are frequent in young-onset autosomal dominant spastic paraplegia. Arch. Neurol. 62, 962–966 (2004).
Kim, S. M., Lee, J. S., Kim, S., Kim, H. J., Kim, M. H., Lee, K. M. et al. Novel compound heterozygous mutations of the SPG11 gene in Korean families with hereditary spastic paraplegia with thin corpus callosum. J. Neurol. 256, 1714–1718 (2009).
Acknowledgements
We thank all the patients and their families for their participation in this study. We also thank the many doctors who kindly provided clinical information and the blood and brain samples of the participants. This work was supported in part by KAKENHI (Grant-in-Aid for Scientific Research) on Priority Areas, Innovative Areas, the Global COE program, and Scientific Research (A) from the Ministry of Education, Culture, Sports, Science and Technology of Japan and a Grant-in-Aid for ‘the Research Committee for Ataxic Diseases’ of the Research on Measures for Intractable Diseases from the Ministry of Health, Welfare and Labour, Japan. HI was supported by a Research Fellowship of the Japan Society for the Promotion of Science for Young Scientists.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Ishiura, H., Takahashi, Y., Hayashi, T. et al. Molecular epidemiology and clinical spectrum of hereditary spastic paraplegia in the Japanese population based on comprehensive mutational analyses. J Hum Genet 59, 163–172 (2014). https://doi.org/10.1038/jhg.2013.139
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.139
Keywords
This article is cited by
-
FBXL17/spastin axis as a novel therapeutic target of hereditary spastic paraplegia
Cell & Bioscience (2022)
-
An integrated modelling methodology for estimating global incidence and prevalence of hereditary spastic paraplegia subtypes SPG4, SPG7, SPG11, and SPG15
BMC Neurology (2022)
-
An autopsied case report of spastic paraplegia with thin corpus callosum carrying a novel mutation in the SPG11 gene: widespread degeneration with eosinophilic inclusions
BMC Neurology (2022)
-
A clinical and genetic study of SPG31 in Japan
Journal of Human Genetics (2022)
-
SPG8 mutations in Italian families: clinical data and literature review
Neurological Sciences (2020)