Abstract
The Japanese Archipelago stretches over 4000 km from north to south, and is the homeland of the three human populations; the Ainu, the Mainland Japanese and the Ryukyuan. The archeological evidence of human residence on this Archipelago goes back to >30 000 years, and various migration routes and root populations have been proposed. Here, we determined close to one million single-nucleotide polymorphisms (SNPs) for the Ainu and the Ryukyuan, and compared these with existing data sets. This is the first report of these genome-wide SNP data. Major findings are: (1) Recent admixture with the Mainland Japanese was observed for more than one third of the Ainu individuals from principal component analysis and frappe analyses; (2) The Ainu population seems to have experienced admixture with another population, and a combination of two types of admixtures is the unique characteristics of this population; (3) The Ainu and the Ryukyuan are tightly clustered with 100% bootstrap probability followed by the Mainland Japanese in the phylogenetic trees of East Eurasian populations. These results clearly support the dual structure model on the Japanese Archipelago populations, though the origins of the Jomon and the Yayoi people still remain to be solved.
Similar content being viewed by others
Introduction
The origins of the people living in the Japanese Archipelago have been studied for a long time (for review, see Suzuki1 and Saitou2). The standard theory based on craniofacial data is the dual structure model proposed by Hanihara.3 According to this model, the first migrants to the Japanese Archipelago came from somewhere in Southeast Asia in the Upper Paleolithic age, who were ancestors of the Jomon people. The second wave of migration took place later in the Yayoi period, and the people came in this time from Northeast Asia. The indigenous Jomon people and the new migrants in and after the Yayoi period gradually mixed with each other. This model provides a reasonable explanation for the morphological similarity between the Ainu people of Hokkaido, the northernmost main island of the Japanese Archipelago, and the Ryukyuan (or Okinawan) people in the Southwest Archipelago, despite a large geographical distance. The similarity of these peoples was already noticed 101 years ago by von Baelz4 as the Ainu–Ryukyuan common origin theory.
A series of studies on genetic polymorphisms of classic markers, such as blood groups, serum proteins and red cell enzymes, were carried out for human populations in the Japanese Archipelago from the 1960s to the 1970s, and the Ainu and the Ryukyuan populations were also studied (for example, Misawa and Hayashida;5, 6 Omoto and Harada;7 Nakajima et al.;8 Omoto et al.9). Omoto10, 11 computed genetic distances among various populations of the world, and by constructing a phylogenetic tree he concluded that the Ainu population may have originated in East Asia, in spite of their unique morphological characters somewhat resembling West Eurasians. Omoto et al.9 estimated genetic distances among the Ainu, the Ryukyuan, the Mainland Japanese and the Chinese populations. Although they did not show a phylogenetic tree, distance relationships indicated a clustering of the Ainu and the Ryukyuan if we apply the neighbor-joining method.12 Omoto13 constructed a phylogenetic tree for various populations of the Mainland Japan, the Ainu and the Ryukyuan, and showed that the Ainu and the Ryukyuan were clustered together. Seven serum protein polymorphism data were used for this tree construction, and this tree was the first one to suggest the genetic similarity between the Ainu and the Ryukyuan. Nei14 constructed a neighbor-joining12 tree of 15 Eurasian populations based on DA distances15 computed from allele frequency data of 18 polymorphic classic markers. The Ainu and the Ryukyuan clustered with 62% bootstrap probability, followed by the Mainland Japanese population.
Omoto and Saitou16 constructed an unrooted tree of three populations in the Japanese Archipelago (Ainu, Ryukyuan and Mainland Japanese) and Korean from allele frequency data of 25 classic polymorphic marker loci, and showed that Ainu and Ryukyuan clustered with 85% and 74% bootstrap probabilities when DA distances15 and Dst distances17 were used, respectively. These probabilities were considerably higher than the random expectation (33%=1/3), and they considered these results as partially supporting the dual structure hypothesis of Hanihara.3 This is because the Ainu–Ryukyuan clustering was expected from the contrasting genetic backgrounds of Jomon vs Yayoi, irrespective of their origins. Saitou18 later constructed a distance-based phylogenetic network19 of these four populations using DA distance values obtained by Omoto and Saitou,16 and found that the length of split separating Ainu and Ryukyuan populations from Mainland Japanese and Korean was much longer than that separating Mainland Japanese and Ryukyuan from the rest of two populations. It confirmed a tree-like structure of the four populations shown by Omoto and Saitou.16
Mitochondrial DNA and Y chromosomal DNA examinations became popular for human population studies from the 1980s, and a series of papers were published for human populations of the Japanese Archipelago.20, 21, 22, 23, 24, 25, 26, 27, 28 All of these studies showed some genetic similarity between the Ainu and the Ryukyuan populations. However, mitochondrial DNA and Y chromosomes are both non-recombining, and the genetic information that can be extracted is limited. Meanwhile, Tokunaga and his colleagues29, 30, 31, 32 conducted DNA typings of highly polymorphic human leukocyte antigen loci for human populations of the Japanese Archipelago, and found a clustering of the Ainu and the Ryukyuan populations.
The situation surrounding DNA polymorphism study of human population drastically changed with the sequencing of the human genome.33 A genome-wide cataloging of single-nucleotide polymorphisms (SNPs) were carried out for four human populations including the Japanese living in Tokyo.34 Nishida et al.35 also conducted an SNP typing of 400 Mainland Japanese individuals, and Yamaguchi-Kabata et al.36 produced and studied SNP data for 7000 individuals living in the various locations of the Japanese Archipelago. They clearly demonstrated the genetic differences between the Mainland Japanese and the Ryukyuan populations. Worldwide surveys of SNP typing were carried out under the Human Genome Diversity Project (HGDP-CEPH)37 including the Mainland Japanese, and the human genetic diversity was analyzed for many human populations in Asia under the HUGO Pan-Asian SNP Consortium,38 including the Mainland and Ryukyu Japanese. However, the Ainu individuals were not included in any of these studies.
One of us (Keiichi Omoto) carried out a series of studies on the genetic polymorphism of many human populations in Asia, and DNA samples have been preserved under his and Momoki Hirai's leadership at the University of Tokyo Kashiwa Campus. We recently formed ‘Asian Archival DNA Repository Consortium’ to keep and utilize these precious DNA samples. This paper is an outcome of activities of this Consortium. The availability of archival DNA in the Ainu population as well as the advancement of high throughput SNP genotyping for this study allowed us to examine in closer detail the genetic substructure and the evolutionary history of the human populations in the Japanese Archipelago. We conducted both individual-based analyses and population-based analyses, and will suggest that the two major waves of migrations during the paleolithic–Jomon period and the Yayoi and the later period produced the unique features of human populations in the Archipelago.
Materials and methods
Sample data, ethical approval and SNP genotyping
Blood samples of the Ainu people were collected in Biratori Town, Hidaka District of Hokkaido in the early 1980s for analysis of DNA by a group from The University of Tokyo, and have since been archived there. A study to use these samples was approved by the Research Ethics Committee of The University of Tokyo. These Ainu DNA samples were the same used in previous studies on mitochondrial DNAs,20, 21, 23, 24 on Y chromosome,24, 26, 28 and on human leukocyte antigen types.29, 30, 31, 32 Two of us (Naruya Saitou and Timothy Jinam) recently visited Biratori Town, and explained these molecular anthropological studies in the past as well as the current study to the representatives of the Ainu people living in that area. DNA samples of the Ryukyuan from the Okinawa main island were collected from 2004 to 2008 by a group from University of the Ryukyus. A study to use these samples was approved by the Ethics Committee of University of the Ryukyus.
A total of 36 Ainu and 38 Ryukyuan samples were genotyped using the Affymetrix genome-wide SNP 6.0 microarray platform (Affymetrix, Santa Clara, CA, USA). All genotyping experiments and their computational analyses were conducted at the Department of Human Genetics, The University of Tokyo. In addition to the Ainu and the Ryukyuan populations, SNP genotype data, generated using the same method, from 200 Mainland Japanese (first set) mostly from the Kanto area35 were used. These three groups (Ainu, Ryukyuan and Mainland Japanese) form the Japanese Archipelago population data set, which was further augmented with HapMap data34 from four populations, namely Yorubans from Africa, Americans of European origins, Han Chinese from Beijing (CHB) and Japanese from Tokyo (JPT). The list of these seven populations is shown in Table 1.
Data filtering and quality checks
SNPs of the mitochondrial DNA, X chromosome and Y chromosome were excluded from the initial SNPs numbering 906 600. Duplicate SNPs and those without a dbSNP ID were also filtered out, resulting in a total of 868 257 remaining SNPs. Individual samples with poor genotyping performances were further filtered out based on the Affymetrix contrast quality control (cQC) threshold of 0.04, as recommended by the manufacturer. Three Ryukyuan and two Mainland Japanese samples were omitted based on this criterion. However, in the Ainu population, only 13 out of 36 individuals passed the cQC threshold. This was probably due to the degradation of DNA quality of the archival samples. To maximize the number of Ainu individuals to be used for downstream analysis, further SNP filtering was done based on confidence scores for each SNP generated during genotype calling using the Affymetrix Birdseed Ver2 algorithm.
In general, SNPs with a confidence score >0.1 are more likely to have failed genotype calling (that is ‘no calls’). By visually inspecting genotype cluster graphs of random SNPs with confidence scores ranging from 0.1 to 0.004, a more stringent cutoff of 0.008 was used to exclude under-performing SNPs while retaining the maximum number of individuals. Thus, based on this criterion, 212 448 SNPs were omitted from the set of 36 Ainu individuals, resulting in 656 237 remaining SNPs. The SNP data in the Ainu and all other populations that were generated using the Affymetrix Genome-Wide 6.0 Assay were further filtered to remove SNPs with call rate <95% and that deviate from Hardy–Weinberg equilibrium (P<0.001). For example, 449 SNP loci were further excluded from the Ainu data, resulting in 655 788 SNP loci (see Supplementary Table S1). The filtering steps were done on each population separately and the number of SNPs filtered out is shown in Supplementary Table S1. After merging SNP data from all populations, the final number of SNPs in the seven population data sets was 641 314.
Merging with other population data
We also included SNP data from 30 other East Asian populations available from public databases in addition to the Japanese Archipelago and HapMap population data sets. These included the HGDP-CEPH data set, which consists of 650 000 SNPs from 51 worldwide populations37 and the Pan-Asian SNP (PASNP) data set, which consists of 54 794 SNPs from 73 populations in Asia.38 The number of SNPs that overlap between the Japanese Archipelago-HapMap 7-population data sets with the HGDP-CEPH panel was 114 001. After applying filters (excluding SNPs with <95% genotype call rate and minor allele frequency <1%) in the merged data set, there were 101 562 SNPs remaining. For merging the Japanese Archipelago-HapMap population data sets with the PASNP data, 15 526 overlapping SNPs from both data sets were extracted and merged. After applying the same filtering criteria as above, the number of remaining SNPs was 14 997. The combination of the Japanese Archipelago, HapMap, HGDP-CEPH and PASNP data sets yielded only 4237 overlapping SNPs. All filtering and merging steps were carried out using PLINK software.39 Supplementary Table S2 shows the list of 16 and 14 populations used from HGDP-CEPH and PASNP data sets, respectively.
Data analysis
Subsequent analysis was carried out using different combinations of the above data sets. For the merged data from all data sets, only populations in East Asia (Supplementary Table S2) were used for analysis. We first conducted individual-based analyses. Principal component analysis (PCA), using the smartpca program in the EIGENSOFT software package,40 was our main strategy. The program frappe41 was used to represent alternative views of population structure and admixture patterns. A maximum-likelihood method is used in this program, and it is computationally more efficient than STRUCTURE.42 Population-based phylogenetic trees were constructed by using CONTML (a maximum-likelihood method for allele frequency data43 was used) for SNP allele frequency data and NEIGHBOR (the neighbor-joining method12 was used) for Dst distance17 matrices computed from SNP allele frequency data by using GENDIST programs, all from the PHYLIP package44 with 5000 bootstrap replicates. Neighbor-net networks45 were also constructed from Dst distance matrices using the software SplitsTree 4.46
Results
Individual-based analyses based on over 640 000 SNP data
The PCA result for the individual SNP data of the seven populations listed in Table 1 is shown in Supplementary Figure S1. The African, the West Eurasian and the East Eurasian (five East Asian populations) were located at the apexes of the triangle in this figure. Because African populations are known to be more distantly related from the two Eurasian populations,47, 48, 49 the second PCA was conducted after excluding the African population (Supplementary Figure S2). Now the first principal component (PC1) separates the West Eurasian and the five East Asian populations, whereas the PC2 separates the Ainu population and the remaining four East Asian populations. Interestingly, the Ainu individuals are linearly aligned, suggesting varying degrees of recent admixture with the mainland Japanese population. It also appears that the population genetically closest to the Ainu is the Ryukyuan, despite the fact that these two populations are geographically located on the northernmost and southernmost poles of the Japanese archipelago, respectively.
When the West Eurasian individuals were further eliminated in the third PCA (Figure 1a), the unique feature of the Ainu individuals and the relationship of the other three populations became prominent. The PC1 and PC2 of this figure explained 1.8% and 0.6%, respectively, of the total variances among the SNP data of 356 individuals. The Mainland Japanese individuals and the HapMap Japanese (JPT) individuals clustered together, as expected. Many Ainu individuals were located at the left side and the Han Chinese in Beijing (HapMap CHB) were distributed at the rightmost side, whereas the Ryukyuan and the Mainland Japanese were sandwiched between them. The PC1 and PC2 coordinates were well correlated with Ryukyuan, Mainland Japanese and Han Chinese individuals. This pattern suggests the existence of the south (Ryukyuan) to the north (Han Chinese in Beijing) geographical cline. However, Ainu individuals, distributed in the northernmost Japanese Archipelago, were found to be closer to the Ryukyuan individuals than to the Mainland Japanese, consistent with the result shown in Supplementary Figure S2.
This finding clearly supports the genetic similarity between the Ainu and the Ryukyuan, in spite of their large geographical distance with each other within the Japanese Archipelago.
Another interesting pattern was the substantial interindividual variation among the Ainu individuals compared with those of the other three populations. Three Ainu individuals were within the cluster of the Mainland Japanese population, whereas the other five Ainu individuals in a red circle constituted a distinct cluster. To further examine this pattern, PCA was performed only for the 36 Ainu individuals (Figure 1b). Now, the PC1 and PC2 explained 6.1% and 5.3% of the total variances, respectively. Interestingly, the high heterogeneity similar to that of Figure 1a was reproduced. This finding indicates that the wide variation observed for the Ainu individuals in Figure 1a was not because of the coexistence of other three populations, but because of the interindividual relationship inherent to the Ainu population. This result may be caused by recent admixtures involving two different source populations.
To see whether this was the case, we calculated the allele-sharing distances between the Ainu and the Mainland Japanese individuals, and compared these with the PC1 coordinates of the Ainu individuals. There was a clear positive correlation (r2=0.542) between the allele-sharing distances and the PC1 coordinates (Supplementary Figure S3). The three Ainu individuals within the Mainland Japanese cluster had the smallest allele-sharing distances, and conversely, the Ainu individuals located farthest from the Mainland Japanese on the PC1 axis tend to have greater allele-sharing distances with the Mainland Japanese population. These observations suggest that the allele sharing between the Ainu and the Mainland Japanese was the result of relatively recent and continuing episodes of gene flow between the two populations. If this is the case, another high degree of variation within the Ainu individuals regarding PC2 may also be explained by the gene flow from a human population genetically distinct from the Mainland Japanese. In particular, the five Ainu individuals in the red circle having the highest PC2 coordinates may be affected by recent admixture events.
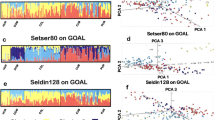
The result of the frappe analysis is shown in Figure 2. When k=2, the two ancestry components corresponds to one (dark blue), which is 100% in 16 Ainu individuals and the other (orange), which is the highest (∼90%) in all the CHB individuals. Interestingly, 20 Ainu individuals showed varied proportions of the orange-colored ancestry component, again suggesting the recent admixture. Most of the Ryukyuan and the Mainland Japanese individuals showed about 30% and 20% blue-colored component, respectively. This difference was consistent with the PCA results in Figure 1a, in that the Ainu was closer to the Ryukyuan than to the Mainland Japanese. The three Ainu individuals who were within the Mainland Japanese cluster in Figure 1a are located at the rightmost columns of the Ainu population, and they also showed ∼20% blue-colored component, similar to the Mainland Japanese.
The result of frappe analysis from k=2 to k=5 for the same individuals of five populations used for the PCA analysis shown in Figure 1a.
As k was increased to 3, the orange-colored component at k=2 divided into two (orange and magenta). Now, all the CHB individuals are almost full of magenta ancestry component, and the Mainland Japanese and JPT individuals also consist of ∼30% magenta-colored ancestry component. There are two Japanese individuals, one in the Mainland Japanese and the other in HapMap JPT, who have >70% magenta component. These two individuals are also outliers in the PCA analysis shown in Figure 1a. If we consider blue- and magenta-colored ancestry components as the Jomon and the Yayoi factors, the intermediate orange-colored component is not easy to comprehend. It is possible that this does not correspond to a real ancestral population, but an artificially inferred component corresponding to the long-term admixture between the Jomon and the Yayoi genetic components. At k=4, the five outlier Ainu individuals observed in the PCA plot (those in red circles in Figures 1a and b) were differentiated from the rest of the Ainu, as indicated in the purple color. The Ryukyuan-specific ancestral component appeared at k=5. This is again not easy to interpret, and could be an artificially inferred component.
Generally speaking, the frappe results appear to be consistent with the PCA analysis in terms of the two patterns: (1) varying amounts of admixture in the Ainu with the Mainland Japanese and (2) the possible presence of another source population, which contributed to the genetic structure of the Ainu. A high correlation between PC1 coordinates of Figure 1a and proportions of the blue-colored ancestry from the frappe analysis (k=2) shown in Supplementary Figure S4 confirms the pattern (1). The pattern (2) can be supported by a high correlation between the purple component frequencies for k=4 and PC2 coordinates in Figure 1a for the Ainu individuals, as shown in Supplementary Figure S5.
Individual-based analyses merged with HGDP-CEPH and PASNP data sets
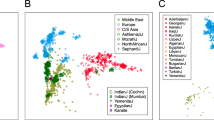
We now move to analyses combined with individuals belonging to the 16 HGDP-CEPH populations. Figure 3a shows the PCA result. The overall distribution of individuals in this figure indicates an L shape. Ainu and Yakut individuals are at the two extremes. The Ryukyuan and the Mainland Japanese populations were located as if they were pulled by the Ainu population, whereas the remaining East Asian populations were closer to northern East Asian populations (Yakut, Mongolian and Oroqen). This clear dichotomy with the L-shaped constellation remains when we further added individuals of the 14-population PASNP data sets (Figure 3b). The Ainu population is located at the one extreme followed by the Ryukyuan, whereas another extreme is now the Uyghur population. The Uyghurs are known to be an admixed population with West Eurasians, so as the Yakuts to a lesser extent.38 This fact indicates that the Ainu, and the two other populations in the Japanese Archipelago (Ryukyuan and Mainland Japanese) to a lesser degree, contain genetic components neither found in the other East Eurasians nor West Eurasians. Another notable addition in Figure 3b is the Korean, who are located between the Mainland Japanese and the Han Chinese in the northern part of China. This result is consistent with that of Tian et al.50
PCA plots of the individuals for the three populations in the Japanese Archipelago (Ainu, Ryukyuans and the Mainland Japanese) and other Asian populations. (a) Result with the HGDP-CEPH data set (see Supplementary Table S2 for the list of populations). (b) Result with the HGDP-CEPH data set and the PASNP data set (see Supplementary Table S2 for the list of populations).
The result of the frappe analysis for populations corresponding to Figure 3a is shown in Supplementary Figure S6. When k=2, the dichotomy pattern is quite similar with that of Figure 2, and additional East Eurasian populations showed similar blue/orange frequencies as in the Han Chinese. As k increased to 3, the light-blue ancestry component, which was almost exclusively found in the Yakut individuals, diverged from the orange component under k=2. The green ancestry component further divided from the orange one at k=4, and this was dominant in the Ryukyuan and the Mainland Japanese. The five ‘outlier’ Ainu individuals in Figure 1 were fully covered by the violet ancestry component at k=5. The fraction of this component was high only in the Ainu. The same situation continues from k=2 to k=6 for the blue ancestry component. This result clearly indicates a unique position of the Ainu population in East Asia. Supplementary Figure S7 shows the result of the frappe analysis for populations corresponding to Figure 3b. The overall pattern is similar to that of Supplementary Figure S6.
Population-based analysis
Individual-based analyses presented in the previous section deciphered the complex structure of each population, especially for the Ainu. Populations are still realistic units of modern human evolution, and we thus constructed phylogenetic trees and networks of populations.
We first used SNP data of the Ainu, the Ryukyuan, the Mainland Japanese and the Han Chinese in Beijing (see Table 1), and constructed an unrooted maximum-likelihood tree (Supplementary Figure S8). The Ainu and the Ryukyuan populations clustered with 100% bootstrap probability, and this is consistent with the PCA result (Figure 1a). The overall pattern is similar to the tree shown by Omoto and Saitou16 based on only 25 polymorphic loci of classic genetic markers, if we equate the Han Chinese in Beijing in the tree of Supplementary Figure S8 with the Korean in their tree; the branch going to the Ainu population is quite long compared with that to the Ryukyuan, whereas the branch going to the Mainland Japanese had zero length.
A neighbor-joining tree was then constructed for the 29 human populations in East Asia, using the merged SNP data with the HGDP-CEPH and PASNP data sets to see the relationships between the three populations in the Japanese Archipelago with other worldwide populations (Figure 4a). Three populations (Japanese, Mainland Japanese and Ryukyu) listed in Supplementary Table S2 were excluded to avoid redundancy of populations. The Korean population is now phylogenetically closest to the human populations in the Japanese Archipelago, though the bootstrap probability to support the clustering of these four populations was 94% (bootstrap value not shown). Four populations (Hezhen, Daur, Oroqen, and Mongolians) that distribute in the northeast Asia as well as the Yakut, the Xibo and the Uyghur formed one cluster in this tree, whereas three populations (Tu, Naxi and Yi) in the southern China formed one cluster, and these two clusters are phylogenetically closer to the Korean and the Japanese Archipelago cluster. In terms of genetic distances, however, some Han Chinese populations (CHB in Beijing and Han-Tw) were closer to populations in the Japanese Archipelago. These smaller genetic distances may be contributed to a smaller random genetic drift caused by large population sizes of these Han populations.
Phylogenetic trees for the three Japanese populations and other Asian populations. (a) A neighbor-joining tree for the three Japanese populations and other Asian populations listed in Supplementary Table S2. (b) A maximum-likelihood tree for the three Japanese populations with Northeast Asians (Hezhen, Daur, Oroqen, Mongolian), Koreans, Han Chinese and populations from central China (Tu, Naxi, Yi). All bootstrap values shown with arrows were obtained from 5000 replications.
We then selected 14 populations in Figure 4a and constructed a neighbor-joining tree (Figure 4b). Because sample sizes of many ethnic minorities in China were small, we merged the data for Hezhen, Daur, Oroqen and Mongolians as the northeast Asian population, and three populations in southern China (Tu, Naxi and Yi) were also merged. The Ainu and the Ryukyuan were clustered with 100% bootstrap probability, followed by the Mainland Japanese. The three populations in the Japanese Archipelago clustered with the Korean with 100% bootstrap probability. The Ainu population has a long branch and is clearly different from the other populations in this figure, confirming the unique phylogenetic position of this population in East Asia. The very short, almost nonexistent branch leading to the Mainland Japanese, as well as its intermediate position between the Ainu-Ryukyu and the cluster for the remaining populations suggest that the Mainland Japanese was formed by the result of admixture between these two ancestral population sources, symbolized as the Jomon and the Yayoi. The northeast Asian population was phylogenetically closest to the Korean–Japanese Archipelago population cluster, followed by the populations in southern China. The three Han Chinese populations in Beijing, Taiwan and Shanghai clustered together with 92% bootstrap probability.
Phylogenetic networks were also constructed for the same genetic distance matrices used for constructing trees in Figures 4a and b, as shown in Supplementary Figures S9 and S10, respectively. The overall pattern of Supplementary Figure S9 was similar to that of Figure 4a, except for the intermediate position of the Yakut, which was not only close to the Uyghur but also close to the four northeast Asian populations (Oroqen, Daur, Hezhen and Mongolian). There are two interesting reticulations in Supplementary Figure S10. Although the southern Chinese populations are close to the three Han Chinese population cluster, they are also close to the northeast Asian population. The Korean is also located in an intermediate position, close to the Japanese Archipelago populations, but also phylogenetically close to the Han Chinese.
Discussion
Genetic heterogeneity of the Ainu population
The s.d. (8.3 × 10−3) of allele-sharing distances for the Ainu population is ∼10 times higher than those (0.75 × 10−3 and 0.85 × 10−3) for the Ryukyuan and the Mainland Japanese, respectively. One possible factor for this variation may be degradation of DNA quality of the Ainu individual samples after preservation of almost 30 years. We thus classified 36 Ainu individuals into two categories (high or low) in terms of Affymetrix Contrast QC (CQC) values. Supplementary Figure S11 shows the distribution of these two categories of individuals on the PCA plot shown in Figure 1a. There is no clear correlation between these two categories and the locations of the Ainu individuals. Furthermore, we performed PCA separately using 13 Ainu individuals who passed the Affymetrix CQC threshold, and the remaining 23 who did not pass the threshold. Supplementary Figure S13 shows that in both cases, the same scattered pattern was observed. We also examined effects of SNP genotyping call heterogeneity among SNP loci. Because the genotyping call rate threshold of 95% was used to produce the PCA plots shown in Figure 1a, we used 90 and 100% call rates and conducted PCA, as shown in Supplementary Figures S12A and S12B. These three PCA results are essentially the same. We further retrieved the top 100 SNPs contributing to PC1 and PC2 in Figure 1a, and a total of 200 SNPs were obtained. After pruning for linkage disequilbrium, 109 SNP loci were left. We used the Digi Tag2 method51 for 96 SNP loci randomly selected from the 109 SNP loci for technical simplicity. The DNA samples used were 36 Ainu, 33 Ryukyuan and 22 Mainland Japanese. SNP call rates for Ainu, Ryukyuan and Mainland Japanese were 0.950, 0.985 and 0.999, respectively. Although the call rate for Ainu was somewhat lower than those for other two populations, the proportions of identical SNP typing with those obtained by using Affymetrix ver. 6.0 were 99.4%, 99.8% and 99.9% for Ainu, Ryukyuan and Mainland Japanese, respectively. We thus conclude that SNP genotypes estimated by using Affymetrix ver. 6.0 were almost identical with those estimated by using the Digi Tag2 method for all the three populations. These results indicate that the DNA degradation was not the reason for the variation among the Ainu individuals.
Interestingly, the average heterozygosity (0.220) of the Ainu population was the least among the seven populations listed in Table 1, whereas those of the other four East Asian populations were more or less the same (∼0.24). This feature is not consistent with a high degree of variation on allele-sharing distances for the Ainu population, if it is a panmictic and isolated population with no gene flow with the surrounding populations. As we showed in this study, however, the Ainu population seems to have experienced gene flow with two different populations, the Mainland Japanese and the yet unknown population in the north. If gene flow happens to occur, the heterogyzosity of admixed individuals should be in the middle of those of the two original populations. To see whether this is the case, the 28 Ainu individuals (three individuals within the Mainland Japanese and five outliers in the red circle in Figure 1a were not included) were divided into two; more admixed and less admixed. When the Ainu individuals were separated according to the normalized PC1 coordinate 0.75 of Supplementary Figure S4, the average heterozygosity of the more admixed 13 Ainu individuals was 0.223, whereas that for the less admixed 15 Ainu individuals was 0.201. When Ainu individuals were separated according to the normalized PC2 coordinate 0.3 of Supplementary Figure S5, average heterozygosities of the less admixed (18), more admixed (10) and outlier (5) Ainu individuals became 0.213, 0.201 and 0.163, respectively. In both cases, the putative admixed Ainu individuals showed the intermediate heterozygosities, and this further confirms that the Ainu population has experienced admixture with the two surrounding populations. In fact, we came to know through discussion with the Ainu people that there were some Mainlaind Japanese individuals who married Ainu people in Biratori Town when blood collection was conducted. These genetically non-Ainu people might have been included in the ‘Ainu’ samples we used. Another information from the Ainu representatives of the Biratori Town was that some Sakhalin Ainu people migrated to that town after the World War II. There is a possibility that the five outliers in the red circle in Figure 1a are Sakhalin Ainu people.
Another possible reason for the low average heterozygosity in the Ainu compared with other populations is ascertainment bias in which SNPs included in the Affymetrix 6.0 genechip were chosen based on their polymorphism in only a few ascertained populations.
Possible mother population for the alternative admixture events with the Ainu population
Unlike admixture with the Mainland Japanese, it is difficult to ascertain the other potential source of admixture in the Ainu without a proper source population. Previous studies did support the idea of contact with Northern populations, which may have contributed to the genetic diversity in the Ainu. Archeological data point to an introduction of a distinct culture, which was quite different from the Satsumon culture, by the Okhotsk people into Hokkaido during the 7th–10th centuries.52 The cultural contact with these northern peoples seemed to continue until recently. Genetic studies using mitochondrial DNA24 and human leukocyte antigen loci32 support this idea by showing close affinities between the Ainu and the Nivkhi who live in the Sakhalin Island and the Amur River region. It would therefore be interesting to collect samples from populations from that area in future studies to have a clearer view of the relationships between the Ainu and Northeast Asian populations.
Perspectives
We still have no clear clues on homelands of the Jomon and the Yayoi people, who constituted the two major genetic components of the modern human populations in the Japanese Archipelago. It should be noted that Omoto53 conducted a pioneering study on the phylogenetic relationship of the Ainu population considering various degrees of admixture. When a 60% admixture with the Mainland Japanese was assumed for the modern Ainu population, the ancestral Ainu population was clustered with Sahulian (Papuan and Australian).53 This sort of simulations based on the real data is needed.
We should also integrate evidences from various research fields such as archeology and morphology. There is a long history of craniofacial studies on Jomon, Yayoi and historical populations in the Japanese Archipelago.1, 3, 54 Although metric characters were mostly used, studies on nonmetric characters are promising,55, 56, 57, 58 for they are expected to be more under genetic controls than metric ones. Hanihara’s analysis58 suggested the existence of gene flows between the Hokkaido Jomon people and the Okhotsk people. An application of the computed tomography images for measuring nonmetric cranial variations in living humans was already started,59 and genome-wide association studies of morphological characters are waiting to be initiated. In fact, the genetic background of shovel-shaped incisors found frequently in East Asian individuals was recently deciphered,60 and the nonsynonymous polymorphism on the EDAR gene showed the greatest difference in SNP genotype frequencies between the Mainland Japanese and Ryukyuan clusters,36 followed by another nonsynonymous polymorphism on the ABCC11 gene that is responsible for the earwax phenotypic differences.61 Similar statistical analyses will be conducted in the near future using the genome-wide SNP data produced by this study. Genomic DNA polymorphisms are not restricted to SNPs, but also include insertion–deletion type polymorphisms such as microsatellite polymorphisms (for example, Li et al.62).
Ancient DNA data are very important for understanding the evolutionary history of the present-day organisms. Because of technical difficulties, most of the ancient DNA data available regarding the origin of peoples in the Japanese Archipelago are mitochondrial DNAs,63, 64, 65, 66, 67, 68, 69, 70, 71 with exceptions of the ABCC11 gene72 and the ABO blood group gene.73 If the genome-wide nuclear DNA polymorphism data can be obtained for ancient DNA samples found from the Japanese Archipelago, we will be able to have a much wider scope on the history of peoples in this Archipelago.
Conclusion
We demonstrated that the Ainu are genetically closer to the Ryukyuan than they are to the Mainland Japanese in this study. The close association between the Ainu and the Ryukyuan, despite their current geographical locations, which is at the two opposing ends of the Japanese Archipelago, may be interpreted as having a shared common ancestry probably dating back to the Jomon period. The population tree in Figure 4b also places the Mainland Japanese in an intermediate position between the Ainu/Ryukyuan and the Continental population clusters. This observation, coupled with the very short external branch in the Mainland Japanese, strongly suggests that they are the result of admixture between the two genetically distinct ancestors, namely the Jomon people and the Yayoi ancestors. Our analysis revealed a great genetic variation within the individuals of the Ainu group, brought about by admixture with the mainland Japanese and possibly another population from Northeast Asia. Figure 5 depicts a plausible time course of the human populations in the three regions of the Japanese Archipelago based on the findings of this study, though many features are still speculative. In conclusion, our results support the more than 100-year-old hypothesis of von Baelz4 that the Ainu and the Ryukyuan have shared genetic ancestry, and the admixture hypothesis (for example, Torii74 and Kanaseki75) that the mainland Japanese are the result of admixture between the ancestral Yayoi people and the indigenous Jomon people.
A scenario of the evolutionary history of the human populations in the three regions of the Japanese Archipelago based on the results of this study and archeological evidences.52 The Northern, the Central and the Southern populations, corresponding to the Ainu, the Mainland Japanese and the Ryukyuan in the present day, were assumed to diverge simultaneously, sometime during the Jomon period, although we do not have a precise time estimate. The admixture of the indigenous Jomon people and Yayoi migrants was assumed to occur sometime after the Yayoi period started 3000 years ago.76 Vertical arrows designate gene flows in historical times, but their timings and frequencies are rather speculative.
The SNP genotype data determined in this study are available upon requests to corresponding authors, under the conditions of collaboration with us and with an appropriate approval of human genomic DNA research ethics committee of institutions to which researchers involved in the data analyses belong.
References
Suzuki, H. The Roots of Japanese Viewed from Bones ( Iwanami Shoten: Tokyo, (1983) {written in Japanese}.
Saitou, N. Japanese Viewed from DNA ( Chikuma Shobo: Tokyo, (2005) {written in Japanese}.
Hanihara, K. Dual structure model for the population history of the Japanese. Jpn Rev. 2, 1–33 (1991).
von Baelz, E. Die Riu-Kiu-Insulaner, die Aino und andere kaukasier-ahnliche Reste in Ostasien. Korres. Blatt. Dtsch. Ges. Anthrop. Ethnol. Urgesch. 42, 187–191 (1911) {written in German}.
Misawa, S. & Hayashida, Y. On the blood groups among the Ainu in Shizunai, Hokkaido. Proc. Jpn. Acad. 44, 83–88 (1968).
Misawa, S. & Hayashida, Y. On the blood groups among the Ainu in Niikappu, Hokkaido. J. Anthrop. Soc. Nippon. 78, 177–186 (1970).
Omoto, K. & Harada, S. The distribution of polymorphic traits in the Hidaka Ainu. II. Red cell enzyme and serum protein groups. J. Fac. Sci. Univ. Tokyo V, IV 2, 171–211 (1972).
Nakajima, H., Ohkura, K., Inafuku, S., Ogura, Y., Koyama, T., Hori, F. et al. The distribution of several serological and biochemical traits in East Asia. II. The distribution of ABO, MNSs, Q, Lewis, Rh, Kell, Duffy and Kidd blood groups in Ryukyu. Jpn. J. Hum. Genet. 12, 29–37 (1967).
Omoto, K., Ishizaki, K., Harada, S., Akaishi, S., Kudo, T. & Takahashi, K. The distribution of serum protein and red cell enzyme types among blood donors of Okinawa Is., the Ryukyus. J. Anthrop. Soc. Nippon. 81, 159–173 (1973).
Omoto, K. Polymorphisms and genetic affinities of the Ainu of Hokkaido. Hum. Biol. Oceania 1, 278–288 (1972).
Omoto, K. The Ainu: a racial isolate? Israel J. Med. Sci. 9, 1195–1215 (1973).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Omoto, K. in The Japanese II (ed. Ikeda J., 217–263 ( Yuzankaku: Tokyo, (1978) {written in Japanese}.
Nei, M. in The Origin and Past of Modern Humans As Viewed from DNA eds Brenner S., Hanihara K., 71–91 World Scientific: Singapore, (1995).
Nei, M., Tajima, F. & Tateno, Y. Accuracy of estimated phylogenetic trees from molecular data. II. Gene frequency data. J. Mol. Evol. 19, 153–170 (1983).
Omoto, K. & Saitou, N. Genetic origins of the Japanese: A partial support for the ‘dual structure hypothesis’. Amer. J. Phys. Anthropol. 102, 437–446 (1997).
Nei, M. Genetic distances between populations. Amer. Nat. 106, 283–292 (1972).
Saitou, N. in Simulations, genetics and human prehistory eds Matsumura S., Forster P., Renfrew C., 89–92 ( McDonald Institute Monographs: Cambridge, (2008).
Bandelt, H. -J. & Dress, A. W. Split decomposition: a new and useful approach to phylogenetic analysis of distance data. Mol. Phyl. Evol. 1, 242–252 (1994).
Harihara, S., Hirai, M. & Omoto, K. Mitochondrial DNA polymorphism in Japanese living in Hokkaido. Jpn. J. Hum. Genet. 31, 73–83 (1986).
Harihara, S., Saitou, N., Hirai, M., Gojobori, T., Park, K. S., Misawa, S. et al. Mitochondrial DNA polymorphism among five Asian populations. Amer. J. Hum. Genet. 43, 134–143 (1988).
Hammer, M. F. & Horai, S. Y chromosomal DNA variation and the peopling of Japan. Amer. J. Hum. Genet. 56, 951–962 (1995).
Horai, S., Murayama, K., Hayasaka, K., Matsubayashi, S., Hattori, Y., Fucharoen, G. et al. MtDNA polymorphism in East Asian Populations, with special reference to the peopling of Japan. Amer. J. Hum. Genet. 59, 579–590 (1996).
Tajima, A., Hayami, M., Tokunaga, K., Juji, T., Matsuo, M., Marzuki, S. et al. Genetic origins of the Ainu inferred from combined DNA analyses of maternal and paternal lineages. J. Hum. Genet. 49, 187–193 (2004).
Tanaka, M., Cabrera, V. M., González, A. M., Larruga, J. M., Takeyasu, T., Fuku, N. et al. Mitochondrial genome variation in eastern Asia and the peopling of Japan. Genome Res. 14, 1832–1850 (2004).
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M. et al. Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes. J. Hum. Genet. 51, 47–58 (2006).
Matsukusa, H., Oota, H., Haneji, K., Toma, T., Kawamura, S. & Ishida, H. A genetic analysis of the Sakishima islanders reveals no relationship with Taiwan aborigines but shared ancestry with Ainu and Main-island Japanese. Amer. J. Phys. Anthropol. 142, 211–223 (2010).
Koganebuchi, K., Katsumura, T., Nakagome, S., Ishida, H., Kawamura, S. & Oota, H., The Asian Archival DNA Repository Consortium Autosomal and Y-chromosomal STR markers reveal a close relationship between Hokkaido Ainu and Ryukyu islanders. Anthropol. Sci. 120 (e-pub ahead of print 2012; doi:10.1537/ase.120322).
Bannai, M., Tokunaga, K., Imanishi, T., Harihara, S., Fujisawa, K., Juji, T. et al. HLA class II alleles in Ainu living in Hidaka District, Hokkaido, northern Japan. Amer. J. Phys. Anthropol. 101, 1–9 (1996).
Hatta, Y., Ohashi, J., Imanishi, T., Kamiyama, H., Iha, M., Simabukuro, T. et al. HLA genes and haplotypes in Ryukyuans suggest recent gene flow to the Okinawa Islands. Hum. Biol. 71, 353–365 (1999).
Bannai, M., Ohashi, J., Harihara, S., Takahashi, Y., Juji, T., Omoto, K. et al. Analysis of HLA genes and haplotypes in Ainu (from Hokkaido, northern Japan) supports the premise that they descent from Upper Paleolithic populations of East Asia. Tissue Antigen 55, 128–139 (2000).
Tokunaga, K., Ohashi, J., Bannai, M. & Juji, T. Genetic link between Asians and native Americans: evidence from HLA genes and haplotypes. Hum. Immunol. 62, 1001–1008 (2001).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Nishida, N., Koike, A., Tajima, A., Ogasawara, Y., Ishibashi, Y., Uehara, Y. et al. Evaluating the performance of Affymetrix SNP Array 6.0 platform with 400 Japanese individuals. BMC Genomics 9, 431–431 (2008).
Yamaguchi-Kabata, Y., Nakazono, K., Takahashi, A., Saito, S., Hosono, N., Kubo, M. et al. Population structure of Japanese based on SNP genotypes from 7,001 individuals in comparison to other ethnic groups: Effects on population-based association studies. Amer. J. Hum. Genet. 83, 445–456 (2008).
Li, J. Z., Absher, D. M., Tang, H., Southwick, A. M., Casto, A. M., Ramachandran, S. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
HUGO Pan-Asian SNP Consortium. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M., Bender, D. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Amer. J. Hum. Genet. 81, 559–575 (2007).
Patterson, N., Price, A. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Tang, H., Peng, J., Wang, P. & Risch, N. Estimation of individual admixture: analytical and study design considerations. Genet. Epidemiol. 28, 289–301 (2005).
Pritchard, J., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Felsenstein, J. Maximum likelihood estimation of evolutionary trees from continuous characters. Amer. J. Hum. Genet. 25, 471–492 (1981).
Felsenstein, J. PHYLIP ( Department of Genome Sciences, University of Washington: Seattle, (2005).
Bryant, D. & Moulton, V. Neighbor-net: an agglomerative method for the construction of phylogenetic networks. Mol. Biol. Evol. 21, 255–265 (2004).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Nei, M. & Roychoudhury, A. Genetic variation within and between the three major races of man, caucasoids, negroids, and mongoloids. Amer. J. Hum. Genet. 26, 421–443 (1974).
Saitou, N. A genetic affinity analysis of human populations. Hum. Evol. 10, 17–33 (1995).
Tishkoff, S., Reed, F. A., Friedlaender, F. R., Ehret, C., Ranciaro, A., Froment, A. et al. The genetic structure and history of Africans and African Americans. Science 324, 1035–1044 (2009).
Tian, C., Kosoy, R., Lee, A., Ransom, M., Belmount, J. W., Gregersen, P. K. et al. Analysis of East Asia genetic structure using genome-wide SNP arrays. PLoS One 3, e3862 (2008).
Nishida, N., Mawatari, Y., Sageshima, M. & Tokunaga, K. Highly parallel and short-acting amplification with locus-specific primers to detect single nucleotide polymorphisms by the DigiTag2 assay. PLoS One 7 e29967 (2012).
Imamura, K. Prehistoric Japan: new perspectives on Insular East Asia ( University of Hawaii Press: Honolulu, (1996).
Omoto, K. in: The Origin and Past of Modern Humans as Viewed from DNA (eds Brenner S., Hanihara K., 92–109 ( World Scientific: Singapore and London, (1995).
Koganei, Y. Beitrage zur physischen Anthropologie der Aino. I. Untersuchungen am Skelett. Mitteil. med. Fac. Kaiser. Univ. Tokyo 2, 1–249 (1893).
Dodo, Y., Ishida, H. & Saitou, N. in The Evolution and Dispersal of Modern Humans in Asia (eds Akazawa T., Aoki K., Kimura T., 479–492 ( Hokusen-sha: Tokyo, (1992).
Fukumine, T., Hanihara, T., Nishima, A. & Ishida, H. Nonmetric cranial variation of early modern human skeletal remains from Kumejima, Okinawa and the peopling of the Ryukyu Islands. Anthropol. Sci. 114, 141–151 (2006).
Komesu, A., Hanihara, T., Amano, T., Ono, H., Yoneda, M., Dodo, Y. et al. Nonmetric cranial variation in human skeletal remains associated with Okhotsk culture. Anthropol. Sci. 116, 33–47 (2008).
Hanihara, T. Nonmetric cranial variation of Jomon Japan: Implications for the evolution of East Asian Diversity. Amer. J. Hum. Biol. 22, 782–790 (2010).
Saitou, N., Kimura, R., Fukase, H., Yogi, A., Murayama, S. & Ishida, H. Advanced CT images reveal nonmetric cranial variations in living humans. Anthropol. Sci. 119, 231–237 (2011).
Kimura, R., Yamaguchi, T., Takeda, M., Kondo, O., Toma, T., Haneji, K. et al. A common variation in EDAR is a genetic determinant of shovel-shaped incisors. Am. J. Hum. Genet. 85, 528–535 (2009).
Yoshiura, K., Kinoshita, A., Ishida, T., Ninokata, A., Ishikawa, T., Kaname, T. et al. A SNP in the ABCC11 gene is the determinant of human earwax type. Nat. Genet. 38, 324–330 (2006).
Li, S. L., Yamamoto, T., Yoshimoto, T., Uchihi, R., Mizutani, M., Kurimoto, Y. et al. Phylogenetic relationship of the populations within and around Japan using 105 short tandem repeat polymorphic loci. Hum. Genet. 118, 695–707 (2006).
Horai, S., Kondo, R., Murayama, K., Hayashi, S., Koike, H. & Nakai, N. Phylogenetic affiliation of ancient and contemporary humans inferred from mitochondrial DNA. Philos. Trans. Royal Soc. London B Biol. Sci. 333, 409–416 (1991).
Oota, H., Saitou, N., Matsushita, T. & Ueda, S. A genetic study of 2,000-year-old human remains from Japan using mitochondrial DNA sequences. Amer. J. Phys. Anthropol. 98, 133–145 (1995).
Oota, H., Saitou, N., Matsushita, T. & Ueda, S. Molecular genetic analysis of remains of a 2,000-year old human population in China-and its relevance for the origin of the modern Japanese population. Amer. J. Hum. Genet. 64, 250–258 (1999).
Shinoda, K. & Kanai, S. Intracemetery genetic analysis at the Nakazuma Jomon Site in Japan by mitochondrial DNA sequencing. Anthropol. Sci. 107, 129–140 (1999).
Wang, L., Oota, H., Saitou, N., Jin, F., Matsushita, T. & Ueda, S. Genetic structure of a 2500-year-old human population in China and its spatiotemporal changes. Mol. Biol. Evol. 17, 1396–1400 (2000).
Sato, T., Amano, T., Ono, H., Ishida, H., Kodera, H., Matsumura, H. et al. Origins and genetic features of the Okhotsk people, revealed by ancient mitochondrial DNA analysis. J. Hum. Genet. 52, 618–627 (2007).
Adachi, N., Shinoda, K., Umetsu, K. & Matsumura, H. Mitochondrial DNA analysis of Jomon skeletons from the Funadomari site, Hokkaido, and its implication for the origins of Native American. Amer. J. Phys. Anthropol. 138, 255–265 (2009).
Adachi, N., Shinoda, K., Umetsu, K., Kitano, T., Matsumura, H., Fujiyama, R. et al. Mitochondrial DNA analysis of Hokkaido Jomon skeletons: remnants of archaic maternal lineages at the southwestern edge of former Beringia. Amer. J. Phys. Anthropol. 146, 346–360 (2011).
Sato, T., Razhev, D., Amano, T. & Masuda, R. Genetic features of ancient West Siberian people of the Middle Ages, revealed by mitochondrial DNA haplogroup analysis. J. Hum. Genet. 56, 602–608 (2011).
Sato, T., Amano, T., Ono, H., Ishida, H., Kodera, H., Matsumura, H. et al. Allele frequencies of the ABCC11 gene for earwax phenotypes among ancient populations of Hokkaido, Japan. J. Hum. Genet. 54, 409–413 (2009).
Sato, T., Kazuta, H., Amano, T., Ono, H., Ishida, H., Kodera, H. et al. Polymorphisms and allele frequencies of the ABO blood group gene among the Jomon, Epi-Jomon and Okhotsk people in Hokkaido, northern Japan, revealed by ancient DNA analysis. J. Hum. Genet. 55, 691–696 (2010).
Torii, R. Japan Before Historic Period ( Isobe: Koyodo, (1918) {written in Japanese}.
Kanaseki, T. The Origin of Japanese Ethnic Group, Hosei University Press: Tokyo, (1976) {written in Japanese}.
Harunari, H. & Imamura, M. The Real Age of the Yayoi Period, Gakuseisha: Tokyo, (2004) {written in Japanese}.
Acknowledgements
We thank all people who donated their DNAs to this study, especially for the Ainu and the Ryukyuan people. We appreciate the effort of Department of Promotion of Measures for Ainu, Biratori Town, for arrangement of the meeting with Ainu representatives. We also thank Ms. Yuko Ogasawara, Natsumi Baba, Rieko Hayashi and Minae Kawashima for technical assistance. We would also like to acknowledge the HUGO Pan-Asian SNP Consortium for providing the SNP genotype data. This work was supported in part by a Strategic Research Project to Naruya Saitou from SOKENDAI, and Grant-in-Aids for Scientific Research (A) (23247040), (A) (22247036) and (B) (21370108) to Hirofumi Matsumura (Katsushi Tokunaga and Naruya Saitou are co-investigators), Shoji Kawamura and Hiroki Oota, respectively, from Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Additional information
Consortium members:
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Japanese Archipelago Human Population Genetics Consortium. The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J Hum Genet 57, 787–795 (2012). https://doi.org/10.1038/jhg.2012.114
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.114
Keywords
This article is cited by
-
An update and frequency distribution of Y chromosome haplogroups in modern Japanese males
Journal of Human Genetics (2024)
-
Prefecture-level population structure of the Japanese based on SNP genotypes of 11,069 individuals
Journal of Human Genetics (2021)
-
Genome-wide SNP data of Izumo and Makurazaki populations support inner-dual structure model for origin of Yamato people
Journal of Human Genetics (2021)
-
Population genomics of East Asian ethnic groups
Hereditas (2020)
-
Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations
Communications Biology (2020)