Abstract
To investigate the pathophysiology of Leber’s hereditary optic neuropathy (LHON), a cohort of 1164 Han Chinese subjects with LHON were screened for ND1 G3460A mutation. A total of 295 subjects from 16 Han Chinese families carrying the G3460A mutation underwent a clinical and genetic evaluation and molecular analysis of mitochondrial (mt)DNA. The incidence of G3460A mutation was 1.4% in this cohort of Chinese subjects with LHON. Twenty-seven (20 males/7 females) of 109 matrilineal relatives among 10 Chinese pedigrees carrying this mutation exhibited a wide range of severity and age-at-onset in visual impairment. Penetrances of optic neuropathy ranged from 7.1% to 50%, with the average of 24.5%. The age-at-onset of 27 affected matrilineal relatives varied from 10 to 40 years, with the average of 22 years. Molecular analysis identified the homoplasmic G3460A mutation and distinct sets of variants belonging to eight haplogroups. Haplogroup M with G3460A mutation was of higher frequency than those in controls. The penetrances of visual loss in families carrying mitochondrial DNA haplogroups A, B and M were higher than those in other families. Furthermore, haplogroup-specific variants tRNASer(AGY) A12223G, tRNAThr G15927A and tRNAGlu A14693G may enhance the penetrance of visual loss in these families. The G3460A mutation occurred through recurrent origins and founder events in Chinese population. Mitochondrial modifiers may modulate the penetrance and expressivity of optic neuropathy among Chinese pedigrees carrying the G3460A mutation. Thus, our findings may provide new insights into the understanding of pathophysiology and valuable information on the management of LHON.
Similar content being viewed by others
Introduction
Leber’s hereditary optic neuropathy (LHON) is a maternally inherited eye disorder that generally affects children to young adults with the rapid, painless, bilateral loss of central vision.1, 2, 3 Mutations in mitochondrial DNA (mtDNA) are associated with this disorder.4, 5, 6 Of these, three primary mutations such as ND1 G3460A, ND4 G11778A and ND6 T14484C, which alter genes encoding the subunits of respiratory chain complex I (NADH dehydrogenase), are responsible for ∼90% of LHON pedigrees in some countries.7, 8, 9, 10 The LHON-associated mtDNA mutation(s) often occurred in nearly homoplasmy or homoplasmy. This hinted mild nature of mutations, evidenced by that fact relatively mild mitochondrial dysfunction was observed in mutant cells carrying one of these mutations.11, 12, 13 Typical features in LHON pedigrees carrying the mtDNA mutation(s) are incomplete penetrance and male bias among the affected subjects, reflecting the complex etiology of this disease.14, 15 In particular, matrilineal relatives within and among families carrying the same mtDNA mutation(s) exhibited a wide range of penetrance and expressivity including severity, age-of-onset and progression in visual impairment. These suggest that these mutations are the primary causative evident, but the secondary events such as environmental factors, nuclear and mitochondrial genetic modifiers are necessary for the manifestation of the optic neuropathy.6, 15, 16 For example, a predominance of male patients presenting with vision loss suggests an X-linked modifier gene for the phenotypic manifestation of the mtDNA mutations.17 Furthermore, a group of so-called ‘secondary’ LHON-associated mtDNA mutations was implicated to act in synergy with these primary mtDNA mutations.18, 19, 20 In addition, mtDNA haplogroups influenced the phenotypic manifestation of the primary mtDNA mutations.21, 22, 23, 24
However, the role of these genetic modifiers, especially mtDNA haplogroups/variants, in the phenotypic expression of these primary mtDNA mutations remains poorly defined. In the previous investigation, we showed that the ND1 T3394C, ND4 G11696A, ND6 T14502C, tRNAMet A4435G and tRNAThr A15951G mutations contributed to the high penetrance and expressivity of optic neuropathy in Chinese families carrying the G11778A mutation.19, 25, 26, 27, 28 In the present investigation, we carried out a systematic and extended mutational screening of ND1 G3460A mutation in a large cohort of 1164 Han Chinese subjects with LHON. This analysis identified 16 subjects harboring the G3460A mutation. We then performed the clinical, genetic and molecular characterization of these affected subjects carrying the G3460A mutation. A wide range of penetrance, severity and age-at-onset of visual loss was observed in the matrilineal relatives within and among these Chinese families. To assess the contribution that mtDNA variants or haplogroups make toward the variable penetrance and expressivity of visual loss in these pedigrees, we performed a PCR amplification of fragments spanning entire mtDNA and subsequent DNA sequence analysis in the matrilineal relatives of those families. These analyses showed that there were distinct sets of mtDNA variants belonging to eight Eastern Asian haplogroups in these Chinese pedigrees carrying the G3460A mutation. Furthermore, we evaluated the potential role of these mtDNA haplogroups and variants in the phenotypic manifestation of the G3460A mutation in these Chinese families.
Materials and methods
Patients and subjects
We ascertained 10 Han Chinese families (Figure 1) through the Eye clinics of Wenzhou Medical College, Zhejiang, Dongfang Hospital, Beijing and Xingtai Eye Hospital, Hebei. Informed consent, blood samples and clinical evaluations were obtained from all participating family members, under protocols approved by the Zhejiang University Institute Review Board and the Wenzhou Medical University Ethic Committee. Members of these pedigrees were interviewed at length to identify both personal or family medical histories of visual impairments, and other clinical abnormalities.
Ophthalmological examinations
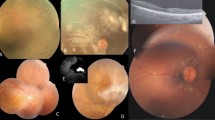
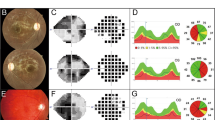
The ophthalmic examinations of the proband and other members of these families were conducted, including visual acuity, visual field examination (Humphrey Visual Field Analyzer IIi, SITA Standard; Carl Zeiss Meditec, Oberkochen, Germany), visual evoked potentials (RETI port gamma, flash visual evoked potentials; Roland Consult, Brandenberg, Germany) and fundus photography (CR6-45NM fundus camera; Canon, Lake Success, NY, USA). The degree of visual impairment was defined according to the visual acuity as follows: normal >0.3, mild=0.3–0.1; moderate <0.1–0.05; severe <0.05–0.02; and profound <0.02.
Mutational analysis of the mitochondrial genome
Genomic DNA was isolated from whole blood of participants using the Puregene DNA Isolation Kits (Puregene DNA Isolation Kit; Gentra Systems, Minneapolis, MN, USA). The presence of the G3460A mutations was examined as detailed elsewhere.7 Briefly, the probands’ DNA fragments carrying the G3460A mutation was amplified by PCR using oligodeoxynucleotides corresponding to mtDNA at positions 3108–3717. The amplified PCR segments were digested with a restriction enzyme BsaHI as the G3460A mutation creates the site for this restriction enzyme.7 Equal amounts of various digested samples were then analyzed by electrophoresis through 7% polyacrylamide gel. The proportions of digested and undigested PCR product were determined by the ImageQuant program after ethidium bromide staining to determine if the G3460A mutation is in the homoplasmy in these subjects.
The entire mitochondrial genomes of 10 probands were PCR amplified in 24 overlapping fragments using sets of the light (L)-strand and the heavy (H)-strand oligonucleotide primers as described previously.29 Each fragment was purified and subsequently analyzed by direct sequencing in an ABI 3700 automated DNA sequencer using the Big Dye Terminator Cycle sequencing reaction kit. These sequence results were compared with the updated consensus Cambridge sequence (GenBank accession number: NC_012920).30
Phylogenetic analysis
A total of 17 vertebrate mtDNA sequences were used in the interspecific analysis. These include the following: Bos Taurus, Cebus albifrons, Gorilla gorilla, Homo sapiens, Hylobates lar, Lemur catta, Macaca mulatta, Macaca sylvanus, Mus musculus, Nycticebus coucang, Pan paniscus, Pongo pygmaeus, Pongo abelii, Papio hamadryas, Tarsius bancanus and Xenopus laevis (Genbank) (Supplementary Table 1). The conservation index was calculated by comparing the human mtDNA variants with other 16 vertebrates.
Haplogroup analyses
The entire mtDNA sequences of the 16 Chinese probands carrying the G3460A mutation were assigned to the Asian mitochondrial haplogroups by using the nomenclature of mitochondrial haplogroups.31, 32
Statistical analysis
Statistical analysis was carried out using the Student’s unpaired, two-tailed t-test contained in the Microsoft-Excel program. Unless indicated otherwise, a P-value <0.05 was considered statistically significant.
Results
Mutational screening of ND1 G3460A mutation in a cohort of Chinese subjects with LHON
To further elucidate the molecular basis of visual loss, we have performed a mutational screening of the G3460A mutation in a cohort of 1164 Han Chinese subjects, who were diagnosed as LHON by the several Eye Clinics in China. First, 609 bp DNA fragments spanning the G3460A mutation were PCR amplified from each affected subject. Each fragment was digested by restriction enzyme MvaI and subsequent electrophoresis analysis. Of those, 16 subjects harbored the homoplasmic G3460A mutation (data not shown). These translate to a frequency of ∼1.4% for the G3460A mutation in this cohort. The presence of the homoplasmic G3460A mutation in those subjects was further confirmed by PCR -amplification of fragments spanning the ND1 gene and subsequent DNA sequence analysis (data not shown).
Clinical data for 16 Chinese probands carrying the G3460A mutation were summarized in Table 1. These consisted of three females and 13 males. Comprehensive medical histories of those probands showed no other clinical abnormalities, including diabetes, muscular diseases, hearing dysfunction and neurological disorders. These subjects exhibited a variety of severity and age-of-onset of visual impairment. Among these, three individuals suffered from profound visual impairment, six subjects exhibited severe visual impairment, four probands had moderate visual impairment and three subjects exhibited mild visual impairment.
Clinical and genetic evaluation of Chinese families carrying the G3460A mutation
In the previous investigations, we have performed the clinical, genetic and molecular characterization of six Chinese pedigrees carrying the G3460A mutation.33, 34 In the current study, a comprehensive history and physical examination as well as ophthalmological examination were performed to identify both personal and family medical histories of visual impairments, and other clinical abnormalities in all available members of another 10 Han Chinese pedigrees carrying the G3460A mutation. Comprehensive family medical histories of those probands and other members of these Chinese families showed no other clinical abnormalities, including diabetes, muscular diseases, hearing impairment and neurological disorders.
As shown in Figure 1 and Table 2, 27 (20 males/7 females) of 109 matrilineal relatives among 10 Chinese pedigrees exhibited the bilateral visual impairment. Ophthalmological examination showed a variable severity of visual impairment in the matrilineal relatives of these families, ranging from profound visual loss, to severe visual impairment, to moderate visual impairment, to mild visual impairment, to completely normal vision. Furthermore, there was a wide range in the age-at-onset of visual impairment in these families. As shown in Table 2, the average age-at-onset of each family in these 10 pedigrees varied from 10 years to 40 years, with the average of 22 years. In addition, there was a wide range of the penetrance of visual loss among these pedigrees. In particular, the penetrances of visual loss in these pedigrees ranged from 7.1% to 50%, with the average of 24.5%.
Analysis of the complete mitochondrial genomes
To assess the contribution that mtDNA variants or haplogroups make toward the variable penetrance and expressivity of optic neuropathy in these Chinese pedigrees, we performed a PCR amplification of fragments spanning entire mtDNA and subsequent DNA sequence analysis in 10 probands carrying the G3460A mutation. The sequence results from these Chinese subjects were aligned with the updated consensus Cambridge sequence.30 In addition to the G3460A mutation, these probands exhibited distinct sets of mtDNA polymorphisms including a number of known variants and six novel variants (Supplemental Table 2). These included 60 known variants in the D-loop region, nine known variants in the 12S rRNA gene, five (one novel and four known) variants in the 16S rRNA gene, the known tRNALeu(UUR) T3290C, tRNAGln T4336C, tRNAGlu T14427C, tRNAThr G15927A and novel tRNASer(AGY) A12223G mutations, as well as 79 (4 novel/75 known) silent variants and 33 known missense mutations in the polypeptide encoding genes. These variants in RNAs and polypeptides were further evaluated by phylogenetic analysis of these variants and sequences from other 16 organisms including mouse,35 bovine36 and Xenopus laevis.37 As shown in Table 3, the known A6 G9010A (A162T) and CO3 C9490T (A95V) variants, as well as novel tRNASer(AGY) A12223G variant showed the high conservation in these species, proposed by Wallace6 and had <1% frequency of 2704 mtDNAs. By contrast, none of other variants showed evolutionary conservation.
Phylogenetic and haplogroup analysis of subjects carrying the G3460A mutation
As shown in Table 4, the mtDNAs from 16 Chinese families carrying the G3460A mutation were distributed among eight different haplogroups. To determine if this distribution corresponding to that of general Chinese population, 104 vision normal Chinese control mtDNAs recruited from the same region were sequenced and assigned to certain Asian haplogroups. Here, all mtDNA lineages fall into two macro-haplogroups M and N, of which Eastern Asian mtDNAs were subdivided.31, 32 Indeed, the G3460A mutation is widely dispersed among 10 common Eastern Asian sub-haplogroups. As shown in Table 4, the frequencies of mtDNA haplogroups A, B, C, D, F, H2, M and R in 16 LHON families were 6.3%, 12.5%, 6.3%, 18.8%, 12.5%, 6.3%, 31.3% and 6.3%, respectively, while those of 104 Chinese controls were 10.6%, 19.2%, 2.9%, 24%, 18.3%, 2.9%, 12.5% and 0%, respectively. Notably, the haplogroup M accounted for 31.3% of the patient’s mtDNA samples but only 12.5% of the Chinese control mtDNA samples in this study and 12.2% of a large cohort of Eastern Asian population.31 Of other haplogroups, the frequency of haplogroups C, H2 and R in the patients’ mtDNA samples were higher than those of control’s mtDNA samples, whereas the frequencies of haplogroups A, B and D in vision-impaired mtDNA samples were lower than those of control mtDNA samples.
Statistical analysis
As shown in Table 4, the average penetrances of optic neuropathy among LHON families carrying mtDNA haplogroups A, B, C, D, F, H2, M and R were 30%, 29.2%, 11.1%, 18.9%, 9.1%, 12.5%, 29.5% and 10.7%, with the average penetrances of 20.9%. Furthermore, the average age-at-onset of optic neuropathy among LHON families carrying mtDNA haplogroups A, B, C, D, F, H2, M, and R were 28.7, 20.9, 20, 21.1, 20, 14, 19.6 and 18 years, with the average age-at-onset of 20.5 years. To further evaluate the role of mitochondrial genetic modifiers in phenotypic expression, statistical analysis was carried out using the unpaired, two-tailed Student’s t-test. The difference of average penetrances (46.5%) of optic neuropathy among the Chinese pedigrees carrying the additional mtDNA variants such as A14693G mutation showed significantly higher than those (17.5%) in other pedigrees lacking the significant mtDNA variants (P=0.008).
Discussion
In the present study, we have performed the clinical, genetic and molecular characterization of 16 Han Chinese families with LHON. Optic neuropathy as a sole clinical phenotype was only present in the maternal lineage of these pedigrees carrying the homoplasmic ND1 G3460A mutation. The incidence of the G3460A mutation in a cohort of 1164 Chinese subjects with LHON was ∼1.4%, whereas the incidences of the mutation in a cohort of 903 LHON Chinese subjects and two cohorts of white subjects with LHON were 0.4% and 13%, respectively.38, 39, 40 A wide range of penetrance and expressivities of optic neuropathy was observed in these Chinese families. The penetrances of optic neuropathy (affected matrilineal relatives/total matrilineal relatives) in 16 Chinese pedigrees ranged from 2.8 to 50%, with the average of 21.2%. On the other hand, 50% males and ∼10% females in Caucasians carrying the G3460A, G11778A or T14484C mutation indeed developed the optic neuropathy.2, 41, 42 Furthermore, the average age-of-onset for optic neuropathy ranged from 10 to 40 years, with the average of 20.5 years in these 16 Chinese families. On the contrary, the average age-of-onset for optic neuropathy was 20 and 29 years in eight and night Caucasian families, respectively.43, 44 Unlike previous reports that the ratios between affected male and female matrilineal relatives were 2.3:1 and 4.3:1 from two cohorts of Caucasian pedigrees carrying the G3460A mutation,43, 44 26 males/11 females of 222 matrilineal relatives in these 16 families exhibited visual impairment.
The variable penetrance and expressivity of visual impairment among these pedigrees carrying the G3460A mutation indicated the involvement of modifier factors. The nuclear background apparently contributed to the phenotypic variability of matrilineal relatives within and among these Chinese families, as described for other pedigrees.15, 42, 45 In fact, the X-linked or autosomal recessive genes such as PARL were implicated to have a role in phenotypic expression of LHON.17, 46 Furthermore, it is possible that environmental factors may also contribute to the phenotypic variability of visual loss in matrilineal relatives of these families.47 The mitochondrial variants/haplotypes have been shown to influence the penetrance and expressivity of visional loss associated with primary mtDNA mutations including G11778A and T14484C mutations.20, 22, 23, 24 In particular, secondary LHON mutations at positions 4216 and 13708 may increase the penetrance and expressivity of LHON associated with the primary LHON mtDNA mutations including G11778A and T14484C.18, 21, 48 Furthermore, our previous investigation showed that the ND1 T3394C, ND4 G11696A, ND6 T14502C, tRNAMet A4435G and tRNAThr A15951G mutations may modulate the phenotypic manifestation of Chinese families with the ND4 G11778A mutation.19, 25, 26, 27, 28
There were distinct sets of mtDNA polymorphisms despite carrying the G3460A mutation in their genomes of 16 Chinese pedigrees. As shown in Table 4, their mtDNAs were distributed among the eight Eastern Asian haplogroups A, B, C, D, F, H2, M and R,31, 32 while mtDNAs of European pedigrees carrying the G3460A mutation belonged to the common European haplogroups H, J and K.5 The presence of the G3460A mutation on various mitochondrial backgrounds suggested that the G3460A mutation occurred sporadically and multiplied through the evolution of the mtDNA in China, as in the cases of Caucasian families.5 Strikingly, the occurrence of haplogroup M-mtDNA carrying the G3460A mutation in LHON patients is much significantly higher frequency than these in controls lacking this mutation.31, 32 The G3460A mutation in the haplogroup M background would have been transmitted by descents to families that are indeed maternally related. In contrast, the lower frequency of haplogroups A and B in these Chinese families suggested that the G3460A mutation on these mitochondrial backgrounds would be the independent mutational events. Thus, the G3460A mutation occurred through recurrent origins and founder events in these Chinese families.
In the present investigation, there was the variable age-of-onset of visual loss on Chinese families on mitochondrial haplogroups. However, this difference was not statistical significance, suggesting that mitochondrial haplogroups may not have an important role in the age-at-onset of visual loss in these Chinese families. However, Chinese pedigrees on mtDNA haplogroups A, B and M exhibited higher penetrance than the average penetrance of visual loss in 16 pedigrees carrying the G3460A mutation. In particular, the haplogroups M7b-specific variant tRNAGlu A14693G, and B5b2-specific variants tRNASer(AGY) A12223G and tRNAThr G15927A likely increased penetrance of optic neuropathy in these families. The A14693G mutation occurs at the first base (conventional position 54) of the TψC-loop of tRNAGlu, the G15927A mutation resides at a highly conserved nucleotide (G42) on the anticodon stem of tRNAThr, and the novel A12223G mutation locates at a highly conserved nucleotide (A31) on the anticodon stem of tRNASe(AGY).49 It is likely that these mutations altered the secondary structure and function of tRNAs, thereby leading to mitochondrial dysfunction. The A4693G and G15927A mtDNA mutations have been shown to influence the phenotypic manifestation of deafness-associated 12S rRNA mutation or LHON-associated ND1 G3460A mutation.33, 50 Therefore, these secondary mtDNA mutations may worsen mitochondrial dysfunctions caused by the G3460A mutation, thereby increasing the penetrance and expressivity of visual loss in these Chinese pedigrees, as in the case of other secondary LHON mutations.25, 26
In summary, our data showed that the incidence of the G3460A mutation was 1.4% in a cohort of 1164 Chinese subjects with LHON. A wide range of penetrances and expressivities of visual loss was observed among 16 Chinese families carrying the G3460A mutation. In particular, the average penetrance of visual loss in these pedigrees was 21.2%. Their mtDNA genomes belonged to eight common Eastern Asian haplogroups, indicating that the G3460A mutation occurred through recurrent origins and founder events in Chinese population. Notably, the mitochondrial haplogroups A, B and M may increase the penetrance of visual loss in these Chinese families. Furthermore, haplogroup-specific variants tRNASer(AGY) A12223G, tRNAThr G15927A and tRNAGlu A14693G may enhance the penetrance of visual loss in these Chinese families. These provided the evidence that mitochondrial genetic modifiers may contribute to the variable penetrance and expressivity of visual loss among Chinese pedigrees carrying the G3460A mutation.
Accession codes
References
Newman, N. J. Leber’s hereditary optic neuropathy. New genetic considerations. Arch. Neurol. 50, 540–548 (1993).
Nikoskelainen, E. K. Clinical picture of lhon. Clin. Neurosci. 2, 115–120 (1994).
Man, P. Y., Griffiths, P. G., Brown, D. T., Howell, N., Turnbull, D. M. & Chinnery, P. F. The epidemiology of Leber hereditary optic neuropathy in the North East of England. Am. J. Hum. Genet. 72, 333–339 (2003).
Wallace, D. C., Singh, G., Lott, M. T., Hodge, J. A., Schurr, T. G., Lezza, A. M. et al. Mitochondrial DNA mutation associated with Leber's hereditary optic neuropathy. Science 242, 1427–1430 (1988).
Howell, N., Oostra, R. J., Bolhuis, P. A., Spruijt, L., Clarke, L. A., Mackey, D. A. et al. Sequence analysis of the mitochondrial genomes from Dutch pedigrees with Leber hereditary optic neuropathy. Am. J. Hum. Genet. 72, 1460–1469 (2003).
Wallace, D. C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: a dawn for evolutionary medicine. Annu. Rev. Genet. 39, 359–407 (2005).
Brown, M. D., Torroni, A., Reckord, C. L. & Wallace, D. Phylogenetic analysis of Lebers hereditary optic neuropathy mitochondrial DNAs indicates multiple independent occurrences of the common mutations. Hum. Mutat. 6, 311–325 (1995).
Mackey, D. A., Oostra, R. J., Rosenberg, T., Nikoskelainen, E., Poulton, J., Barratt, T. et al. Primary pathogenic mtDNA mutations in multigeneration pedigrees with Leber hereditary optic neuropathy. Am. J. Hum. Genet. 59, 481–485 (1996).
Mashima, Y., Yamada, K., Wakakura, M., Kigasawa, K., Kudoh, J., Shimizu, N. et al. Spectrum of pathogenic mitochondrial DNA mutations and clinical features in Japanese families with Leber’s hereditary optic neuropathy. Cur. Eye Res. 17, 403–408 (1998).
Carelli, V., La Morgia, C., Valentino, M. L., Barboni, P., Ross-Cisneros, F. N. & Sudan, A. A. Retinal ganglion cell neurodegeneration in mitochondrial inherited disorders. Biochim. Biophys. Acta 1787, 518–528 (2009).
Hofhaus, G., Johns, D. R., Hurko, O., Attardi, G. & Chomyn, A. Respiration and growth defects in transmitochondrial cell lines carrying the 11778 mutation associated with Leber's hereditary optic neuropathy. J. Biol. Chem. 271, 13155–13161 (1996).
Brown, M. D., Trounce, I. A., Jun, A. S., Attardi, G. & Chomyn, A. Functional analysis of lymphoblast and cybrid mitochondria containing the 3460, 11778, or 14484 Leber's hereditary optic neuropathy mitochondrial DNA mutation. J. Biol. Chem. 275, 39831–39836 (2000).
Qian, Y., Zhou, X., Liang, M., Qu, J. & Guan, M. X. The altered activity of complex III may contribute to the high penetrance of Leber's hereditary optic neuropathy in a Chinese family carrying the ND4 G11778A mutation. Mitochondrion 11, 871–877 (2011).
Riordan-Eva, P., Sanders, M. D., Govan, G. G., Sweeney, M. G., Da Costa, J., Harding, A. E. et al. The clinical features of Leber's hereditary optic neuropathy defined by the presence of a pathogenic mitochondrial DNA mutation. Brain 118 (Pt 2), 319–337 (1995).
Yu-Wai-Man, P., Griffiths, P. G., Hudson, G. & Chinnery, P. F. Inherited mitochondrial optic neuropathies. J. Med. Genet. 46, 145–158 (2009).
Yen, M. Y., Wang, A. G. & Wei, Y. H. Leber’s hereditary optic neuropathy: a multifactorial disease. Prog. Retin. Eye Res. 25, 381–396 (2006).
Hudson, G., Keers, S., Yu Wai Man, P., Griffiths, P., Huoponen, K., Savontaus, M. L. et al. Identification of an X-chromosomal locus and haplotype modulating the phenotype of a mitochondrial DNA disorder. Am. J. Hum. Genet. 77, 1086–1091 (2005).
Torroni, A., Petrozzi, M., Durbano, L., Sellitto, D., Zeviani, M., Carrara, F. et al. Haplotype and phylogenetic analyses suggest that one european-specific mtdna background plays a role in the expression of Leber hereditary optic neuropathy by increasing the penetrance of the primary mutations 11778 and 14484. Am. J. Hum. Genet. 60, 1107–1121 (1997).
Qu, J., Li, R., Zhou, X., Tong, Y., Lu, F., Qian, Y. et al. The novel A4435G mutation in the mitochondrial tRNAMet may modulate the phenotypic expression of the LHON-associated ND4 G11778A mutation. Invest. Ophthalmol. Vis. Sci. 47, 475–483 (2006).
Ji, Y., Zhang, A. M., Jia, X., Zhang, Y. P., Xiao, X., Li, S. et al. Mitochondrial DNA haplogroups M7b1'2 and M8a affect clinical expression of Leber hereditary optic neuropathy in Chinese families with the m.11778G>A mutation. Am. J. Hum. Genet. 83, 760–768 (2008).
Brown, M. D., Starikovskaya, E., Derbeneva, O., Hosseini, S., Allen, J. C., Mikhailovskaya, I. E. et al. The role of mtdna background in disease expression: a new primary LHON mutation associated with Western Eurasian haplogroup J. Hum. Genet. 110, 130–138 (2002).
Hudson, G., Carelli, V., Spruijt, L., Gerards, M., Mowbray, C., Achilli, A. et al. Clinical expression of Leber hereditary optic neuropathy is affected by the mitochondrial DNA-haplogroup background. Am. J. Hum. Genet. 81, 228–233 (2007).
Pello, R., Martin, M. A., Carelli, V., Nijtmans, L. G., Achilli, A., Pala, M. et al. Mitochondrial DNA background modulates the assembly kinetics of OXPHOS complexes in a cellular model of mitochondrial disease. Hum. Mol. Genet. 17, 4001–4011 (2008).
Kaewsutthi, S., Phasukkijwatana, N., Joyjinda, Y., Chuenkongkaew, W., Kunhapan, B. & Tun, A. W. Mitochondrial haplogroup background may influence Southeast Asian G11778A Leber hereditary optic neuropathy. Invest. Ophthalmol. Vis. Sci. 52, 4742–4748 (2011).
Li, R., Qu, J., Zhou, X., Tong, Y., Hu, Y., Qian, Y. et al. The mitochondrial tRNAThr A15951G mutation may influence the phenotypic expression of the LHON-associated ND4 G11778A mutation in a Chinese family. Gene 376, 79–86 (2006).
Qu, J., Li, R., Zhou, X., Tong, Y., Yang, L., Chen, J. et al. Cosegregation of the ND4 G11778A mutation with the LHON-associated ND4 G11778A mutation in a four generation Chinese family. Mitochondrion 7, 140–146 (2007).
Zhang, J., Zhou, X., Zhou, J., Li, C., Zhao, F., Wang, Y. et al. Mitochondrial ND6 T14502C variant may modulate the phenotypic expression of LHON-associated G11778A mutation in four Chinese families. Biochem. Biophys. Res. Commun. 399, 647–653 (2010).
Zhang, M., Zhou, X., Li, C., Zhao, F., Zhang, J., Yuan, M. et al. Mitochondrial haplogroup M9a specific variant ND1 T3394C may have a modifying role in the phenotypic expression of the LHON-associated ND4 G11778A mutation. Mol. Genet. Metab. 101, 192–199 (2010).
Rieder, M. J., Taylor, S. L., Tobe, V. O. & Nickerson, D. A. Automating the identification of DNA variations using quality-based fluorescence re-sequencing: analysis of the human mitochondrial genome. Nucleic Acids Res. 26, 967–973 (1998).
Andrews, R. M., Kubacka, I., Chinnery, P. F., Lightowlers, R. N., Turnbull, D. M., Howell, N. et al. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat. Genet. 23, 147 (1999).
Tanaka, M., Cabrera, V. M., Gonzalez, A. M., Larruga, J. M., Takeyasu, T., Fuku, N. et al. Mitochondrial genome variation in Eastern Asia and the peopling of Japan. Genome Res. 14, 1832–1850 (2004).
Kong, Q. P., Bandelt, H. J., Sun, C., Yao, Y. G., Salas, A., Achilli, A. et al. Updating the East Asian mtDNA phylogeny: a prerequisite for the identification of pathogenic mutations. Hum. Mol. Genet. 15, 2076–2086 (2006).
Tong, Y., Mao, Y., Zhou, X., Yang, L., Zhang, J., Cai, W. et al. The mitochondrial tRNAGlu A14693G mutation may influence the phenotypic manifestation of ND1 G3460A mutation in a Chinese family with Leber's hereditary optic neuropathy. Biochem. Biophys. Res. Commun. 357, 524–530 (2007).
Tong, Y., Sun, Y. H., Zhou, X., Zhao, F., Mao, Y., Wei, Q. P. et al. Very low penetrance of Leber's hereditary optic neuropathy in five Han Chinese families carrying the ND1 G3460A mutation. Mol. Genet. Metab. 99, 417–424 (2010).
Bibb, M. J., Van Etten, R. A., Wright, C. T., Walberg, M. W. & Clayton, D. A. Sequence and gene organization of mouse mitochondrial DNA. Cell 26, 167–180 (1981).
Gadaleta, G., Pepe, G., De Candia, G., Quagliariello, C., Sbisà, E. & Saccone, C. The complete nucleotide sequence of the Rattus norvegicus mitochondrial genome: cryptic signals revealed by comparative analysis between vertebrates. J. Mol. Evol. 28, 497–516 (1989).
Roe, B. A., Ma, D. P., Wilson, R. K. & Wong, J. F. The complete nucleotide sequence of the Xenopus laevis mitochondrial genome. J. Biol. Chem. 260, 9759–9774 (1985).
Huoponen, K., Vilkki, J., Aula, P., Nikoskelainen, E. K. & Savontaus, M. L. A new mtDNA mutation associated with Leber hereditary optic neuroretinopathy. Am. J. Hum. Genet. 48, 1147–1153 (1991).
Howell, N., Kubacka, I., Xu, M. & McCullough, D. A. Leber hereditary optic neuropathy - involvement of the mitochondrial ND1 gene and evidence for an intragenic suppressor mutation. Am. J. Hum. Genet. 48, 935–942 (1991).
Jia, X., Li, S., Xiao, X., Guo, X. & Zhang, Q. Molecular epidemiology of mtDNA mutations in 903 Chinese families suspected with Leber hereditary optic neuropathy. J. Hum. Genet. 51, 851–856 (2006).
Brown, M. D. & Wallace, D. C. Spectrum of mitochondrial-DNA mutations in Lebers hereditary optic neuropathy. Clin. Neurosci. 2, 138–145 (1994).
Macmillan, C., Kirkham, T., Fu, K., Allison, V., Andermann, E., Chitayat, D. et al. Pedigree analysis of French Canadian families with T14484C Leber's hereditary optic neuropathy. Neurology 50, 417–422 (1998).
Johns, D. R., Smith, K. H. & Miller, N. R. Leber's hereditary optic neuropathy. Clinical manifestations of the 3460 mutation. Arch. Ophthalmol. 110, 1577–1581 (1992).
Harding, A. E., Sweeney, M. G., Govan, G. G. & Riordan-Eva, P. Pedigree analysis in Leber hereditary optic neuropathy families with a pathogenic mtdna mutation. Am. J. Hum. Genet. 57, 77–86 (1995).
Johns, D. R., Heher, K. L., Miller, N. R. & Smith, K. H. Leber’s hereditary optic neuropathy. Clinical manifestations of the 14484 mutation. Arch. Ophthalmol. 111, 495–498 (1993).
Phasukkijwatana, N., Kunhapan, B., Stankovich, J., Chuenkongkaew, W. L., Thomson, R., Thornton, T. et al. Genome-wide linkage scan and association study of PARL to the expression of LHON families in Thailand. Hum. Genet. 128, 39–49 (2010).
Kirkman, M. A., Yu-Wai-Man, P., Korsten, A., Leonhardt, M., Dimitriadis, K. & De Coo, I. F. Gene-environment interactions in Leber hereditary optic neuropathy. Brain 132, 2317–2326 (2009).
Johns, D. R. & Berman, J. Alternative, simultaneous complex I mitochondrial DNA mutations in Leber's hereditary optic neuropathy. Biochem. Biophys. Res. Commun. 174, 1324–1330 (1991).
Suzuki, T., Nagao, A. & Suzuki, T. Human mitochondrial tRNAs: biogenesis, function, structural aspects, and diseases. Annu. Rev. Genet. 45, 299–329 (2011).
Lu, J., Li, Z., Zhu, Y., Yang, A., Li, R., Zheng, J. et al. Mitochondrial 12S rRNA variants in 1642 Han Chinese pediatric subjects with aminoglycoside-induced and nonsyndromic hearing loss. Mitochondrion 10, 380–390 (2010).
Acknowledgements
This work was supported by National Key Technologies R&D Program Grant 2012BAI09B03 from the Ministry of Science and Technology of China (M-XG and PJ) and a Chinese Young Scholar Award (81200724) from National Science Foundation of China (JZ) and a research award (Y201225490) from the Ministry of Education of Zhejiang Province (YJ).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Ji, Y., Liang, M., Zhang, J. et al. Mitochondrial haplotypes may modulate the phenotypic manifestation of the LHON-associated ND1 G3460A mutation in Chinese families. J Hum Genet 59, 134–140 (2014). https://doi.org/10.1038/jhg.2013.134
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.134
Keywords
This article is cited by
-
Haplogroup Context is Less Important in the Penetrance of Mitochondrial DNA Complex I Mutations Compared to mt-tRNA Mutations
Journal of Molecular Evolution (2018)