Abstract
Although evidence suggests an importance of genetic factors in the development of breast cancer in Taiwanese (ethnic Chinese) women, including a high incidence of early-onset and secondary contralateral breast cancer, a major breast cancer predisposition gene, BRCA1, has not been well studied in this population. In fact, the carcinogenic impacts of many genetic variants of BRCA1 are unknown and classified as variants of uncertain significance (VUS). It is therefore important to establish a method to characterize the BRCA1 VUSs and understand their role in Taiwanese breast cancer patients. Accordingly, we developed a multimodel assessment strategy consisting of a prescreening portion and a validated functional assay to study breast cancer patients with early-onset, bilateral or familial breast cancer. We found germ-line BRCA1 mutations in 11.1% of our cohort and identified one novel missense mutation, c.5191C>A. Two genetic variants were initially classified as VUSs (c.1155C>T and c.5191C>A). c.1155C>T is not predicted to be deleterious in the prescreening portion of our assessment strategy. c.5191C>A, on the other hand, causes p.T1691K, which is predicted to have high deleterious probability because of significant structural alteration, a high deleterious score in the predictive programs and, clinically, triple negative characteristics in breast tumors. This mutant is confirmed by transcription activation and yeast growth-inhibition assays. In conclusion, we show as high a prevalence of germ-line BRCA1 mutation in high-risk Taiwanese patients as in Caucasians and demonstrate a useful strategy for studying BRCA1 VUSs.
Similar content being viewed by others
Introduction
Breast cancer is one of the most common cancers and is the leading cause of cancer-related death in women throughout the world. Interestingly, the prevalence of breast cancer is lower in Taiwanese (ethnic Chinese) populations than in Caucasian populations,1 and the epidemiological patterns are different in the two patient groups. First, the median age of breast cancer patients in Taiwan (45–49 years) is lesser than that in Western countries (70–74 years).2 Second, a rise in incidence has been observed consistently in Asian countries,2, 3, 4 and one age-period-cohort analysis showed that the increase in breast cancer incidence was largely due to a rise in young patients.2 Third, a longitudinal cohort revealed that young breast cancer patients in Taiwan carried a greater risk for and shorter interval to progression to secondary contralateral primary breast cancer than in western women.5, 6 As the young age and bilateral breast cancer are more likely to be related to genetic predisposal of factors,7, 8 it is important to identify potential predisposing factors in Taiwanese patients.
However, genetic factors related to breast cancer, including BRCA1, have not been well studied in the Asia-Pacific areas. Of the reported studies conducted in China, Korea, Japan and Malaysia, germ-line BRCA1 mutation occurred in approximately 4–9% of breast cancer patients.9, 10, 11, 12 This is lower than the ∼10% reported in non-Jewish Caucasian women with early-onset or a familial history of breast cancer.8 In Taiwan, two small studies found a few benign polymorphisms of BRCA1 without the identification of pathological mutants associated with breast cancer development.13, 14 Of these studies, only deletion or frame-shift mutation of the BRCA1 gene could be clearly defined as a deleterious mutant; other mutations, especially missense changes, were of uncertain significance with regard to breast cancer development and remained uncharacterized.
The BRCA1 gene contains 24 exons, spanning more than 70 kb in the genomic DNA, and encodes 1863 amino acids.15 This large gene displays several thousand different alleles, and most of them are missense changes according to the Breast Cancer Information Core (BIC) database. Categorizing missense changes as deleterious mutations associated with breast cancer development vs benign polymorphisms is difficult16 and poses significant obstacles in genetic studies.17 Currently, these mutations are generally classified as deleterious, benign polymorphisms or variants of uncertain significance (VUS).
Approaches to studying the mutation effects of disease genes usually proceeds in one of the two directions: clinical epidemiological studies and functional assays.18 However, evaluation of the phenotype–genotype correlation based on epidemiological data is often problematic because the sample size of rare genetic variants is insufficient for meaningful epidemiological analysis. Although functional assays provide supportive, qualitative evidence to characterize the mutant proteins, they are usually not available in clinical practice. Therefore, to better characterize germ-line BRCA1 mutations, here we establish an effective method to distinguish the deleterious mutation and benign polymorphism, and apply this method in the assessment of our breast cancer patients. This method is described as multimodel assessment, including mutation prescreening and validated functional assays. The prescreening aims to identify the high probable deleterious mutations. After initial database searching, the VUSs are studied followed by evaluation with pedigree analysis, tumor histopathology and bioinformatics. Finally, functional assays are performed with the possible deleterious mutations to prove their pathogenicity (Figure 1). In this way, we successfully identified and confirmed a novel mutation and demonstrated that the prevalence of BRCA1 mutation in Taiwanese high-risk women is similar as in Caucasians.
Materials and methods
Patients
A total of 36 women diagnosed with breast cancer between 2005 July and 2007 December in the National Taiwan University Hospital Breast Cancer Center were enrolled in this study. All patients were required to meet one of the following criteria: (1) early-onset breast cancer (age before 35 years) or bilateral breast cancer, (2) breast cancer onset before age 50 years together with at least one first or second-degree relative having breast cancer or ovarian cancer or (3) breast cancer after age 50 years, but with three relatives having breast cancer or one relative having ovarian cancer. Written consent was obtained from each participant. This study was approved by the Research and Ethical Committee of National Taiwan University Hospital (IRB 0941171001).
Mutation screening of BRCA1 gene
BRCA1 gene mutation screening was performed by using BRCAChip, a resequencing microarray for the detection of mutations in BRCA1, BRCA2 and TP53 genes and designed by Vita Genomics and manufactured by Affymetrix (Santa Clara, CA, USA) using photolithography and solid-phase DNA synthesis. Each microarray contained 28.2 kb of coding exon and flanking intron sequence (both sense and antisense) of BRCA1, BRCA2 and TP53 genes. In order to expand the detection of insertion/deletion mutation in BRCAChip, specific probes for 456 previously reported insertion/deletion mutations in BRCA1 and BRCA2 genes were also tiled on the BRCAChip.
Resequencing array assay was performed following Affymetrix's instructions (PN 701231). In brief, BRCA1 genes were amplified from genomic DNA by conventional PCR,19 and the amplified products were purified, quantitated, and pooled in equimolar amount for fragmentation, labeling and hybridization with BRCA resequencing chip. After 16 h of hybridization, the chip was washed and stained for scanning. The scanned image was analyzed with Affymetrix GeneChip Operating Software. The intensity of each probe set and base calling of each nucleotide were analyzed by GeneChip Sequence Analysis Software (Affymetrix, Santa Clara, CA, USA) followed by visual inspection and confirmation. Genetic variants were detected by comparison with a consensus wild-type sequence constructed for each gene. All potential genetic variants were independently confirmed by repeated PCR amplification of the indicated gene region(s) and direct sequencing in both forward and reverse directions.
Mutation database searching
The genetic variants found in our patients were compared with on-line published databases: BIC (http://www.research.nhgri.nih.gov/bic/) and ENTREZSNP (http://www.ncbi.nlm.nih.gov). Large-scale deletion, frame-shift mutation and nonsense mutation of BRCA1 are considered to be deleterious mutations. The missense changes could be classified as deleterious, benign polymorphism and VUS. Some specific deleterious or benign genetic variants have been documented, such as S1613G (benign polymorphism) and M1775R (deleterious mutation).16 For VUS, we used multimodel assessment to evaluate the pathogenicity (Figure 1).
Family history and tumor histopathology
Patients' clinical information including age, sex, tumor histological type and cancer history of family members were analyzed. Tumor stage was defined according to the sixth edition of the American Joint Committee on Cancer staging system. The tumor histological grade was evaluated by the Nottingham combined histological grading system. The expression of estrogen receptor (ER), progesterone receptor (PR) and HER2 in the tumor tissues was determined by immunohistochemical staining. ER or PR was considered positive when more than 10% of tumor cells showed positive staining. For HER-2 staining, a score of 3+ was considered positive and a specimen with a score of 2+ was determined by fluorescence in situ hybridization analysis.
Bioinformatics analysis
Sequence alignment
Highly conserved amino-acid sequences were considered as important sites of protein function.20 Variants appearing in the high-degree conservation sequence are more likely to be pathogenic than those in the low-degree conservation sequence. We searched the full-length BRCA1 sequence of the Vertebrata in the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/). Sequence alignment was performed by using the ClustalX 2.0.12 software (University College Dublin, Dublin, Ireland, http://www.clustal.org).
Structural modeling
For comparative structural modeling, we created mutants using the MODELLER program21 based on the template of human wild-type BRCA1 BRCT domain interacting with a BACH1 phosphorylated peptide (PDB code: 1T29). The 3-dimensional structure of the mutation was built using UCSF Chimera program (University of California, San Francisco, CA, USA, http://www.cgl.ucsf.edu/chimera/).
Web-servers prediction programs
We not only accessed the pathogenicity of missense mutations by the above predictive methods but also compared the results with three bioinformatics missense variants prediction web-servers: PANTHER (Protein ANalysis THrough Evolutionary Relationships22), PolyPhen-2 (Polymorphism Phenotyping, v223) and CanPredict.24
PANTHER (http://www.pantherdb.org/tools/csnpScoreForm.jsp) is a classification system used to determine the association of protein sequence relationships with function relationships. It uses the Hidden Markov Model for scoring the ‘functional likelihood’ of a missense mutation based on the evolutionarily conserved sequences. It generates a ‘substitution position-specific evolutionary conservation’ (subPSEC) score for an amino-acid substitution. The scores are continuous from 0 (benign) to –10 (high probability of deleterious). PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2/) is a second generation of PolyPhen capable of predicting the pathogenicity of an amino-acid substitution mainly by the structural alteration. This tool calculates the PSIC score (Position-Specific Independent Counts) of the amino-acid substitution to probabilistically classify the mutation as benign, possibly damaging or probably damaging. The higher the PSIC score (ranging from 0 to 1), the greater is the probability of pathological damage to the protein structure and function. CanPredict (http://www.cgl.ucsf.edu/Research/genentech/canpredict/) is a program used to distinguish cancer-associated missense changes from those that are benign polymorphisms. By using two computational metrics (Sorting Intolerant from Tolerant (SIFT) and the Pfam-based LogR.E-value method), as well as Gene Ontology Similarity Score, this approach increases the ability to predict the cancer relevance of missense mutations. SIFT, a sequence homology-based tool, shows that well-conserved positions between protein species tend to be predicted as deleterious. The LogR.E-value is the calculated difference between wild-type and variant proteins as based on the Pfam motif model. CanPredict categorizes missense mutations as likely cancer, likely non-cancer or not determined.
Functional assays
Quantitative assessment of transcriptional activation in mammalian cells
Transcription-based assay could characterize the deleterious mutations of BRCA1 COOH-terminus.16 To establish this functional assay, we constructed a wild-type cDNA of BRCA1 into pcDNA3.1+ vector, then introduced the BRCA1 C-terminus (cDNA 4186–5792, amino acid 1396-1863) to fuse with GAL4 DBD (DNA binding domain) into pGBT9 vector by the restriction site of BamHI and EcoRI. Then, the fusion segment of GAL4 DBD-BRCA1 C-terminus was subcloned into pcDNA 3.1+ vector using the restriction site of HindIII and BamHI. Site-directed mutagenesis was performed to create T1691K, M1775R and S1613G mutants (Supplementary Table 1), according to the manual of QuikChange Site-Directed Mutagenesis Kit (Stratagene, Agilent Technologies, La Jolla, CA, USA).
The pGL4.31[luc2P/GAL4UAS/Hygro] vector (Promega, Madison, WI, USA) was used as a reporter in this assay, and transfection was normalized with the internal control pGL4.74[hRluc/TK] Vector (Promega). Human 293T cells were cultured in 24-well plates with Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum. After 24 h, 293T cells were co-transfected with pcDNA 3.1+ constructed with GAL4 DBD-BRCA1 C-terminus (wild type and mutants), pGL4.31 and pGL4.74 by using calcium phosphate transfection method. Luciferase assay was performed 36-h post transfection by using a Dual-GLO luciferase assay system (Promega), and cells were also harvested for western blot. Cell lysates were electrophoresed in 10% SDS-PAGE, and probed with an anti-GAL4 DBD monoclonal antibody (Clontech, TAKARA BIO, Mountain View, CA, USA).
Yeast growth assay
We constructed wild type and mutations (S1613G, M1775R and T1691K) of BRCA1 COOH-terminus (amino acid 1396–1863) into pACT2 vectors. The diploid strain BY4741 of Saccharomyces cerevisiae, kindly provided by Professor Lee FJ, has the genotype of MATa his3, leu2, met15, ura3. Yeast was transformed with plasmid DNA by using the lithium acetate method with single-stranded DNA as a carrier, and plated on solid synthetic medium with 2% glucose lacking leucine. Colonies grew for 3 days at 30 °C. BY4741 transformants containing pACT2 constructs (vector alone, wild type, S1613G, M1775R and T1691K) were grown overnight in liquid medium lacking leucine to reach saturationed , and then inoculated into the fresh medium from the initial OD600 of 0.2 at 30 °C in a shaker. The OD600 was checked at 0, 3, 6 and 9 h. Yeast transformants were collected to determine the expression of BRCA1 C-terminus by using western blot probed with anti-BRCA1 antibody C-20 (Santa Cruz Biotechnology, Santa Cruz, CA, USA).
Results
BRCA1 sequence analysis
Among the 36 patients, 18 genetic variants were found, including 14 variants located in exons and 4 variants in exon–intron boundaries (Table 1). The 14 exon sequence variations consisted of two frame-shift mutations, one nonsense mutation, six missense mutations and five synonymous nucleotide polymorphisms with no amino-acid substitution (Table 1). We first searched the BIC database for the 18 genetic variants. Three genetic variants (c.5149del4, c.5454delC and c.5655C>T) were defined as deleterious mutations associated with breast cancer. Four genetic variants had not been described in the BIC database.
Among the six missense mutations, four were classified as benign polymorphisms according to the BIC database: c.2731C>T, c.3432A>G, c.3667A>G and c.4956A>G, which led to P871L, E1038G, K1183R and S1613G, respectively. Another two missense mutations (c.1155C>T and c.5191C>A) were classified as VUS.
Analysis of the two BRCA1 VUS
Pedigree analysis
Two missense mutations of VUS were c.1155C>T in exon 11 and c.5191C>A in exon 17, which resulted in P346S and T1691K in BRCA1, respectively. The c.1155C>T missense mutation occurred in a 44-year-old woman, whose mother developed breast cancer at the age of 63 years. T1691K was identified form a 32-year-old woman, and breast cancer familial history was found in her mother and two maternal aunts at the age of 44, 48 and 57 years, respectively (Table 2).
Tumor histopathology
The dominant histopathological feature of breast cancer arising from BRCA1 mutation is high grade infiltrating ductal carcinoma (IDC), and the immunohistochemical staining is negative expression of ER, PR and HER2 (triple negative). In the modified Manchester scoring prediction system, grade 3 IDC with triple negative immunotyping contributes a significant impact in the prediction of BRCA1 mutant, but positive expression of HER2 diminishes the predictive possibility of BRCA1 mutation.25 The patient with c.1155C>T developed a grade 2 IDC with negative staining of ER, PR and positive expression of HER2; the other patient with c.5191C>A mutation had a grade 3 IDC with triple negative immunohistochemical staining. On the basis of the pathological feature, c.1155C>T of BRCA1 is less likely to be a pathological mutant and c.5191C>A mutation is of strong deleterious significance.
Bioinformatics study
Sequence alignment in BRCA1
We identified 13 full-length BRCA1 sequences of the Vertebrata in the National Center for Biotechnology Information (Supplementary Table 2). Multi-alignment comparisons show high levels of amino-acid identity in the ring domain, BRCT-N and BRCT-C region between humans and other vertebrata (Figure 2a). Amino acid threonine of codon 1691 (T1691) is located in the BRCT-N, and is 100% conserved between Homo sapiens and Tetraodon. Codon 346 is not located in the ring domain or BRCT regions; the conservation is lower at its nearby region between humans and other vertebrata. Codon 346 proline (P346) is only conserved in primates and Bovidae, and changed to serine in Rattus, Mus and Gallus. On the basis of the levels of conservation observed, missense mutation T1691K is likely to affect protein function whereas P346S is not.
(a) Sequence alignment in BRCA1. P346 is conserved only in the primates and Bovidae, and changed to Ser in Rattus, Mus and Gallus. T1691 is located in the BRCT-N, and conserved in all Vertebrata. (b) Ribbon presentation of the T1691K mutant structure modeling (purple) superimposed with wtBRCA1 (white) structure interacting with BACH1 phosphorylated peptide; the interacting surface is shown as the mesh. T1691 is located in the loop between the third β-strand and second α-helix of BRCT-N, which contribute to protein surface interactions with the phosphorylated BACH1 peptide. (c) Two hydrogen bonds (blue dotted lines) from the T1691 to S1715 and D1692 may contribute to stabilizing the structure and the binding surface. Superposition of lysine substitution disrupts the hydrogen bonds and causes steric hindrance between K1691 and S1715 side chains.
Structure modeling
The BRCA1 missense mutation T1691K lies at the loop between the third β-sheet and second α-helix of BRCT-N. To investigate the structural consequences of this missense mutation, we used the X-ray crystal structure of wild type (wt) BRCA1 COOH terminus (PDB:1T29) as a template to generate T1691K point mutation on it. In the wtBRCA1, the loop in which T1691 is located forms the protein surface and is a part of the binding pocket interacting with the BACH1 phosphorylated peptide26, 27 (Figure 2b). The buried T1691 residue makes the hydrogen bond to S1715 and D1692, as a contribution to hold the structure of protein surface and the binding pocket (Figure 2c). As a result of T1691K mutation, the two hydrogen bonds are disrupted. Moreover, steric hindrance develops because of the significantly greater length of the lysine residue compared with threonine. The bump check confirmed that the interatomic distances are too short between the side chains of K1691 and S171528 (Supplementary Table 3). Therefore, T1691K causes steric hindrance and losses of hydrogen bonds, which would no longer stabilize the local structure but significantly alter the binding surface. The structural analysis also compares well with the interspecies sequence alignment; the sequence of this loop and S1715 are 100% conserved between humans and Tetraodon. This fact may reflect that the loop has the important role for interacting with the phosphorylated peptide.26
Web-servers prediction programs
Bioinformatics analysis of missense mutation was performed by PolyPhen-2, PANTHER and CanPredict to calculate the impact of the mutations. For the two VUSs, T1691K mutation was calculated as 0.993 (probably damaging) by PolyPhen-2, 0.90404 (deleterious) by PANTHER and ‘likely cancer’ by CanPredict; P346S was calculated as 0 (benign) by PolyPhen-2, 0.24245 (not deleterious) by PANTHER and not determined by CanPredict (not found in the Pfam database). Thus, T1691K was consistently determined to be a deleterious mutation by the three programs, and P346S was considered as a benign polymorphism. The results of the web-servers predictions support the result of the two bioinformatics analyses described above. Therefore, P346S was not analyzed further.
Functional assay
Quantitative assessment of transcriptional activation in mammalian cells
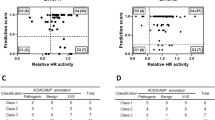
The known mutation M1775R was used as a deleterious control, and benign polymorphism S1613G and wt BRCA1 (amino acids 1396–1863) were used as neutral controls. The expression levels of DBD–BRCA1 fusion protein were comparable for wild type and variants in mammalian cells. Variant M1775R displayed marked reduction of relative luciferase activity (∼2% of wt) and S1613G displayed slightly increased relative luciferase activity (∼120% of wt). The test VUS T1691K also showed significant reduced activity (∼2% of wt), similar to the known mutation M1775R (Figure 3b). This quantitative assessment revealed that T1691K mutant protein lacked transcriptional activation activity, indicating that T1691K is a deleterious mutation.
Functional assays of BRCA1 genetic variants. (a) Location of missense variant (T1691K), benign polymorphism (S1613G) and deleterious control (M1775R). The fusion protein DBD–BRCA1 C-terminus was subcloned into pcDNA3.1+ vector for quantitative transcriptional activation assay. The wt BRCA1 or mutant BRCA1 COOH-terminus was constructed into pACT2 for yeast growth curve assay. (b) Quantitative transcriptional activation assay in mammalian cells. Cells were cotransfected with a pGL4.31vector (GAL4-responsive firefly luciferase reporter), pGL4.74 vector (Renilla luciferase driven by a constitutive promoter as internal control) and a pcDNA3.1+ construct containing GAL4 DBD fused with wt BRCA1 or mutant BRCA1 COOH-terminus. Measurements were done in triplicates and normalized against the internal transfection Renilla luciferase. Wt BRCA1 was used as the control (100%) of relative luciferase activity. (c) Determination of the expression of BRCA1 C-terminus by using western blot with mouse anti-GAL4 DBD monoclonal antibody in mammalian cell extracts. (d) S. cerevisiae was transformed with pACT2 contained with wt BRCA1 or mutant BRCA1 COOH-terminus. Colonies were grown for 3 days at 30 °C on solid synthetic medium with 2% glucose lacking leucine. Grossly, the colony size of transformants with pACT2 vector alone or T1691K was larger than that with wt BRCA1 or S1613G. (e) Growth curves of S. cerevisiae transformed with pACT2 constructs (vector alone, wild type, S1613G, M1775R and T1691K) in YPD medium lacking leucine. The initial OD600 was 0.2, and all transformants grew in a 30 °C shaker. The OD600 was checked at 0, 3, 6 and 9 h. (f) Determination of the expression of BRCA1 C-terminus by using western blot probed with anti-BRCA1 antibody C-20.
Yeast growth assay
The expression of wt BRCA1 protein inhibits S. cerevisiae growth.29 Taking the phenotype, the suppression of growth curve can be used to distinguish deleterious mutants from benign polymorphisms. Protein expression levels were similar for all constructs by western blotting. Yeasts containing wt BRCA1 or S1613G constructs were fully suppressed in the growth curves, indicating that S1613G is actually a benign polymorphism. The known mutant M1775R showed a partial growth suppression phenotype. Yeast transformed with T1691K construct exhibited a growth curve similar to that of the transformants containing pACT2 vector alone (Figure 3e), which was consistent with the evidence that T1691K has no effect on growth inhibition. This phenotype suggests that T1691K is a deleterious mutant.
Clinical patient characteristics
In the 36 patients, four deleterious mutations of BRCA1 were confirmed by the multimodel assessment, resulting in an 11.1% incidence in this case series. The four mutants consisted of one novel (c.5191C>A) and three known mutations (c.5149del4, c.5454delC and c.5655C>T). All four patients with deleterious BRCA1 mutation were diagnosed with IDC before age 35, and have familial breast cancer histories. Two of them had developed secondary contralateral breast cancer at the time of this report. The patient with c.5149del4 had grade 3 bilateral IDC, and one first degree relative and one second degree relative with breast cancer. The patient with c.5454delC had one first degree relative and one second degree relative with breast cancer. Mutation c.5655C>T occurred in a patient presenting with bilateral breast cancer; the sister of this patient also had breast cancer. Mutation c.5191C>A presented in a patient with one first degree relative and two second degree relatives having breast cancer (Table 2).
Among the 36 patients, patients with BRCA1 deleterious mutation are significantly younger (32.7 vs 37.7 years, P=0.023) and are more likely to have a history of familial breast cancer (P=0.025) than patients without deleterious mutation. By using univariate analysis, negativity of ER expression (P=0.005) and triple negativity of IDC (P=0.019) comprises the significant dominant pathology types of patients with deleterious mutation. Other characteristics, including pathological stage, histological type, grade and HER2 expression are not significantly different.
Discussion
In this study, we analyzed germ-line BRCA1 mutation in 36 high-risk breast cancer patients and found 11.1% prevalence in this patient cohort. By the multimodel assessment, we identify a novel deleterious mutation c.5191C>A (p.T1691K) that affects BRCA1 protein function and could be relevant to breast cancer development. We also demonstrate in this cohort that patients with BRCA1 deleterious mutation are younger, have a more dominant familial history and are more likely to have triple negative (negative expression of ER, PR and HER2) histopathological characteristics when compared with non-BRCA1 mutants.
This is the first study to provide evidence that germ-line BRCA1 mutation has an important genetic impact in breast cancer patients with early-onset and/or familial history in the Taiwanese population. Among the four mutants, c.5149del4 has been widely reported in Western countries.30 c.5454delC is also found in Caucasians and reported as a founder mutation in Malay and Philippine breast cancer patients.31, 32 c.5655C>T and c. 5191C>A are recorded twice and once in the BIC database, respectively. Other genetic variants, such as c.2201C>T, c.2430T>C, c.2731C>T, c.3432A>G, c.3667A>G, c.4427T>C and c.4956A>G, have been reported in Western and Asian countries.12 Our cohort shows that the genetic spectrum of germ-line BRCA1 in the Chinese population of Taiwan is similar with the other races of the world.
The 11.1% prevalence of this case series is similar with reports of non-Ashkenazi Jewish Caucasian women, but the prevalence is higher compared with prior Asia-Pacific studies. Prior studies from the Chinese population showed 5.9 or 2.8% of germ-line BRCA1 mutation in women with familial breast cancer or with breast cancer before 40 years.9 Malaysia presented 2.8% germ-line BRCA1 mutation in early-onset breast cancer patients without familial history.10 One Korean study demonstrated 6.8% prevalence of germ-line BRCA1 mutation from 354 patients with familial breast cancer.11 Possible reasons for the higher prevalence in our cohort are the small patient number and slightly different enrollment criteria used in the study. Also, VUS of BRCA1 was not characterized in the prior Chinese, Korean and Malaysia studies, and these studies only included frame-shift mutations and little known missense mutations as deleterious mutants. This reflects the fact that VUS (especially missense changes) of BRCA1 poses a problem in the genetic study of breast cancer.17
Currently, functional assays are required to characterize the contribution of BRCA1 VUS for breast cancer development, and at least five functional assays have been reported.33 However, the sensitivity and specificity of the different types of functional assays used to determine the VUS of BRCA1 are variable.33 It is a difficult and impractical task to perform all of the functional assays, and characterizing all the VUSs identified in patients using the functional assays is not an effective and efficient strategy. Therefore, we devised the multimodel assessment using clinical information (pedigree and pathological study) and bioinformatics analysis for pre-screening the VUS, excluding the obvious non-deleterious VUS, and selecting the possible deleterious VUS for further assessment by validated functional assays. First, to improve the prediction accuracy, we add tumor histopathology to the clinical pedigree assessment. Negative expression of ER or triple negative type is the major feature in breast cancer arising from germ-line BRCA1 mutation.25 Second, we use several bioinformatics methods to definite the impact of amino-acid substitution on the protein function. In our study, we demonstrate that the novel mutation T1691K is located in the high-degree sequence conserved region, and the three-dimensional simulation structure shows that change of this region influences the interaction with BACH1. Unlike previous simulation models in which M1775R or M1775K directly cause steric hindrance in binding with the phosphorylated peptide,34 T1691K may significantly alter the protein surface leading to disruption in BACH1 interaction. This observation is consistent with the previous observation that the loop containing T1691 is critical for BACH1 binding.26
Web-severe programs in our multimodel assessment strategy of mutants shows that T1691K was predicted as a deleterious mutant based on a high score. However, PolyPhen and PANTHER may overpredict damaging classes and SIFT (CanPredict) may overpredict tolerated classes of mutations.35 These limitations of the classification systems are observed in the benign control S1613G, which is predicted as damaging, although the impact score is lower than true mutants (Table 2, T1691K and M1775R). Therefore, bioinformatics analyses provide the predictive value, but the functional assay is still required to validate the mutation. Two sets of functional assays, including transcriptional activation assay and yeast growth inhibition, are important to provide qualitative evidence.33 T1691K is confirmed as a deleterious mutation, and S1613G is proved to be a benign polymorphism, consisted with previous observations. Therefore, an effective and robust assessment model is established for the analysis of BRCA1 VUS.
The development of multimodel assessment is based on the well-known protein structure and functions. The highly conserved sequence, well-known structure and functions of BRCA1 are only in the C-terminus (two BRCT domains) and the Ring domain. Functional assay of transcriptional activation is used to evaluate mutations in the C-terminus of BRCA1, and yeast growth inhibition is used for two BRCT domains.33 For Ring domain, changing the functional analysis to BARD1 binding assay may be a feasible way.36 Although the currently designed multimodel assessment is limited in the C-terminus of BRCA1, most functionally compromised mutants are located in the C-terminus and Ring domain. Therefore, the strategy of multimodel assessment is clinically applicable for evaluating most VUSs of BRCA1.
The multimodel assessment strategy described here not only improves genetic counseling but also identifies the specific patient group with BRCA1 mutation that would be appropriate to receive new targeted therapy. A poly(ADPribose) polymerase (PARP) inhibitor, for example, was shown to have promising activity in the treatment of BRCA-deficient breast and ovary cancers in phase I and II clinical trials.37, 38 These facts underscore the growing importance of establishing comprehensive methods for the accurate classification of BRCA1 alleles. Though the patient number is limited in the current study, this effective assessment model will be used and further evaluated in future large-scale studies.
References
Porter, P . ‘Westernizing’ women′s risks? Breast cancer in lower-income countries. N. Engl. J. Med. 358, 213–216 (2008).
Shen, Y. C., Chang, C. J., Hsu, C., Cheng, C. C., Chiu, C. F. & Cheng, A. L. Significant difference in the trends of female breast cancer incidence between Taiwanese and Caucasian Americans: implications from age-period-cohort analysis. Cancer Epidemiol. Biomarkers Prev. 14, 1986–1990 (2005).
Sim, X., Ali, R. A., Wedren, S., Goh, D. L., Tan, C. S., Reilly, M. et al. Ethnic differences in the time trend of female breast cancer incidence: Singapore, 1968-2002. BMC Cancer 6, 261 (2006).
Matsuno, R. K., Anderson, W. F., Yamamoto, S., Tsukuma, H., Pfeiffer, R. M., Kobayashi, K. et al. Early- and late-onset breast cancer types among women in the United States and Japan. Cancer Epidemiol. Biomarkers Prev. 16, 1437–1442 (2007).
Kuo, W. H., Yen, A. M., Lee, P. H., Hou, M. F., Chen, S. C., Chen, K. M. et al. Incidence and risk factors associated with bilateral breast cancer in area with early age diagnosis but low incidence of primary breast cancer: analysis of 10-year longitudinal cohort in Taiwan. Breast Cancer Res. Treat. 99, 221–228 (2006).
Kuo, W. H., Yen, A. M., Lee, P. H., Chen, K. M., Wang, J., Chang, K. J. et al. Cumulative survival in early-onset unilateral and bilateral breast cancer: an analysis of 1907 Taiwanese women. Br. J. Cancer 100, 563–570 (2009).
Malone, K. E., Begg, C. B., Haile, R. W., Borg, A., Concannon, P., Tellhed, L. et al. Population-based study of the risk of second primary contralateral breast cancer associated with carrying a mutation in BRCA1 or BRCA2. J. Clin. Oncol. 28, 2404–2410 (2010).
Lynch, H. T., Silva, E., Snyder, C. & Lynch, J. F. Hereditary breast cancer: part I. Diagnosing hereditary breast cancer syndromes. Breast J. 14, 3–13 (2008).
Chen, W., Pan, K., Ouyang, T., Li, J., Wang, T., Fan, Z. et al. BRCA1 germline mutations and tumor characteristics in Chinese women with familial or early-onset breast cancer. Breast Cancer Res. Treat. 117, 55–60 (2009).
Toh, G.T.K.P., Lee, S. S., Lee, D. S., Lee, S. Y., Selamat, S., Mohd, T. N. A. et al. BRCA1 and BRCA2 germline mutations in Malaysian women with early-onset breast cancer without a family history. PLoS ONE 3, e2024 (2008).
Ahn, S. H., Son, B. H., Yoon, K. S., Noh, D. Y., Han, W., Kim, S. W. et al. BRCA1 and BRCA2 germline mutations in Korean breast cancer patients at high risk of carrying mutations. Cancer Lett. 245, 90–95 (2007).
Liede, A. & Narod, S. A. Hereditary breast and ovarian cancer in Asia: genetic epidemiology of BRCA1 and BRCA2. Hum. Mutat. 20, 413–424 (2002).
Li, S. S., Tseng, H. M., Yang, T. P., Liu, C. H., Teng, S. J. & Huang, H. W. Molecular characterization of germline mutations in the BRCA1 and BRCA2 genes from breast cancer families in Taiwan. Hum. Genet. 104, 201–204 (1999).
Chen, S. T., Chen, R. A., Kuo, S. J. & Chien, Y. C. Mutational screening of breast cancer susceptibility gene 1 from early onset, bi-lateral, and familial breast cancer patients in Taiwan. Breast Cancer Res. Treat. 77, 133–143 (2003).
Venkitaraman, A. R. Linking the cellular functions of BRCA genes to cancer pathogenesis and treatment. Annu. Rev. Pathol. 4, 461–487 (2009).
Carvalho, M. A., Marsillac, S. M., Karchin, R., Manoukian, S., Grist, S. & Swaby, R. F. Determination of cancer risk associated with germ line BRCA1 missense variants by functional analysis. Cancer Res. 67, 1494–1501 (2007).
Dombernowsky, S. L., Weischer, M., Freiberg, J. J., Bojesen, S. E., Tybjaerg-Hansen, A. & Nordestgaard, B. G. Missense polymorphisms in BRCA1 and BRCA2 and risk of breast and ovarian cancer. Cancer Epidemiol. Biomarkers Prev. 18, 2339–2342 (2009).
Couch, F. J., Rasmussen, L. J., Hofstra, R., Monteiro, A. N., Greenblatt, M. S. & de Wind, N. Assessment of functional effects of unclassified genetic variants. Hum. Mutat. 29, 1314–1326 (2008).
Frolov, A., Prowse, A. H., Vanderveer, L., Bove, B., Wu, H. & Godwin, A. K. DNA array-based method for detection of large rearrangements in the BRCA1 gene. Genes Chromosomes Cancer 35, 232–241 (2002).
Fleming, M. A., Potter, J. D., Ramirez, C. J., Ostrander, G. K. & Ostrander, E. A. Understanding missense mutations in the BRCA1 gene: an evolutionary approach. Proc. Natl. Acad. Sci. USA 100, 1151–1156 (2003).
Fiser, A. & Sali, A. Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol. 374, 461–491 (2003).
Thomas, P. D., Campbell, M. J., Kejariwal, A., Mi, H., Karlak, B. & Daverman, R. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Adzhubei, I. A., Schmidt, S., Peshkin, L., Ramensky, V. E., Gerasimova, A. & Bork, P. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Kaminker, J. S., Zhang, Y., Waugh, A., Haverty, P. M., Peters, B. & Sebisanovic, D. Distinguishing cancer-associated missense mutations from common polymorphisms. Cancer Res. 67, 465–473 (2007).
Evans, D. G., Lalloo, F., Cramer, A., Jones, E. A., Knox, F., Amir, E. et al. Addition of pathology and biomarker information significantly improves the performance of the Manchester scoring system for BRCA1 and BRCA2 testing. J. Med. Genet. 46, 811–817 (2009).
Gough, C. A., Gojobori, T. & Imanishi, T. Cancer-related mutations in BRCA1-BRCT cause long-range structural changes in protein-protein binding sites: a molecular dynamics study. Proteins 66, 69–86 (2007).
Williams, R. S., Lee, M. S., Hau, D. D. & Glover, J. N. Structural basis of phosphopeptide recognition by the BRCT domain of BRCA1. Nat. Struct. Mol. Biol. 11, 519–525 (2004).
Hooft, R. W., Vriend, G., Sander, C. & Abola, E. E. Errors in protein structures. Nature 381, 272 (1996).
Humphrey, J. S., Salim, A., Erdos, M. R., Collins, F. S., Brody, L. C. & Klausner, R. D. Human BRCA1 inhibits growth in yeast: potential use in diagnostic testing. Proc. Natl. Acad. Sci. USA 94, 5820–5825 (1997).
Górski, B., Jakubowska, A., Huzarski, T., Byrski, T., Gronwald, J., Grzybowska, E. et al. A high proportion of founder BRCA1 mutations in Polish breast cancer families. Int. J. Cancer 110, 683–686 (2004).
De Leon Matsuda, M. L., Liede, A., Kwan, E., Mapua, C. A., Cutiongco, E. M., Tan, A. et al. BRCA1 and BRCA2 mutations among breast cancer patients from the Philippines. Int. J. Cancer 98, 596–603 (2002).
Lee, A. S., Ho, G. H., Oh, P. C., Balram, C., Ooi, L. L., Lim, D. T. et al. Founder mutation in the BRCA1 Gene in Malay Breast Cancer Patients From Singapore. Hum. Mutat. Mutation in Brief #633 22, 178 (2003).
Carvalho, M. A., Couch, F. J. & Monteiro, A. N. Functional assays for BRCA1 and BRCA2. Int. J. Biochem. Cell Biol. 39, 298–310 (2007).
Tischkowitz, M., Hamel, N., Carvalho, M. A., Birrane, G., Soni, A., van Beers, E. H. et al. Pathogenicity of the BRCA1 missense variant M1775K is determined by the disruption of the BRCT phosphopeptide-binding pocket: a multi-modal approach. Eur. J. Hum. Genet. 16, 820–832 (2008).
Carvalho, M., Pino, M. A., Karchin, R., Beddor, J., Godinho-Netto, M., Mesquita, R. D et al. Analysis of a set of missense, frameshift, and in-frame deletion variants of BRCA1. Mutat. Res. 660, 1–11 (2009).
Hashizume, R., Fukuda, M., Maeda, I., Nishidawa, H., Oyake, D., Yabuki, Y. et al. The RING heterodimer BRCA1-BARD1 is a ubiquitin ligase inactivated by a breast cancer-derived mutation. J. Biol. Chem. 276, 14537–14540 (2001).
Fong, P. C., Boss, D. S., Yap, T. A., Tutt, A., Wu, P., Mergui-Roelvink, M. et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N. Engl. J. Med. 361, 123–134 (2009).
Fong, P. C., Yap, T. A., Boss, D. S., Carden, C. P., Mergui-Roelvink, M., Gourley, C. et al. Poly(ADP)-ribose polymerase inhibition: frequent durable responses in BRCA carrier ovarian cancer correlating with platinum-free interval. J. Clin. Oncol. 28, 2512–2519 (2010).
Acknowledgements
We thank Dr Huamei Chang and Vita Genomics, Inc. for help with technological aspects of the BRCAChip microarray sequencing; Dr Fang-Jen Lee and Gootle Chen for technological assistance with the yeast assay; Dr Ming-Shi Shiao and Dr Chi-Yuan Chou for help with structural modeling and Dr Long-Yuan Li for help in homologous recombination assay. This work was supported in part by the National Science Council (grant NSC 95-2314-B-002-154-MY3), and Taiwan Department of Health Clinical Trial and Research Center of Excellence (DOH100-TD-B-111-004), Taiwan, Republic of China.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Kuo, WH., Lin, PH., Huang, AC. et al. Multimodel assessment of BRCA1 mutations in Taiwanese (ethnic Chinese) women with early-onset, bilateral or familial breast cancer. J Hum Genet 57, 130–138 (2012). https://doi.org/10.1038/jhg.2011.142
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.142
Keywords
This article is cited by
-
Prevalence of BRCA1, BRCA2, and PALB2 genomic alterations among 924 Taiwanese breast cancer assays with tumor-only targeted sequencing: extended data analysis from the VGH-TAYLOR study
Breast Cancer Research (2023)
-
Impact of BRCA mutation on the survival and risk of contralateral breast cancer in Asian breast cancer patients
Breast Cancer Research and Treatment (2022)
-
Germline breast cancer susceptibility gene mutations and breast cancer outcomes
BMC Cancer (2018)
-
iRegNet3D: three-dimensional integrated regulatory network for the genomic analysis of coding and non-coding disease mutations
Genome Biology (2017)
-
High prevalence and predominance of BRCA1 germline mutations in Pakistani triple-negative breast cancer patients
BMC Cancer (2016)