Abstract
In the nonrecombining region of the Y-chromosome, there are single-nucleotide polymorphisms (Y-SNPs) that establish haplogroups with particular geographical origins (European, African, Native American, etc.). The complex process of admixture that gave rise to the majority of the current Mexican population (∼93%), known as Mestizos, can be examined with Y-SNPs to establish their paternal ancestry and population structure. We analyzed 18 Y-SNPs in 659 individuals from 10 Mexican-Mestizo populations from different regions of the country. In the total population sample, paternal ancestry was predominately European (64.9%), followed by Native American (30.8%) and African (4.2%). However, the European ancestry was prevalent in the north and west (66.7–95%) and, conversely, Native American ancestry increased in the center and southeast (37–50%), whereas the African ancestry was low and relatively homogeneous (0–8.8%). Although this paternal landscape concurs with previous studies based on genome-wide SNPs and autosomal short tandem repeats (STRs), this pattern contrasts with the maternal ancestry, mainly of Native American origin, based on maternal lineages haplogroups. In agreement with historical records, these results confirm a strong gender-biased admixture history between European males and Native American females that gave rise to Mexican-Mestizos. Finally, pairwise comparisons and analysis of molecular variance tests demonstrated significant population structure (FST=4.68%; P<0.00005), delimiting clusters that were geographically defined as the following: north–west, center–south and southeast.
Similar content being viewed by others
Introduction
In Mexico, European colonization of the New World gave rise to a complex biological admixture process mainly between Native Americans, Spaniards and African slaves. Mestizos are the result of this process and presently constitute ∼93% of the total Mexican population.1 The complex amalgamation that characterizes the current Mexican population derives from several events that have taken place from the European conquest to the present day. During colonial times, three devastating epidemics of smallpox (1520–1521), measles (1545–1548) and typhus (1576–1581) drastically diminished the indigenous population in the 16th century.2 Although Native population density was recovered some time later, an important amount of communities and ethnic groups disappeared completely. Another event contributing to the admixture was the industrial development promoted by discovery of silver mines in Northern Mexico from the 16th to the 18th century. This encouraged foreign settlements throughout the Mexican territory and, consequently, the Mestizo population has increased significantly over the last 500 years.2, 3 Additionally, in recent years, immigration from rural to urban centers has increased regional differences of admixture, principally increasing Native American ancestry in urban centers.3 The resulting high genetic diversity has been evaluated by means of genome-wide analyses both in Mexican and in Latin-American populations.4, 5, 6
Single-nucleotide polymorphisms (SNPs) in the non-recombining region of the human Y-chromosome (NRY) generate paternal lineages or haplogroups that may be related with a specific geographical origin, typically the place where the mutation originated and/or expanded.7 This information allows reconstructing the evolutionary and demographic paternal history of human populations, such as migrations, genetic relationships and admixture components.8 Furthermore, haploid uniparental inheritance of SNPs in the NRY (Y-SNPs) determine a lower effective population size, rendering this more sensitive to genetic drift effects, thus making it easier to analyze population structure.9 In America, four Y-chromosome lineages have been described in Native groups: (1) the most common is Q1a3a* (QM3), followed by (2) the ancestral Q1* (P36), and two rare lineages, including (3) C* (RPS4Y) located in North America, and (4) Q1a3a1 (M19), observed in South America.10 Additional paternal lineages described in Native Americans have been defined as a result of recent migration as colonial times, such as the European haplogroup R* (M207), among others.10 Additionally, biallelic markers have been employed to detect additional ancestries, such as the insertion/deletion locus (indel) known as YAP, which has been employed to identify the African paternal ancestry in Latin America.11, 12
The study of NRY biallelic variation for defining paternal lineages is particularly limited in Mestizos from Mexico and is even scarcer in Native American groups from this country.13, 14 Although the Y-STRs used in human identification have been employed to estimate paternal admixture in different Mexican–Mestizo populations,15 these markers do not constitute the ideal tool for this purpose.16 Available estimates based on Y-SNPs have been generated from a small number of populations, such as that of Mexico City,17 and those of the states of Guerrero,18 Jalisco13 and Nuevo León.19 Moreover, with the exception of the latter report on Nuevo León, the remaining studies were restricted to two or three Y-linked biallelic markers; thus, the wide diversity of paternal lineages evaluated by Y-SNPs is relatively unknown throughout the Mexican territory. In this work, we examined 18 Y-SNPs in order to establish a confident landscape of the paternal ancestry in Mestizo populations from different regions of the country. Our aims were the following: (1) to estimate the frequency of Y-linked haplogroups to define the main paternal ancestries of Mexican–Mestizo populations (that is, European, Native American and African) and (2) to ascertain their genetic relationships and population structure. Results revealed the prevalence of European and Native American ancestries, with regional differences supporting significant paternal population structure.
Materials and methods
Population sampling
A total of 659 unrelated males from 10 Mestizo populations located in the following four different regions of Mexico were studied: (1) north: Chihuahua (80), Sinaloa (80) and Durango (20); (2) west: Aguascalientes (81) and Jalisco (80); (3) central–south: Veracruz (55), Guerrero (71), and Oaxaca (30) and (4) southeast: Chiapas (81) and Yucatán (81). The geographic location of these population samples is depicted in Figure 1. Because of the small population sample size of Durango and Oaxaca (n⩽30), they were grouped with Sinaloa and Guerrero to estimate haplogroup frequencies, respectively. The state of Chiapas, located between the south and southeast of Mexico, was classified into the southeast due to the shared Mayan ethnological and linguistic affiliations between ethnic groups from the Yucatán Peninsula and Chiapas.20 All individuals were self-classified as Mestizos because they speak Spanish, have Mexican parents and do not belong to any specific ethnic group. Before inclusion in our study, all volunteers signed an informed consent form, according to the ethical guidelines of the Helsinki Declaration and approval by the Ethical Research Committee at the CUCiénega (UdeG). DNA was extracted from liquid blood samples by the conventional salting-out method.21
PCR multiplex amplification
We amplified 17 Y-SNPs in two reactions based on the report of Brion et al.22 but with primer concentration modifications specified in Figure 2. Primers M3 and M242 were designed for coupling into the PCR multiplex 1 (Table 1). Additionally, in samples with an undefined haplogroup, YAP marker was analyzed using primers and conditions suggested by Hammer and Horai.23 PCR multiplexes were performed in a final volume of 10 μl with PCR master mix of QIAGEN Kit Multiplex PCR (QIAGEN, Hilden, Germany), primers and 10 ng of genomic DNA. PCR products (2.5 and 0.625 μl for PCR multiplex 1 and 2, respectively) were purified with ExoSAP-IT (Amersham Biosciences) to remove primers and unincorporated dNTPs.
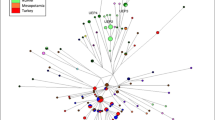
The Y-chromosome phylogenetic tree constructed from the Y-SNPs analyzed in this study based on the nomenclature reported by Karafet et al.30 The color of Y-SNPs indicates PCR multiplex 1 and 2. The (insertion/deletion locus) indel marker YAP was amplified separately. The primer concentration (μM) standardized for PCR and SBE reactions is described. A full color version of this figure is available at the Journal of Human Genetics journal online.
Single-base extension (SBE) reaction
SBE reactions were performed in a total volume of 5 μl with 2.5 μl SNaPshot reaction mix (Applied Biosystems, Foster City, CA, USA), 1 μl SBE primer mix (Figure 2), 0.3 and 1.5 μl of the purified PCR product for Multiplex 1 and 2, respectively. We followed the SBE cycling conditions specified by Brion et al.22 Later, SBE products were purified with 1 μl of SAP (Amersham Biosciences). For capillary electrophoresis, 1 μl of SBE product was mixed with 9 μl of HiDi formamide and 0.3 μl of GeneScan-120 LIZ size standard (Applied Biosystems). Afterward, the products were injected electrokinetically for 16 s at 1.2 kV and 5 μA at 60 °C in a 36-cm-length capillary filled with POP-7 on an ABI Prism 3130® Genetic Analyzer. Finally, electropherograms were analyzed with GeneMapper® v3.2 software (Applied Biosystems).
Data analyses
Frequencies of NRY haplogroups and haplotype diversity (D) were estimated individually and in the total population. Exact tests were carried out in order to compare paternal admixture estimates (Y-SNPs) with those obtained from Y-STRs.15 Exact tests were performed to check homogeneity of ancestral components among Mexican-Mestizos using RxC software.24 Population structure was evaluated by analysis of molecular variance,25 in addition to spatial analysis of molecular variance, which allows establishing population clusters incorporating genetic and geographical criteria.26 Pairwise comparisons (FST p-values) of haplogroup distribution were computed with Arlequin 3.11 software (Berne, Switzerland).25 Nei’s genetic distances27 were calculated with the program Genetic Data Analysis (GDA) version 1.1.28 Genetic distances were displayed on a multidimensional scaling (MDS) plot and on a neighbor-joining tree (NJ). The SPSS 10.0 and Tree View29 software were employed for these purposes, respectively.
Results
Haplogroup distribution and paternal ancestry
Among 19 potential haplogroups to be identified with the Y-SNPs analyzed, we found 16 in the total Mexican-Mestizo population sample. Haplogroups O*, C* and C3* were not detected (Table 2). Total paternal ancestry was predominantly European (49.9%), including haplogroups R*, R1a1*, R1b1*, I* and E1b1b1*, whose frequency increased toward north and west (range, 43.2–65%). Among these, R1b1* was the most frequent haplogroup, representing 79.3% of these European lineages, whereas R*, R1a1*, I* and E1b1b1* showed maximum frequency of 10%. Eurasian haplogroups FH*, G*, J*, K* and P* were found in 14.7% of the total sample; outstanding among these, haplogroup J*, which represented nearly one-half of this ancestry (7.6%). European and Eurasian lineages were merged to estimate the paternal European ancestry in each Mexican–Mestizo population (see Discussion), reaching a total frequency of 64.6% (Figure 3).
Paternal Native American ancestry was represented by haplogroups Q1a3a* and Q*, with a total frequency of 30.8% in Mexican-Mestizos and increasing toward the Center and southeast (37–50%) (Table 2) (Figure 3). Among Native American lineages, Q1a3a* was the principal one (81.3%), whereas the proportion of Q* was much lower (18.7%). In fact, Q* lineage was not observed in Durango and Oaxaca (data not shown), probably owing to the limited sample size of these states. African ancestry was represented by haplogroups AB* and E* with a frequency of 4.2% in Mexican-Mestizos (Figure 3), with predominance of lineage E* (4.1%) (Table 2). Although African ancestry was distributed homogeneously throughout the country (P=0.5876), this was higher in Jalisco (8.75%). In fact, in Jalisco, we found a single male AB* (xCT) (0.15%); these two haplogroups are the most ancestral African lineages restricted to sub-Saharan Africa and are characterized by the ancestral state for M168.30 On the other hand, Asian haplogroups D* and DE* were observed in a single male from Chiapas and Aguascalientes, respectively (0.3%). Finally, haplotype diversity (D) did not exhibit a clear geographical pattern (Table 2).
Comparison between paternal admixture estimates (Y-SNPs vs Y-STRs)
We compared our paternal admixture estimates with those previously estimated by means of 12 Y-STRs (included in Y-filer (Foster City, CA, USA) and Powerplex-Y (Madison, WI, USA) human identification kits) in the following five Mexican–Mestizo populations: Chihuahua, Aguascalientes, Jalisco, Chiapas and Yucatán.15 Interestingly, both studies based on Y-chromosome markers displayed similar increasing gradients of European and Native American ancestries toward the north, west, center–south and southeast, respectively. However, in the majority of the aforementioned states (3/5), we found differences in paternal ancestry proportions between Y-STR and Y-SNP admixture estimates (p ⩽0.010480). This result corroborates that slowly evolving Y-SNPs are suitable tools to define the paternal geographical origin with respect to Y-STRs, which present higher mutation rates and render difficult establishing ancestral states.16 In brief, although Y-STRs provide a paternal admixture landscape of human populations, caution must be taken regarding these estimates that should preferably be obtained with Y-SNPs.
Population structure and genetic relationships
The analysis of molecular variance test showed a low but significant differentiation among Mexican-Mestizos (FST=4.68%; P<0.00005), demonstrating population structure of paternal lineages. The spatial analysis of molecular variance test allowed defining the best population clustering with three levels (K=3; FCT=5.75%, P<0.00005; FSC=0.43%, P=0.0635), which was in close agreement with the previously established geographic classification: (1) north and west (now clustered): Chihuahua, Sinaloa-Durango, Aguascalientes and Jalisco; (2) center–south: Veracruz, Guerrero-Oaxaca and (3) southeast: Chiapas and Yucatán. Pairwise comparisons and genetic distances (data not shown) showed a lesser differentiation between populations from the same cluster, in agreement with spatial analysis of molecular variance results, as represented on a MDS plot (Figure 4a) and an NJ tree (Figure 4b). The principal exception was Jalisco, which remained separated in an MDS quadrant (Figure 4a), and is probably consequence of the highest frequency of African haplogroups in this western state (8.7%).
Genetic distances based on the non-recombining region of the human Y-chromosome (NRY) haplogroup distribution among 10 Mexican–Mestizo populations represented on the multidimensional scaling (MDS) plot (a) and the neighbor-joining (NJ) tree (b). Population clustering depicted in the MDS plot is in agreement with spatial analysis of molecular variance (SAMOVA) results. For abbreviations, please see information below Table 2. A full color version of this figure is available at the Journal of Human Genetics journal online.
Discussion
To our knowledge, this study represents the first and greatest effort to define the main paternal lineages and population structure of Mexican-Mestizos throughout the country. In fact, the paternal ancestry estimates obtained herein are more reliable and include a large number of populations compared to the previous Y-STR report.15 This information is important for association studies concerning Y-linked diseases, such as infertility, some cancers, alcohol dependence, among others.31, 32 For human identification purposes, our results are relevant for selecting the correct population database for interpretation of Y-linked DNA profiles that generally are obtained with Y-STRs. The scarce availability of these databases from Mexican–Mestizo populations compel the use of as much information as possible, such as the population clusters inferred herein, which constitute a guide for forensic genetics casework. Finally, in anthropology, these results contribute to historical research, providing an overview of male migration effects after European contact in Mexican territory.33
European and Native American paternal ancestry
The frequency of European haplogroups R1b1*, R*, R1a*, I* and E1b1b1* (49.9%) plus Eurasian haplogroups G*, J* and K* (14.7%) defined the total European ancestry of Mexican-Mestizos in 64.6% Eurasian lineages; that is, those found both in Asia and in Europe, were considered European rather than Asian based on the following: (1) historical records on Mexican population that largely involve Spanish conquerors;3, 34 (2) the elevated frequency of the cited Eurasian lineages in Europe, particularly in the Iberian Peninsula, where Spain is located,22, 35, 36, 37, 38 and (3) the scarce frequency of Asian ancestry (range, 1–1.4%) detected in Mexican-Mestizos according to genome-wide SNP results.5
A higher frequency of European ancestry was observed in the north and west and vice versa; Native American ancestry increased toward Center–South and southeast (Table 2) (Figure 3). This pattern is in close agreement with the conclusions based on 13 autosomal STRs of the Combined DNA Index System (CODIS) widely used for human identification purposes,39 and genome-wide SNPs in Mexican–Mestizo populations.5 This landscape is explained by three main historical facts: (1) lower pre-Hispanic population density in the north of Mexico (Aridoamerica), contrasting with the highest population density of the Center–South and southeast (Mesoamerica) (Figure 1), as previously described.39 The adequate climatic conditions of Mesoamerica explain its preferential peopling, because this environment allowed humans to produce different crops (for example, maize, beans, squash and cocoa), contrary to Aridoamerica that displayed limited vegetation.40 This fact explains the current high concentration of ethnic groups in Mesoamerica, specifically in the states of Chiapas, Oaxaca, Guerrero, Hidalgo, Yucatán, Campeche and Veracruz;41 (2) the demographic decrease of Native American population due to overwork, warfare (for example, ‘‘Chichimeca warfare’’), and the presence of 14 epidemic diseases in Mexico between 1520 and 160042 and (3) the economic development of north and west regions, for instance the silver mines in Zacatecas that since 1548 led to the generation of new mining cities, such as Durango City and Chihuahua City, which attracted large numbers of Spanish males.2, 3 We consider these phenomena to be major events that contributed to diminish the frequency of Native American lineages in some regions and to disperse differentially European lineages throughout the country.
African paternal ancestry
The observed homogeneous distribution of African ancestry throughout the Mexican territory (range, 0–8.8%) contrasts with previous genetic studies and admixture models suggesting an appreciable African ancestry in coastal regions.34, 43, 44 For example, African haplotypes of the β-globin gene have been detected at high frequencies in the Costa-Chica regions of Guerrero and Oaxaca.45 Although the population samples analyzed in our study did not derive from these coastal regions, our results suggest a limited African gene flow of males to the surrounding cities. This conclusion is in agreement with previous results suggesting that the increasing African ancestry is limited to particular coastal regions, but not to wide coastal areas of Mexico.13 Similarly, states with presumable higher African ancestry (that is, Veracruz and Guerrero) have not shown a significant increase; they have only presented the highest interpopulation genetic variance of this ancestry.5 This also could be described as a ‘‘diluting effect’’, in which the settlement of African slaves has not significantly affected the African-ancestry average of nearby cities, but it is indeed possible to find a few individuals with a higher African ancestry. Regarding the relatively high African ancestry in Jalisco (8.8%), this result is attributable to its economic influence during the colonial period when African slaves were brought to work in mines and on sugar plantations, after the indigenous population diminished owing to different causes, such as diseases and Mixtón warfare in Jalisco.2, 46 In addition, African ancestry may have come to Mexico via the Spaniards, who also received lineages from Middle East and North Africa,34 as described in different parts of Europe (that is, the Balkans, Italy, Portugal and the south of Spain).47
Asymmetrical admixture or differential gene flow
As mentioned previously, European paternal lineages are the most frequent haplogroups in the majority of Mexican–Mestizo populations. Conversely, the maternal lineages (mtDNA) found in Mestizos from different regions including north,48 center and south17, 18, 49 are mainly of Native American ancestry (⩾90%), displaying a strong gender-biased admixture pattern. This phenomenon has been widely described as asymmetrical admixture or differential gene flow.4, 11, 50, 51, 52 Although our admixture estimates in Mexican-Mestizos are similar to Latin American populations regarding the predominance of paternal European ancestry, a number of Latin American populations display low levels of Native American ancestry (0–11.9%), with variable amounts of African ancestry (range, 0.9–30%).12, 51, 52, 53, 54, 55, 56 Conversely, maternal ancestry in Latin America is mainly of Native American origin, except in specific locations such as Cuba and Melo City (Uruguay), where the African ancestry prevails with 45.3% and 52%, respectively.53, 54
Genetic relationships and population structure
In general, the significant paternal population structure inferred here was similar to that observed in a previous study based on Y-STRs in Mexican-Mestizos,15 in which three main regions are distinguished, including the following: (1) north–west, (2) center–south and (3) southeast. Coincidentally, analysis of autosomal STRs has shown a similar structural pattern,39 and for practical purposes this could be relevant for epidemiological or human identification purposes. As could be expected, genetic relationships among Mexican-Mestizos were consistent with the inferred population structure and in agreement with higher European ancestry in the north and west than in the rest of the country (Figure 3).
In conclusion, the paternal ancestry of Mexican–Mestizo males was mainly European, Native American and African, respectively. However, a contrasting pattern of genetic variation based on European and Amerindian ancestry throughout the country was observed, resulting in population structure. A gender-biased admixture pattern was demonstrated in Mexican-Mestizos, which was characterized by a greater proportion of European men and Native American women.
References
INEGI. Conteo de población y vivienda, Tabulados Basicos, Tomo I. 2006: México, (2005).
García-Martinez, B. in Nueva Historia Mínima de México (eds Escalante-Gonzalbo P., García-Martínez B., Jáuregui L., Zoraida-Vázquez J., Speckman-Guerra E., Garciadiego J. 58–112 Colegio de México: México, (2009).
Moreno-Toscano, A. in Historia mínima de México (ed. Cosio.Villegas D. 53–86 Colegio de México: México, (2003).
Wang, S., Ray, N., Rojas, W., Parra, M. V., Bedoya, G., Gallo, C. et al. Geographic patterns of genome admixture in Latin American Mestizos. PLoS Genet. 4, e1000037 (2008).
Silva-Zolezzi, I., Hidalgo-Miranda, A., Estrada-Gil, J., Fernandez-Lopez, J. C., Uribe-Figueroa, L., Contreras, A. et al. Analysis of genomic diversity in Mexican Mestizo populations to develop genomic medicine in Mexico. Proc. Natl Acad. Sci. USA 106, 8611–8616 (2009).
Bryc, K., Velez, C., Karafet, T., Moreno-Estrada, A., Reynolds, A., Auton, A. et al. Colloquium paper: genome-wide patterns of population structure and admixture among Hispanic/Latino populations. Proc. Natl Acad. Sci. USA 107, 8954–8961 (2010).
Underhill, P. A., Passarino, G., Lin, A. A., Shen, P., Mirazón Lahr, M., Foley, R. A. et al. The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations. Ann. Hum. Genet. 65, 43–62 (2001).
Rangel-Villalobos, H in FÓSILES Y MOLÉCULAS: Aproximaciones a La Historia Evolutiva del Homo sapiens (ed. González-Martín A. 120–171 Memorias Real Sociedad Española de Historia Natural, Madrid, España, (2010).
Shriver, M. D., Kittles, R. A. Genetic ancestry and the search for personalized genetic histories. Nat. Rev. Genet. 5, 611–618 (2004).
Zegura, S. L., Karafet, T. M., Zhivotovsky, L. A., Hammer, M. F. High-resolution SNPs and microsatellite haplotypes point to a single, recent entry of Native American Y chromosomes into the Americas. Mol. Biol. Evol. 21, 164–175 (2004).
Carvajal-Carmona, L. G., Soto, I. D., Pineda, N., Ortíz-Barrientos, D., Duque, C., Ospina-Duque, J. et al. Strong Amerind/white sex bias and a possible Sephardic contribution among the founders of a population in northwest Colombia. Am. J. Hum. Genet. 67, 1287–1295 (2000).
Carvalho-Silva, D. R., Santos, F. R., Rocha, J., Pena, S. D. The phylogeography of Brazilian Y-chromosome lineages. Am. J. Hum. Genet. 68, 281–286 (2001).
Rangel-Villalobos, H., Muñoz-Valle, J. F., González-Martín, A., Gorostiza, A., Magaña, M. T., Páez-Riberos, L. A. Genetic admixture, relatedness and structure patterns among mexican populations revealed by the Y-chromosome. Am. J. Phys. Anthrop. 135, 448–461 (2008).
Kemp, B. M., González-Oliver, A., Malhi, R. S., Monroe, C., Schroeder, K. B., McDonough, J. et al. Evaluating the Farming/Language Dispersal Hypothesis with genetic variation exhibited by populations in the Southwest and Mesoamerica. Proc. Natl Acad. Sci. USA 107, 6759–6764 (2010).
Salazar-Flores, J., Dondiego-Aldape, R., Rubi-Castellanos, R., Anaya-Palafox, M., Nuño-Arana, I., Canseco-Avila, L. M. et al. Population structure and paternal admixture landscape on present-day Mexican-Mestizos revealed by Y-STR haplotypes. Am. J. Hum. Biol. 22, 401–409 (2010).
de Knijff, P. Messages through bottlenecks: on the combined use of slow and fast evolving polymorphic markers on the human Y chromosome. Am. J. Hum. Genet. 67, 1055–1061 (2000).
Martínez-Marignac, V. L., Valladares, A., Cameron, E., Chan, A., Perera, A., Globus-Goldberg, R. et al. Admixture in Mexico City: implications for admixture mapping of Type 2 diabetes genetic risk factors. Hum. Genet. 120, 807–819 (2007).
Bonilla, C., Gutierrez, G., Parra, E. J., Kline, C., Shriver, M. D. Admixture analysis of a rural population of the state of Guerrero. Mexico. Am. J. Phys. Anthropol. 128, 861–869 (2005).
Martínez-Fierro, M. L., Beuten, J., Leach, R. J., Parra, E. J., Cruz-Lopez, M., Rangel-Villalobos, H. et al. Ancestry informative markers and admixture proportions in northeastern Mexico. J. Hum. Genet. 54, 504–509 (2009).
Duverger, C. El primer mestizaje: La clave para entender el pasado mesoamericano 1 ed Santillana: México, (2007).
Miller, S. A., Dikes, D. D., Polesky, H. P. A simple salting out procedure for extrating DNA from human nucleated cells. Nucleic Acid Res. 16, 1215 (1989).
Brion, M., Sanchez, J. J., Balogh, K., Thacker, C., Blanco-Verea, A., Borating, C. et al. Introduction of an single nucleotide polymorphism-based ‘Major Y-chromosome haplogroup typing kit’ suitable for predicting the geographical origin of male lineages. Electrophoresis 26, 4411–4420 (2005a).
Hammer, M. F., Horai, S. Y chromosomal DNA variation and the peopling of Japan. Am. J. Hum. Genet. 56, 951–962 (1995).
Raymond, M., Rousset, F. An exact test for population differentiation. Evolution 49, 1280–1283 (1995).
Excoffier, L., Laval, G., Schneider, S. Arlequin ver. 3.11: an integrated software package for population genetics data analysis. Evol. Bioinform. Online pp 145 (2007).
Dupanloup, I., Schneider, S., Excoffier, L. A simulated annealing approach to define the genetic structure of populations. Mol. Ecol. 11, 2571–2581 (2002).
Nei, M. Genetic distance between populations. Am. Nat. 106, 238–292 (1972).
Lewis, P. O., Zaykin, D. Genetic Data Analysis: computer progam for the analysis of allelic data. Ver. 1.0 (d16c). Free program distributed by the authors over the internet from GDA (2001).
Page Roderic, D. M. Available at http://taxonomy.zoology.gla.ac.uk/rod/rod.html TreeView (2001).
Karafet, T. M., Mendez, F. L., Meilerman, M. B., Underhill, P. A., Zegura, S. L., Hammer, M. F. New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res. 18, 830–838 (2008).
Jobling, M. A., Tyler-Smith, C. New uses for new haplotypes the human Y chromosome, disease and selection. Trends Genet. 16, 356–362 (2000).
Krausz, C., Quintana-Murci, L., Forti, G. Y chromosome polymorphisms in medicine. Ann. Med. 36, 573–583 (2004).
Aguirre-Beltrán, G. La población negra de México, Fondo de cultura económica: México, (1989).
Gorodezky, C., Alaez, C., Vázquez-García, M. N., De la Rosa, G., Infante, E., Balladares, S. et al. The genetic structure of Mexican Mestizos of different locations: tracking back their origins through MHC genes, blood group systems, and microsatellites. Hum. Immunol. 62, 979–991 (2001).
Brion, M., Sobrino, B., Blanco-Vera, A., Lareu, M. V., Carrecedo, A. Hierarchical analysis of 30 Y-Chromosome SNPs in European Populations. Int. J. Legal Med. 119, 10–15 (2004).
Semino, O., Magri, C., Benuzzi, G., Lin, A. A., Al-Zahery, N., Battaglia, V. et al. Origin, diffusion, and differentiation of Y-chromosome haplogroups E and J: inferences on the neolithization of Europe and later migratory events in the Mediterranean area. Am. J. Hum. Genet. 74, 1023–1034 (2004).
Brion, M., Dupuy, B. M., Heinrich, M., Hohoff, C., Hoste, B., Ludes, B. et al. A collaborative study of the EDNAP group regarding Y-chromosome binary polymorphism analysis. Forensic Sci. Int. 153, 103–108 (2005b).
Adams, S.M., Bosch, E, Balaresque, P.L., Ballereau, S.J., Lee, A.C., Arroyo, E. et al. The genetic legacy of religious diversity and intolerance: paternal lineages of Christians, Jews, and Muslims in the Iberian Peninsula. Am. J. Hum. Genet. 83, 725–736 (2008).
Rubi-Castellanos, R., Martínez-Cortés, G., Muñoz-Valle, J. F., González-Martín, A., Cerda-Flores, R. M., Anaya-Palafox, M. et al. Pre-Hispanic Mesoamerican demography approximates the present-day ancestry of Mestizos throughout the territory of Mexico. Am. J. Phys. Anthropol. 139, 284–294 (2009).
Nárez, J . Aridamérica y Oasisamérica, Historia Antigua de México. En Preclásico eds Manzanilla L., López-Luján L. 121–157 INAH, Coordinación de humanidades y Instituto de investigaciones antropológicas: México, (2000).
Cisneros, I. H. In. En Derechos humanos de los pueblos indígenas en México (ed. Cisneros I. H. 85–88 CDHDF: México, (2004).
Serrano-Sánchez, C. 500 años de historia: La conquista y el mestizaje biológico en México. En Conquista, transculturación y mestizaje, raíz y origen de México (ed. Ochoa, L. 37–45 Instituto de Investigaciones antropológicas: México, (1995).
Lisker, R. Estructura Genética de la Población Mexicana: aspectos médicos y antropológicos, Salvat: México, (1985).
Gorostiza, A., Acunha, V., Gené, M., Calderón, R., González-Martín, A. in Diversidad Biológica y Salud Humana (ed. Martínez-Almagro A. A. 157–163 Universidad Católica: San Antonio, (2006).
Magaña, M. T., Ongay, Z., Tagle, J., Bentura, G., Cobian, J. G., Perea, F. J. et al. Analysis of betaS and betaA genes in a Mexican population with African roots. Blood Cells Mol. Dis. 28, 12–16 (2002).
Muriá, J. M. Historia de Jalisco, UNED: Guadalajara, México, (1980).
Cruciani, F., La Fratta, R., Trombetta, B., Santolamazza, P., Sellitto, D., Colomb, E. B. et al. Tracing past human male movements in northern/eastern Africa and western Eurasia: new clues from Y-chromosomal haplogroups E-M78 and J-M12. Mol. Biol. Evol. 24, 1300–1311 (2007).
Green, L. D., Derr, J. N., Knight, A. mtDNA affinities of the peoples of North-Central Mexico. Am. J. Hum. Genet. 66, 989–998 (2000).
Guardado-Estrada, M., Juarez-Torres, E., Medina-Martinez, I., Wegier, A., Macías, A., Gomez, G. et al. A great diversity of Amerindian mitochondrial DNA ancestry is present in the Mexican mestizo population. J. Hum. Genet. 54, 695–705 (2009).
Torroni, A., Chen, Y. S., Semino, O., Santachiara-Beneceretti, A.S., Scott, C. R., Lott, M. T. et al. mtDNA and Y-chromosome polymorphisms in four Native American populations from Southern Mexico. Am. J. Hum. Genet. 54, 303–318 (1994).
Carvajal-Carmona, L. G., Ophoff, R., Service, S., Hartiala, J., Molina, J., Leon, P. et al. Genetic demography of Antioquia (Colombia) and the Central Valley of Costa Rica. Hum. Genet. 112, 534–541 (2003).
Rojas, W., Parra, M. V., Campo, O., Caro, M. A., Lopera, J. G., Arias, W. et al. Genetic make up and structure of Colombian populations by means of uniparental and biparental DNA markers. Am. J. Phys. Anthropol. 143, 13–20 (2010).
Sans, M., Weimer, T. A., Franco, M. H., Salzano, F. M., Bentancor, N., Alvarez, I. et al. Unequal contributions of male and female gene pools from parental populations in the African descendants of the city of Melo, Uruguay. Am. J. Phys. Anthropol. 118, 33–44 (2002).
Mendizabal, I., Sandoval, K., Berniell-Lee, G., Calafell, F., Salas, A., Martínez-Fuentes, A. et al. Genetic origin, admixture, and asymmetry in maternal and paternal human lineages in Cuba. BMC Evol. Biol. 8, 213 (2008).
Salas, A., Jaime, J. C., Alvarez-Iglesias, V., Carracedo, A. Gender bias in the multiethnic genetic composition of central Argentina. J. Hum. Genet. 53, 662–674 (2008).
Corach, D., Lao, O., Bobillo, C., van Der Gaag, K., Zuniga, S., Vermeulen, M. et al. Inferring continental ancestry of argentineans from Autosomal, Y-chromosomal and mitochondrial DNA. Ann. Hum. Genet. 74, 65–76 (2010).
Acknowledgements
We thank the Consejo Nacional de Ciencia y Tecnología-México (CONACyT-México; grant 129693) and the Consejo Estatal de Ciencia y Tecnología de Jalisco (COECyT-JAL) for grant 25-2008-32) to HR-V. We also are grateful for the Doctoral fellowship from CONACyT-México awarded to GM-C. Authors state that there is not conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Martínez-Cortés, G., Salazar-Flores, J., Gabriela Fernández-Rodríguez, L. et al. Admixture and population structure in Mexican-Mestizos based on paternal lineages. J Hum Genet 57, 568–574 (2012). https://doi.org/10.1038/jhg.2012.67
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.67
Keywords
This article is cited by
-
Genome wide association study on development and evolution of glutinous rice
BMC Genomic Data (2022)
-
COVID-19 and gastrointestinal symptoms in Mexico, a systematic review: does location matter?
BMC Infectious Diseases (2021)
-
“Distribution of paternal lineages in Mestizo populations throughout Mexico: an in silico study based on Y-STR haplotypes”
Journal of Human Genetics (2021)
-
Paternal lineages and forensic parameters based on 23 Y-STRs (Powerplex® Y23) in Mestizo males from Mexico City
International Journal of Legal Medicine (2020)
-
BLK and BANK1 polymorphisms and interactions are associated in Mexican patients with systemic lupus erythematosus
Inflammation Research (2019)