Abstract
Expression of cyclic adenosine monophosphate-specific phosphodiesterase 7B (PDE7B) mRNA is increased in patients with chronic lymphocytic leukemia (CLL), thus suggesting that variation may occur in the PDE7B gene in CLL. As genetic variation in other PDE family members has been shown to associate with numerous clinical disorders (reviewed in this manuscript), we sought to identify single-nucleotide polymorphisms (SNPs) in the PDE7B gene promoter and coding region of 93 control subjects and 154 CLL patients. We found that the PDE7B gene has a 5′ non-coding region SNP −347C>T that occurs with similar frequency in CLL patients (1.9%) and controls (2.7%). Tested in vitro, −347C>T has less promoter activity than a wild-type construct. The low frequency of this 5′ untranslated region variant indicates that it does not explain the higher PDE7B expression in patients with CLL but it has the potential to influence other settings that involve a role for PDE7B.
Similar content being viewed by others
Introduction
Cyclic nucleotide phosphodiesterases (PDEs), a superfamily of enzymes divided into 11 families (PDE1–11), several with multiple isoforms,1, 2 hydrolyze cyclic adenosine monophosphate (cAMP) and cyclic guanosine monophosphate (cGMP), and thereby have a crucial role in termination of cyclic nucleotide-mediated signaling, in particular signaling by drugs that regulate the generation of cAMP and cGMP.3 In addition, PDEs, including specific PDE isoforms, have been used as drug targets. Several PDEs, including PDE4, PDE7 and PDE8, preferentially hydrolyze cAMP.4 In spite of their importance in the regulation of cyclic nucleotides, the mechanisms that regulate the expression of PDE genes are not well defined. For example, regions in the promoter that regulate activity have only been identified for a limited number of PDE genes5 and details regarding events involved in PDE gene expression are not well understood. Increases in cAMP can regulate the expression of certain PDEs, in particular via action of the cAMP-response element-binding protein or the inducible cAMP early repressor,6, 7, 8, 9 and in addition, glucocorticoids can transcriptionally inhibit PDE expression.10, 11
Our interest in PDE gene expression arose from studies of patients with chronic lymphocytic leukemia (CLL), the most common leukemia in North American and European adults.12 We found that mRNA and protein expression of PDE7B, a cAMP-specific PDE predominantly expressed in brain and lymphocytes,13, 14, 15 are increased >10-fold in peripheral blood mononuclear cells of patients with CLL.16 We thus set out to determine if the increased expression of PDE7B in CLL patients might result from genetic variation in the coding sequence or 5′ untranslated region (5′ UTR) of the PDE7B gene. We began these studies with the knowledge that genetic variation, in particular single-nucleotide polymorphisms (SNPs), had been identified in certain PDE family members, and suggested to contribute to clinical disorders, such as schizophrenia (for example, PDE4B),17, 18, 19 stroke (for example, PDE4D)20 and retinitis pigmentosa (for example, PDE6A and PDE6B),21 but evidence regarding such genetic variation for other PDEs is limited. We thus set out to identify genetic variants in the PDE7B gene and to test the hypothesis that genetic variation might contribute to the increased PDE7B mRNA expression in CLL and to the pathophysiology of CLL with implications for the possible use of inhibitors of PDE7B in the treatment of CLL. By sequencing the 5′ UTR of the PDE7B gene, we identified a variant that decreases promoter activity. In addition, in this article, we place our findings for PDE7B in the context of genetic variation of other human PDEs.
Materials and methods
Patient selection and sample preparation
We evaluated samples from 154 CLL patients followed by the CLL Research Consortium and 93 healthy donors at the University of California, San Diego. Following informed consent, blood was collected from the healthy donors and from patients who satisfied diagnostic and immunophenotypic criteria for CLL. The University of California, San Diego, Institutional review board approved these studies and the CLL Research Consortium approved the procurement of the samples, with all approvals in accordance with the Declaration of Helsinki.
Diagnosis of CLL was based on morphological and immunophenotyping criteria.22 Patients with CLL were also categorized by the expression of 70-kDa zeta-associated protein.23, 24, 25 Mononuclear cells were isolated using a Ficoll gradient and then frozen in fetal calf serum plus 10% dimethyl sulfoxide before storage in liquid N2. The 70-kDa zeta-associated protein expression was determined by flow cytometry of mononuclear cells. PCR was used to assess the immunoglobulin heavy chain variable gene family. Table 1 shows demographic data, heavy chain variable and the 70-kDa zeta-associated protein status.
Genomic DNA and RNA extraction, PCR and sequencing
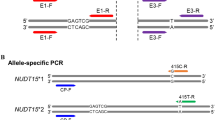
Genomic DNA and RNA were prepared using isolation kits for blood cells (Qiagen, Valencia, CA, USA). The complementary DNA (cDNA) was generated using the Superscript III cDNA synthesis system (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. We sought to identify genetic variants within the putative PDE7B (NC_000006.11) promoter and coding regions as follows: (a) PDE7B gene promoter, the region 700 bp upstream and 300 bp downstream of the transcription start site of PDE7B was amplified; this resulted in a 1-kb PCR product (P1; forward primer: 5′-CACCATAGCCTTGCTTCTTGA-3′; reverse primer, 5′-TGGCAGTTCTGTGACCTTTG-3′). (b) PDE7B coding region (304 to 1656 nt), the full-length cDNA of PDE7B was amplified in two PCR products, P2 from 276 to 1165 bp (forward primer: 5′-CTGGAGAAGTTGCTGGATTCT-3′; reverse primer: 5′-ATTCATTCTGCCTGTTGATGTCT-3′) and P3 from 1074 to 1870 bp (forward primer: 5′-CTCTGGACATCATGCTTGGA-3′; reverse primer: 5′-CTCCCACGTTACTGAATGGAG-3′). PCR amplifications were performed (AccuPower PCR Premix; Bioneer, Alameda, CA, USA) using standard procedures with 50 ng of genomic DNA product (P1) or 200 ng of cDNA (P2–3), in a total volume of 20 μL containing 10 mM Tris-HCl (pH 9.0), 40 mM KCl, 1.5 mM MgCl2, 1 U DNA polymerase, 1.5 mM deoxyribonucleotide triphosphate and 0.5 μM each specific primer. PCR cycling conditions were: initial denaturing at 95 °C for 5 min (P1) or 2 min (P2–3) followed by 35 cycles of denaturing at 94 °C for 40 s; annealing at 59 °C (P1), 55 °C (P2) or 57 °C (P3) for 40 s; extension at 72 °C for 57 s (P1), 59 s (P2) or 58 s (P3); and a final extension step at 72 °C for 5 min. DNA Clean & Concentrator TM-5 kit (D4003 lt 0940 BW) and Centri-sep columns (Zymo Research, Orange, CA, USA) were used to purify PCR products. Sequence was determined on a 3130 × l Genetic Analyzer (Applied Biosytems, Carlsbad, CA, USA), with the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems). Polymorphisms were manually confirmed. The data were validated visually and the files were inspected to identify heterozygotes. Each sequence was imported into the Biology Workbench (http://workbench.sdsc.edu/) for analysis. SNPs were confirmed by re-sequencing from the reverse direction.
Dual luciferase assays
To investigate the functional impact of −347 C>T polymorphism on PDE7B promoter activity, we constructed reporter plasmids with 5′-flanking regions that contained either wild type (−347C) or SNP (−347T). The 1-kb fragment encompassing the PDE7B transcription start site (from −700 to +300) was amplified from patients with the −347 CC or −347 TT genotype, and inserted into the firefly luciferase reporter plasmid pGL4-basic (Promega, Madison, WI, USA). HEK293 cells were transfected with 3 μg of pGL4-Luc containing PDE7B promoter regions of either the wild-type or SNP variants and 0.1 μg of pRL-TK-Luc using the Optifect transfection reagent (Invitrogen, Carlsbad, CA, USA). pRL-TK-Luc encodes the Renilla luciferase under the control of an herpes simplex virus thymidine kinase promoter and serves as an internal control to normalize for transfection efficiency. After 48 h, cells were harvested and assessed using a dual-luciferase reporter assay system (Promega, Madison, WI, USA). Firefly and Renilla luciferase activities were measured using a TD-20/20 Luminometer (Turner Design, Sunnyvale, CA, USA). The results shown were found in three independent experiments.
Statistical analyses
All determinations were performed in duplicate or triplicate. Values are expressed as mean±s.d. Comparison between groups was based on 2 × 2 contingency table and use of the χ2 statistic; P<0.05 was considered to be statistically significant.
Results
CLL study cohort
To determine whether the PDE7B promoter possesses genetic variants, we sequenced samples prepared from 154 CLL patients and 93 control subjects. This cohort represents a typical population of CLL patients (that is, 62% male and 38% female subjects; mean age at diagnosis, 62 years). We also evaluated patient samples for expression of the immunoglobulin heavy chain variable mutation and 70-kDa zeta-associated protein, both of which are prognostic markers in CLL.26, 27 (Table 1).
Identification of a variant in PDE7B and comparison of its frequency in control and CLL subjects
In initial studies designed to identify genetic variants in PDE7B, we sequenced ∼1.3 kb of its coding region in 43 CLL patients and 7 normal subjects and found limited genetic variation in this region: a SNP in the 5′ upstream region, −347C>T, occurred with ∼2% frequency (1.9% in CLL patients 6/308 alleles (all heterozygotes) and 2.7% in 5/186, (3 heterozygotes and 1 homozygote) in controls). Thus, the overall SNP frequency was 2.2% (11/494 alleles). Subgroup analysis of CLL patients classified as having either aggressive or indolent CLL28, 29 revealed a comparable frequency of SNP expression in the two groups of patients (data not shown). Males had a slightly higher, statistically nonsignificant (P>0.1) SNP frequency (2.6%; 9/346) than did females (1.4%; 2/148; Table 2). Expression of the SNP was not associated with other CLL markers (Table 1).
Use of the TFSEARCH program (http://www.cbrc.jp/research/db/TFSEARCH.html) to predict transcription factor binding sites revealed that the PDE7B promoter containing the SNP has putative binding sites for multiple transcription factors, including the sex determination region Y, which encodes a sex-determining transcription factor of the high-mobility group box family and two members of the forkhead box (FOX) family, FOXA2 and FOXD3.30, 31 The TFSEARCH program predicted that the −347C>T SNP abolishes the binding site for sex determination region Y and FOXA2 but not FOXD3. Although binding of FOXD3 was not altered by the SNP, this transcription factor is down-regulated in CLL and in a mouse model of CLL through a nuclear factor-κB p50:histone deacetylase 1 co-repressor complex 3.32
Functional assessment of the PDE7B promoter SNP
On the basis of possible impact of the PDE7B promoter SNP on transcription, we assessed its activity using a dual luciferase assay and found that the 347C>T variant had decreased transcriptional activity compared with the wild-type construct (P<0.001; Figure 1).
Impact of the −347C and −347T genotypes on PDE7B promoter function. pGL4 luciferase reporter constructs containing a 1-kb phosphodiesterase 7B promoter with the −347C (WT) or −347T (SNP) genotype were transfected into HEK293 cells. Vec, the promoter-less pGL4, was used as a control and its luciferase activity was defined as 1. The data represent the mean±s.d. of three independent experiments. **P<0.001 compared with vec alone and *P<0.001 between WT and SNP. SNP, single-nucleotide polymorphism; vec, vector; WT, wild type.
Discussion
Our results indicate that the coding sequence and 5′ upstream region of the PDE7B gene has a SNP (−347C>T) that occurs with low frequency (that is, P<3%), is located in a possible putative binding sites for sex determination region Y, FOXA2 or FOXD3 and decreases transcriptional activity. Thus, although relatively rare, the SNP has the potential to contribute to inter-individual differences in PDE7B expression.
On the basis of information available at the dbSNP (http://www.ncbi.nlm.nih.gov/SNP/), PDE7B is predicted to contain 1616 SNPs within the 345 718-bp contig reported in Build 129. Only two of these are coding SNPs (found in exons 8 and 12), both synonymous, and the rest are found in regulatory regions and introns. There are nine SNPs identified between −473 and −1756 located in the 5′ UTR that are deposited in dbSNP. Thus, the −347C>T SNP discovered in this study is novel, although overall, PDE7B has a relative dearth of potentially functional polymorphisms. Such conservation may imply that the function of PDE7B is important, such that it has undergone minimal genetic change during evolution.
Table 3 summarizes data for genetic variation in PDEs. Genetic variants of PDE6, a retina-specific PDE, occur in certain retinal diseases while other data implicate associations between PDE polymorphisms and certain central nervous system, metabolic, cardiovascular or neoplastic disorders. Inter-subject variability in response to PDE-interacting drugs associates both positively and negatively with genetic variations in PDEs. For example, genetic variations in PDE1A, PDE9A and PDE11A associate with the response to antidepressants but apparently only in certain populations.33, 34, 35 Polymorphisms in PDE5A appear unable to explain differences in cardiovascular effects in subjects who are administered sildenafil or other PDE5 inhibitors, implicating other mechanisms besides such genetic variation as contributing to variable responses.33, 34, 36
Could our results that fail to show an association of genetic variants for PDE7B in CLL be falsely negative? Might copy number variation explain the prominent increase in gene expression of PDE7B in CLL patients?16, 37 Copy number variation (1000 bp segments of genomic DNA with inter-individual variability in copy number38, 39) has been associated with numerous diseases.40, 41, 42 Copy number variations may affect gene expression43 and thus might increase expression of PDE7B transcripts in patients with CLL. An additional mechanism for the increase in PDE7B expression in CLL could be non-coding RNAs, including miRNAs, expression of certain of which is increased in CLL and may contribute to the difference in aggressive and indolent forms of the disease.44 Further studies will be needed to assess gene structure in CLL45 and also whether the SNP alters methylation of the PDE7B promoter, thereby changing chromosomal structure and transcriptional activity.35, 46, 47, 48
References
Beavo, J. A. Cyclic nucleotide phosphodiesterases: functional implications of multiple isoforms. Phys. Rev. 75, 725 (1995).
Conti, M. & Beavo, J. Biochemistry and physiology of cyclic nucleotide phosphodiesterases: essential components in cyclic nucleotide signaling. Annu. Rev. Biochem. 76, 481–511 (2007).
Goodwin, J. S. & Ceuppens, J. Regulation of the immune response by prostaglandins. J. Clin. Immunol. 3, 295–315 (1983).
Moore, A. R. & Willoughby, D. A. The role of cAMP regulation in controlling inflammation. Clin. Exp. Immunol. 101, 387 (1995).
Osawa, H., Niiya, T., Onuma, H., Murakami, A., Ochi, M., Nishimiya, T. et al. Others systematic search for single nucleotide polymorphisms in the 5′ flanking region of the human phosphodiesterase 3B gene: absence of evidence for major effects of identified polymorphisms on susceptibility to Japanese type 2 diabetes. Mol. Genet. Metab. 79, 43–51 (2003).
Ding, B., Abe, J., Wei, H., Xu, H., Che, W., Aizawa, T. et al. A positive feedback loop of phosphodiesterase 3 (PDE3) and inducible cAMP early repressor (ICER) leads to cardiomyocyte apoptosis. Proc. Natl. Acad. Sci. USA 102, 14771 (2005).
Erdogan, S. & Houslay, M. D. Challenge of human Jurkat T-cells with the adenylate cyclase activator forskolin elicits major changes in cAMP phosphodiesterase (PDE) expression by up-regulating PDE3 and inducing PDE4D1 and PDE4D2 splice variants as well as down-regulating a novel PDE4A. Biochem. J. 321, 165 (1997).
McCahill, A., Campbell, L., McSorley, T., Sood, A., Lynch, M. J., Li, X. et al. In cardiac myocytes, cAMP elevation triggers the down-regulation of transcripts and promoter activity for cyclic AMP phosphodiesterase-4A10 (PDE4A10). Cell Signal 20, 2071–2083 (2008).
Verghese, M. W., McConnell, R. T., Lenhard, J. M., Hamacher, L. & Jin, S. L. Regulation of distinct cyclic AMP-specific phosphodiesterase (phosphodiesterase type 4) isozymes in human monocytic cells. Mol. Pharmacol. 47, 1164 (1995).
Ahlström, M., Pekkinen, M., Huttunen, M. & Lamberg-Allardt, C. Dexamethasone down-regulates cAMP-phosphodiesterase in human osteosarcoma cells. Biochem. Pharmacol. 69, 267–275 (2005).
Hermsdorf, T., Richter, W. & Dettmer, D. Effects of dexamethasone and glucagon after long-term exposure on cyclic AMP phosphodiesterase 4 in cultured rat hepatocytes. Cell. Signal. 11, 685–690 (1999).
Goldin, L. R., Pfeiffer, R. M., Li, X. & Hemminki, K. Familial risk of lymphoproliferative tumors in families of patients with chronic lymphocytic leukemia: results from the Swedish family-cancer database. Blood 104, 1850 (2004).
Gardner, C., Robas, N., Cawkill, D. & Fidock, M. Cloning and characterization of the human and mouse PDE7B, a novel cAMP-specific cyclic nucleotide phosphodiesterase. Biochem. Biophys. Res. Commun. 272, 186–192 (2000).
Hetman, J. M., Soderling, S. H., Glavas, N. A. & Beavo, J. A. Cloning and characterization of PDE7B, a cAMP-specific phosphodiesterase. Proc. Natl. Acad. Sci. USA 97, 472 (2000).
Li, L., Yee, C. & Beavo, J. A. CD3- and CD28-dependent induction of PDE7 required for T cell activation. Science 283, 848–851 (1999).
Zhang, L., Murray, F., Zahno, A., Kanter, J. R., Chou, D., Suda, R. et al. Cyclic nucleotide phosphodiesterase profiling reveals increased expression of phosphodiesterase 7B in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 105, 19532–19537 (2008).
Fatemi, S. H., King, D. P., Reutiman, T. J., Folsom, T. D., Laurence, J. A., Lee, S. et al. PDE4B polymorphisms and decreased PDE4B expression are associated with schizophrenia. Schizophr. Res. 101, 36–49 (2008).
Numata, S., Ueno, S. I., Iga, J. I., Song, H., Nakataki, M., Tayoshi, S. et al. Positive association of the PDE4B (phosphodiesterase 4B) gene with schizophrenia in the Japanese population. J. Psychiatr. Res. 43, 7–12 (2008).
Pickard, B. S., Thomson, P. A., Christoforou, A., Evans, K. L., Morris, S. W., Porteous, D. J. et al. The PDE4B gene confers sex-specific protection against schizophrenia. Psychiatr. Genet. 17, 129 (2007).
Nakayama, T., Asai, S., Sato, N. & Soma, M. PDE4D gene in the STRK1 region on 5q12: susceptibility gene for ischemic stroke. Curr. Med. Chem. 14, 3171–3178 (2007).
Hartong, D. T., Berson, E. L. & Dryja, T. P. Retinitis pigmentosa. Lancet 368, 1795–1809 (2006).
Cheson, B. D., Bennett, J. M., Grever, M., Kay, N., Keating, M. J., OBrien, S. et al. National Cancer Institute-sponsored Working Group guidelines for chronic lymphocytic leukemia: revised guidelines for diagnosis and treatment. Blood 87, 4990–4997 (1996).
Kipps, T. J. The B-cell receptor and ZAP-70 in chronic lymphocytic leukemia. Best Pract. Res. Clin. Haematol. 20, 415–424 (2007).
Rassenti, L. Z., Huynh, L., Toy, T. L., Chen, L., Keating, M. J., Gribben, J. G. et al. ZAP-70 compared with immunoglobulin heavy-chain gene mutation status as a predictor of disease progression in chronic lymphocytic leukemia. N. Engl. J. Med. 351, 893–901 (2004).
Wiestner, A., Rosenwald, A., Barry, T. S., Wright, G., Davis, R. E., Henrickson, S. E. et al. ZAP-70 expression identifies a chronic lymphocytic leukemia subtype with unmutated immunoglobulin genes, inferior clinical outcome, and distinct gene expression profile. Blood 101, 4944 (2003).
Cruse, J. M., Lewis, R. E., Webb, R. N., Sanders, C. M. & Suggs, J. L. Zap-70 and CD38 as predictors of IgVH mutation in CLL. Exp. Mol. Pathol. 83, 459–461 (2007).
Moreno, C. & Montserrat, E. New prognostic markers in chronic lymphocytic leukemia. Blood Rev. 22, 211–219 (2008).
Abbott, B. L. Chronic lymphocytic leukemia: recent advances in diagnosis and treatment. Oncologist 11, 21 (2006).
Ghia, P., Ferreri, A. J. M. & Caligaris-Cappio, F Chronic lymphocytic leukemia. Crit. Rev. Oncol. Hematol. 64, 234–246 (2007).
Abel, E. V. & Aplin, A. E. FOXD3 is a mutant B-RAF-regulated inhibitor of G1-S progression in melanoma cells. Cancer Res. 70, 2891 (2010).
Sutton, J., Costa, R., Klug, M., Field, L., Xu, D., Largaespada, D. A. et al. Genesis, a winged helix transcriptional repressor with expression restricted to embryonic stem cells. J. Biol. Chem. 271, 23126 (1996).
Chen, S. S., Raval, A., Johnson, A. J., Hertlein, E., Liu, T. H., Jin, V. X. et al. Epigenetic changes during disease progression in a murine model of human chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 106, 13433 (2009).
Cabanero, M., Laje, G., Detera-Wadleigh, S. & McMahon, F. J. Association Study of Phosphodiesterase Genes in the Sequenced Alternatives to Relieve Depression (STAR* D) sample. Pharmacogenet. Genomics 19, 235 (2009).
Wong, M. L., Whelan, F., Deloukas, P., Whittaker, P., Delgado, M., Cantor, R. M. et al. Phosphodiesterase genes are associated with susceptibility to major depression and antidepressant treatment response. Proc. Natl. Acad. Sci. USA 103, 15124 (2006).
Teranishi, K. S., Slager, S. L., Garriock, H., Kraft, J. B., Peters, E. J., Reinalda, M. S. et al. Variants in PDE11A and PDE1A are not associated with citalopram response. Mol. Psychiatry 12, 1061–1063 (2007).
Salvi, F., Sarzani, R., Giorgi, R., Donatelli, G., Pietrucci, F., Micheli, A. et al. Cardiovascular effects of sildenafil in hypertensive men with erectile dysfunction and different alleles of the type 5 cGMP-specific phosphodiesterase (PDE5). Int. J. Impot. Res. 16, 412–417 (2004).
Zhang, L., Murray, F., Rassenti, L. Z., Pu, M., Kelly, C., Kanter, J. R. et al. Cyclic nucleotide phosphodiesterase 7B mRNA: an unfavorable characteristic in chronic lymphocytic leukemia. Int. J. Cancer 129, 1162–1169 (2011).
Jakobsson, M., Scholz, S. W., Scheet, P., Gibbs, J. R., VanLiere, J. M., Fung, H. C. et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451, 998–1003 (2008).
Pinto, D., Marshall, C., Feuk, L. & Scherer, S. W. Copy-number variation in control population cohorts. Hum. Mol. Genet. 16, R168 (2007).
Cho, E. K., Tchinda, J., Freeman, J. L., Chung, Y. J., Cai, W. W. & Lee, C. Array-based comparative genomic hybridization and copy number variation in cancer research. Cytogenet. Genome Res. 115, 262–272 (2006).
McCarroll, S. A. & Altshuler, D. M. Copy-number variation and association studies of human disease. Nat. Genet. 39, S37–S42 (2007).
Rodriguez-Revenga, L., Mila, M., Rosenberg, C., Lamb, A. & Lee, C. Structural variation in the human genome: the impact of copy number variants on clinical diagnosis. Genet. Med. 9, 600 (2007).
Stranger, B. E., Forrest, M. S., Dunning, M., Ingle, C. E., Beazley, C., Thorne, N. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848 (2007).
Nana-Sinkam, S. P. & Croce, C. M. MicroRNAs as therapeutic targets in cancer. J. Lab. Clin. Med. 157, 216–225 (2011).
Grubor, V., Krasnitz, A., Troge, J. E., Meth, J. L., Lakshmi, B., Kendall, J. T. et al. Novel genomic alterations and clonal evolution in chronic lymphocytic leukemia revealed by representational oligonucleotide microarray analysis (ROMA). Blood 113, 1294 (2009).
Costello, J. F., Frühwald, M. C., Smiraglia, D. J., Rush, L. J., Robertson, G. P., Gao, X. et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat. Genet. 24, 132–138 (2000).
Lübbert, M. DNA methylation inhibitors in the treatment of leukemias, myelodysplastic syndromes and hemoglobinopathies: clinical results and possible mechanisms of action. Curr. Top. Microbiol. Immunol. 249, 135 (2000).
Raval, A., Tanner, S. M., Byrd, J. C., Angerman, E. B., Perko, J. D., Chen, S. S. et al. Downregulation of death-associated protein kinase 1 (DAPK1) in chronic lymphocytic leukemia. Cell 129, 879–890 (2007).
Rabionet, R., Jaworski, J. M., Ashley-Koch, A. E., Martin, E. R., Sutcliffe, J. S., Haines, J. L. et al. Analysis of the autism chromosome 2 linkage region: GAD1 and other candidate genes. Neurosci. Lett. 372, 209–214 (2004).
Bhuiyan, Z. A., Hamdan, M. A., SHAMSI, E. T. A., Postma, A. V., Mannens, M.M.A.M., WILDE, A. A. M. et al. A novel early onset lethal form of catecholaminergic polymorphic ventricular tachycardia maps to chromosome 7p14-p22. J. Cardiovasc. Electrophysiol. 18, 1060–1066 (2007).
Sano, R., Miki, T., Suzuki, Y., Shimada, F., Taira, M., Kanatsuka, A. et al. Analysis of the insulin-sensitive phosphodiesterase 3B gene in type 2 diabetes. Diabetes Res. Clin. Pract. 54, 79–88 (2001).
Rastogi, A., Zai, C., Likhodi, O., Kennedy, J. L. & Wong, A. H. Genetic association and post-mortem brain mRNA analysis of DISC1 and related genes in schizophrenia. Schizophr. Res. 114, 39–49 (2009).
Munshi, A. & Kaul, S. Stroke genetics–focus on PDE4D gene. Int. J. Stroke 3, 188–192 (2008).
Waldkirch, E., Ückert, S., Sigl, K., Langnaese, K., Richter, K., Stief, C. G. et al. Expression of cAMP-dependent protein kinase isoforms in the human prostate: functional significance and relation to PDE4. Urology 76, 515.e8-14 (2010).
Gretarsdottir, S., Thorleifsson, G., Reynisdottir, S. T., Manolescu, A., Jonsdottir, S., Jonsdottir, T. et al. The gene encoding phosphodiesterase 4D confers risk of ischemic stroke. Nat. Genet. 35, 131–138 (2003).
Ross, O. A., Worrall, B. B. & Meschia, J. F. Advancing stroke therapeutics through genetic understanding. Curr. Drug Targets 8, 850–859 (2007).
Newcombe, P. J., Verzilli, C., Casas, J. P., Hingorani, A. D., Smeeth, L. & Whittaker, J. C. Multilocus Bayesian meta-analysis of gene-disease associations. Am. J. Hum. Genet. 84, 567–580 (2009).
Xue, H., Wang, H., Song, X., Li, W., Sun, K., Zhang, W. et al. Phosphodiesterase 4D gene polymorphism is associated with ischaemic and haemorrhagic stroke. Clin. Sci. 116, 335 (2009).
Lövkvist, H., Smith, J. G., Luthman, H., Höglund, P., Norrving, B., Kristoffersson, U. et al. Ischaemic stroke in hypertensive patients is associated with variations in the PDE4D genome region. Eur. J. Hum. Genet. 16, 1117–1125 (2008).
Kim, M. K., Kim, J. T., Choi, S. M., Lee, S. H., Park, M. S. & Cho, K. H. Phosphodiesterase4D gene and risk of noncardiogenic ischemic stroke in a Korean population. J. Korean Med. Sci. 24, 307–310 (2009).
Sun, Y., Huang, Y., Chen, X., Liu, Y., Lu, X., Shi, Y. et al. Association between the PDE4D gene and ischaemic stroke in the Chinese Han population. Clin. Sci. 117, 265–272 (2009).
Bondarenko, E. A., Tupitsyna, T. V., Slominski, P. A., Shetova, I. M., Shamalov, N. A., Botsina, A. I. et al. [Phosphodiesterase 4D (PDE4D) gene polymorphism in patients with acute stroke from Moscow]. Genetika 46, 861–864 (2010).
Bevan, S., Dichgans, M., Gschwendtner, A., Kuhlenbäumer, G., Ringelstein, E. B. & Markus, H. S. Variation in the PDE4D gene and ischemic stroke risk: a systematic review and meta-analysis on 5200 cases and 6600 controls. Stroke 39, 1966–1971 (2008).
Matsushita, T., Kubo, M., Yonemoto, K., Ninomiya, T., Ashikawa, K., Liang, B. et al. Lack of association between variations of PDE4D and ischemic stroke in the Japanese population. Stroke 40, 1245–1251 (2009).
Munshi, A., Babu, M. S., Kaul, S., Shafi, G., Anila, A. N., Alladi, S. et al. Phosphodiesterase 4D (PDE4D) gene variants and the risk of ischemic stroke in a South Indian population. J. Neurol. Sci. 285, 142–145 (2009).
Quarta, G., Stanzione, R., Evangelista, A., Zanda, B., Di Angelantonio, E., Marchitti, S. et al. Phosphodiesterase 4D and 5-lipoxygenase activating protein genes and risk of ischemic stroke in Sardinians. Eur. J. Hum. Genet. 17, 1448–1453 (2009).
Homma, S., Sakamoto, T., Hegab, A. E., Saitoh, W., Nomura, A., Ishii, Y. et al. Association of phosphodiesterase 4D gene polymorphisms with chronic obstructive pulmonary disease: relationship to interleukin 13 gene polymorphism. Int. J. Mol. Med. 18, 933–939 (2006).
Gottlieb, D. J., O’Connor, G. T. & Wilk, J. B. Genome-wide association of sleep and circadian phenotypes. BMC Med. Genet. (8 Suppl 1), S9 (2007).
Calboli, F. C. F., Tozzi, F., Galwey, N. W., Antoniades, A., Mooser, V., Preisig, M. et al. A genome-wide association study of neuroticism in a population-based sample. PloS One 5, e11504 (2010).
Heck, A., Lieb, R., Unschuld, P. G., Ellgas, A., Pfister, H., Lucae, S. et al. Evidence for associations between PDE4D polymorphisms and a subtype of neuroticism. Mol. Psychiatry 13, 831–832 (2008).
Yoshida, T., Kato, K., Yokoi, K., Oguri, M., Watanabe, S., Metoki, N. et al. Association of gene polymorphisms with chronic kidney disease in high- or low-risk subjects defined by conventional risk factors. Int. J. Mol. Med. 23, 785–792 (2009).
Liao, Y. C., Lin, H. F., Guo, Y. C., Yu, M. L., Liu, C. K. & Juo, S.- H. H. Sex-differential genetic effect of phosphodiesterase 4D (PDE4D) on carotid atherosclerosis. BMC Med. Genet. 11, 93 (2010).
Melén, E., Himes, B. E., Brehm, J. M., Boutaoui, N., Klanderman, B. J., Sylvia, J. S. et al. Analyses of shared genetic factors between asthma and obesity in children. J. Allergy Clin. Immunol. 126, 631.e1-8–637.e1-8 (2010).
Bryniarski, L., Rzepecki, M., Klocek, M. & Wyczołkowski, M. [The safety of 5-phosphodiesterase inhibitors in the treatment of erectile dysfunction in patients with cardiovascular disease]. Przegl. Lek. 66, 192–197 (2009).
Roberts, K. E., Fallon, M. B., Krowka, M. J., Brown, R. S., Trotter, J. F., Peter, I. et al. Genetic risk factors for portopulmonary hypertension in patients with advanced liver disease. Am. J. Respir. Crit. Care Med. 179, 835–842 (2009).
Hahn, W- H, Suh, J- S & Cho, B- S Phosphodiesterase-5gene (PDE5A) polymorphisms are associated with progression of childhood IgA nephropathy. Pediatr. Nephrol. 25, 1663–1671 (2010).
Corton, M., Blanco, M. J., Torres, M., Sanchez-Salorio, M., Carracedo, A. & Brion, M. Identification of a novel mutation in the human PDE6A gene in autosomal recessive retinitis pigmentosa: homology with the nmf28/nmf28 mice model. Clin. Genet. 78, 495–498 (2010).
Riazuddin, S. A., Zulfiqar, F., Zhang, Q., Yao, W., Li, S., Jiao, X. et al. Mutations in the gene encoding the alpha-subunit of rod phosphodiesterase in consanguineous Pakistani families. Mol. Vis. 12, 1283–1291 (2006).
Gal, A., Orth, U., Baehr, W., Schwinger, E. & Rosenberg, T. Heterozygous missense mutation in the rod cGMP phosphodiesterase beta-subunit gene in autosomal dominant stationary night blindness. Nat. Genet. 7, 64–68 (1994).
Danciger, M., Blaney, J., Gao, Y. Q., Zhao, D. Y., Heckenlively, J. R., Jacobson, S. G. et al. Mutations in the PDE6B gene in autosomal recessive retinitis pigmentosa. Genomics 30, 1–7 (1995).
Sakamoto, K., McCluskey, M., Wensel, T. G., Naggert, J. K. & Nishina, P. M. New mouse models for recessive retinitis pigmentosa caused by mutations in the Pde6a gene. Hum. Mol. Genet. 18, 178–192 (2009).
Hmani-Aifa, M., Benzina, Z., Zulfiqar, F., Dhouib, H., Shahzadi, A., Ghorbel, A. et al. Identification of two new mutations in the GPR98 and the PDE6B genes segregating in a Tunisian family. Eur. J. Hum. Genet. 17, 474–482 (2008).
Gao, Y. Q., Danciger, M., Longmuir, R., Piriev, N. I., Zhao, D. Y., Heckenlively, J. R. et al. Screening of the gene encoding the alpha-subunit of cone cGMP-PDE in patients with retinal degenerations. Invest. Ophthalmol. Vis. Sci. 40, 1818–1822 (1999).
Wyszynski, D. F., Baldwin, C. T., Cleves, M. A., Amirault, Y., Nolan, V. G., Farrell, J. J. et al. Polymorphisms near a chromosome 6q QTL area are associated with modulation of fetal hemoglobin levels in sickle cell anemia. Cell. Mol. Biol. 50, 23–33 (2004).
Chen, C., Wickenheisser, J., Ewens, K. G., Ankener, W., Legro, R. S., Dunaif, A. et al. PDE8A genetic variation, polycystic ovary syndrome and androgen levels in women. Mol. Hum. Reprod. 15, 459–469 (2009).
Arnaud-Lopez, L., Usala, G., Ceresini, G., Mitchell, B. D., Pilia, M. G., Piras, M. G. et al. Phosphodiesterase 8B gene variants are associated with serum TSH levels and thyroid function. Am. J. Hum. Genet. 82, 1270–1280 (2008).
Shields, B. M., Freathy, R. M., Knight, B. A., Hill, A., Weedon, M. N., Frayling, T. M. et al. Phosphodiesterase 8B gene polymorphism is associated with subclinical hypothyroidism in pregnancy. J. Clin. Endocrinol. Metab. 94, 4608–4612 (2009).
Libé, R., Fratticci, A., Coste, J., Tissier, F., Horvath, A., Ragazzon, B. et al. Phosphodiesterase 11A (PDE11A) and genetic predisposition to adrenocortical tumors. Clin. Cancer Res. 14, 4016–4024 (2008).
Horvath, A., Korde, L., Greene, M. H., Libe, R., Osorio, P., Faucz, F. R. et al. Functional phosphodiesterase 11A mutations may modify the risk of familial and bilateral testicular germ cell tumors. Cancer Res. 69, 5301–5306 (2009).
Peverelli, E., Ermetici, F., Filopanti, M., Elli, F. M., Ronchi, C. L., Mantovani, G. et al. Analysis of genetic variants of phosphodiesterase 11A in acromegalic patients. Eur. J. Endocrinol. 161, 687–694 (2009).
DeWan, A. T., Triche, E. W., Xu, X., Hsu, L.- I., Zhao, C., Belanger, K. et al. PDE11A associations with asthma: results of a genome-wide association scan. J. Allergy Clin. Immunol. 126, 871–873.e9 (2010).
Acknowledgements
This work was supported by grants from the Lymphoma and Leukemia Society, NIH and FCVI-HGUA (Fundación de la Comunidad Valenciana para la investigación en el Hospital General Universitario de Alicante) and Conselleria de Sanitat Valenciana, Spain.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Peiró, A., Tang, CM., Murray, F. et al. Genetic variation in phosphodiesterase (PDE) 7B in chronic lymphocytic leukemia: overview of genetic variants of cyclic nucleotide PDEs in human disease. J Hum Genet 56, 676–681 (2011). https://doi.org/10.1038/jhg.2011.80
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.80