Abstract
MLXIPL is a transcription factor integral to the regulation of glycolysis and lipogenesis in the liver. Common variants of the MLXIPL gene (MLXIPL) are known to influence plasma triglyceride levels in people of European descent. As MLXIPL has a key role in energy storage, genetic variations of the MLXIPL may be relevant to physiological adaptations to nutritional stresses that have occurred during the evolution of modern humans. In the present study, we assessed the phenotypic consequences of the Q241H variant of MLXIPL in populations of Asian and Oceanian origin and also surveyed the prevalence of Q241H variant in populations worldwide. Multiple linear regression models based on 2373 individuals of Asian origin showed that the H allele was significantly associated with decreased concentrations of plasma triglycerides (P=0.0003). Direct genotyping of 1455 individuals from Africa, Asia and Oceania showed that the triglyceride-lowering H allele was found at quite low frequencies (0.00–0.16) in most of the populations examined. The exceptions were some Central Asian populations, including Mongolians, Tibetans and Uyghurs, which exhibited much higher frequencies of the H allele (0.21–0.26). The high prevalence of the H allele in Central Asia implies that the Q241H variant of MLXIPL might have been significant for utilization of carbohydrates and fats in the common ancestors of these populations, who successfully adapted to the environment of Central Asia by relying on nomadic livestock herding.
Similar content being viewed by others
Introduction
Triglycerides now receive much attention for their importance in the etiology of cardiovascular disease, one of the leading causes of death in industrialized countries. Hypertriglyceridemia (a condition in which plasma triglyceride levels are elevated) can increase the risk of developing atherosclerosis. However, triglycerides are the principal energy source that is suitable for long-term storage. Therefore, triglycerides have been essential molecules for the evolution of modern humans. Abnormalities associated with triglycerides result in part from the mismatch between modern lifestyles and an innate human physiology that evolved to efficiently store excessive calories as triglycerides.
Many genetic studies of lipid abnormalities and cardiovascular disease have discovered loci associated with plasma triglyceride concentrations, high-density lipoprotein (HDL) cholesterol, and low-density lipoprotein (LDL) cholesterol1 Familial severe hyperlipidemia is caused by rare, deleterious mutations in genes that have central roles in lipoprotein metabolism, including those that encode lipoprotein lipase, LDL receptor, apolipoprotein B and cholesteryl ester transfer protein. Common genetic variants with more moderate effects have also been identified. These variants, such as the ɛ4 allele of the apolipoprotein E gene, contribute significantly to the variation of plasma lipid traits and the risk for polygenic hyperlipidemia in the general population.1
Recent genome-wide association studies identified at least 30 loci that were associated with plasma lipid traits in populations of European descent.2, 3, 4, 5, 6 One of these newly identified genes, MLXIPL (MLX-interacting protein-like, located in 7q11), is involved in the regulation of glycolysis and lipogenesis in humans and rodents.7 MLXIPL, also known as the carbohydrate-responsive element-binding protein, is a basic helix-loop-helix leucine zipper transcription factor expressed predominantly in the liver.8 The influx of glucose into hepatocytes and the resulting generation of xylulose-5-phosphates trigger nuclear localization and activation of MLXIPL.9 Heterotetramers of activated MLXIPL and MLX (Max-like protein X) upregulate the expression of the gene encoding liver-type pyruvate kinase, the enzyme that regulates the final step of glycolysis.10, 11, 12 MLXIPL also regulates the expression of genes involved in the NADPH supply system, gluconeogenesis and lipogenesis and consequently favors the conversion of excess carbohydrates into triglycerides rather than glycogen11, 13 MLXIPL-knockout mice were viable and showed reduced hepatic lipogenesis from glucose and increased liver glycogen content.11
In the genome-wide association study conducted by Kooner et al., a common missense variant in the exon 6 of MLXIPL (rs3812316), resulting in a change from glutamine (Q) to histidine (H) at residue 241 of MLXIPL, was strongly associated with low plasma triglyceride concentrations.3 We successfully replicated the association between the Q241H variant and low plasma triglyceride concentrations in a population of over 20 000 Japanese individuals,14 and Pan et al. reported a significant association between the Q241H variant and risks for coronary artery disease in a Chinese population.15 In contrast, Vrabilk et al. found no evidence for an association between Q241H variant and low plasma triglyceride concentrations in a Caucasian population.16 Repeating these studies in a variety of human populations would thus help to substantiate the association between the Q241H variant and plasma triglyceride concentrations. Additionally, Kooner et al. postulated that because the major Q allele of MLXIPL, which was associated with higher triglyceride concentrations, seems to permit more efficient conversion of carbohydrates into fats when food is abundant, MLXIPL is a plausible candidate for a ‘thrifty gene’ in the evolution of modern humans.3 Information on the worldwide prevalence of the Q241H variant may provide insights into the evolutionary role of MLXIPL; however, very little is known about the frequencies of the Q241H variant in the human populations worldwide. In the present study, we assessed the association between the Q241H variant and plasma triglyceride concentrations in Asian and Oceanian populations with different lifestyles and also surveyed the distribution of the Q241H variant in populations worldwide.
Materials and methods
Association study
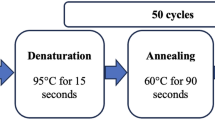
Blood specimens were collected from 2492 unrelated individuals, including 570 Japanese, 1203 Mongolian, 388 Chinese, 212 Thai and 119 Palauans. Written informed consent was obtained from the all participants. Plasma triglyceride, total cholesterol and high-density lipoprotein (HDL) cholesterol concentrations were determined using standard methods. LDL cholesterol concentrations were estimated according to the Friedewald's formula. For individuals exhibiting >400 mg dl−1 triglycerides, a null value was assigned for the LDL cholesterol concentration. Genotyping of the Q241H variant was performed using TaqMan genotyping assay systems and an ABI PRISM 7900HT Sequence Detection System (Applied Biosystems). The association between the Q241H variant and plasma lipid concentrations was tested using multiple linear regression models assuming an additive model of inheritance. Genotypes were coded as 0, 1 or 2 depending on the copy numbers of the H allele. Triglyceride concentration values were transformed using a -1/sqrt of log transformation to remove skew.3, 14 Individuals under treatment for diabetes mellitus or with HbA1c ⩾6.5% were excluded. Sex (male=0; female=1), age and body mass index were included as independent variables. We also tested the association between the Q241H variant and plasma triglyceride concentrations in combined samples by means of a linear multiple regression model including genotypes, sex, age, body mass index and ethnicity.
Estimation of the worldwide distribution of Q241H variant
We determined Q241H variant genotypes of 1455 unrelated individuals from Africa (Ghanaians, Xhosans and South Africans of European descent), South Asia (Tamils, Sinhalese, Nepalis and Bangladeshis), East Asia (two Han Chinese populations and Japanese in Okinawa), South-East Asia (Indonesians), Central Asia (Uyghurs and Tibetans) and Oceania (Samoans). Genotyping was performed as described above. We also retrieved H allele frequencies of East Asian populations from the HGDP-CEPH Human Genome Diversity Cell Line Panel.17 Genotype data including 650 000 single-nucleotide polymorphisms (SNPs) from the HGDP-CEPH individuals were downloaded from the HGDP-CEPH database.18 As the Q241H variant was not found in the SNP array in the previous study,18 rs17147538, which is separated from the Q241H variant by 37.4 kb and in absolute linkage disequilibrium with the Q241H variant in Mongolians and Japanese (r2=1.0), was used to infer Q241H variant genotypes of the HDGP-CEPH individuals.
Determination of the ancestral allele of the Q241H variant
To determine the human ancestral allele of the Q241H variant, the MLXIPL coding sequences from eight chimpanzees (Pan troglodytes) were determined using PCR amplification and direct sequencing. Exon 6 and flanking intronic sequences of MLXIPL were amplified with FastStart Taq Polymerase (Roche Applied Science, Tokyo, Japan). PCR products were purified using Agencourt AMPure (Beckman Coulter, Tokyo, Japan). The dye termination reaction was performed using the DYEnamic ET dye terminator kit (GE Healthcare, Tokyo, Japan). DNA sequences were determined using a MegaBACE 1000 (GE Healthcare). We also queried two world-wide web-based services, Polyphen (http://genetics.bwh.harvard.edu/pph/) and SIFT (http://sift.jcvi.org/), for the functional significance of the Q to H substitution at residue 241 of MLXIPL.
Analysis of polymorphism patterns surrounding the Q241H variant
The genotypes of 46 SNPs near the Q241H variant were determined in 47 randomly selected Mongolian individuals by using the high resolution melting analysis.19 A list of the genotyped SNPs and PCR primer sequences may be available by request. PCR was performed using ExTaq polymerase version HS (Takara, Tokyo, Japan) and LCGreen+ (Idaho Technology, Salt Lake City, UT, USA). High resolution melting analysis of the amplified products was performed using a LightScanner instrument (Idaho Technology). The linkage disequilibrium (LD) status of the Q241H variant with the 46 SNPs was analyzed using Haploview software (http://www.broadinstitute.org/).20 We also genotyped 95 Japanese, 121 Ghanaians and 100 South Africans of European descent for rs17145738 and determined its LD status with the Q241H variant in each population.
Results
Association between the Q241H variant and plasma triglyceride concentrations
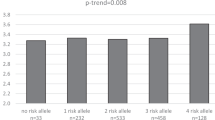
The frequencies of the H allele in the five Asia-Pacific populations were estimated as follows: 0.23 in the Mongolian, 0.10 in the Japanese, 0.09 in the Thai, 0.13 in the Chinese and 0.03 in the Palauan population (Table 1). Deviation from the Hardy–Weinberg equilibrium was not observed in any of the populations (P>0.05). Individuals homozygous for the H allele were not detected in the Palauan population. We therefore excluded the Palauan population from further association analysis. The results of the multiple linear regression analyses are shown in Table 2. In the combined samples analysis, body mass index and sex showed strong associations with plasma triglyceride concentrations, and the H allele was strongly associated with lower plasma triglyceride concentrations (β=−0.08, P=0.0003). Linear multiple regression models based on the individual populations showed that the association between the H allele and the plasma triglyceride concentration was significant in the Mongolian, Japanese and Chinese populations (P<0.05). The H allele also showed a triglyceride-lowering effect in the Thai population, although the association was not significant. In the combined samples, the H allele was not associated with body mass index or with plasma concentrations of total cholesterol, HDL cholesterol or LDL cholesterol (data not shown).
Worldwide distribution of the H allele
We determined genotypes of Q241H variant of 1455 individuals from Africa, South Asia, East Asia, Central Asia, South-East Asia and Oceania in order to know the prevalence of the H allele in populations worldwide. We also estimated the frequencies of the H allele in the HGDP-CEPH Asian populations by inference from their genotypes of rs17145738, in which we confirmed absolute linkage disequilibrium with Q241H variant in Japanese and Mongolian populations. The distribution of the H allele in the tested populations is illustrated in Figure 1. Among the directly genotyped populations, the H allele was found at low frequencies in Africa (0.05), South Asia (0.06), East Asia (0.11) and South-East Asia (0.12). Central Asian populations including Mongolians, Tibetans and Uyghurs showed a higher prevalence of the H allele (0.21–0.26). Among the HGDP-CHEP Asian populations, the Yakut (n=25) showed a high prevalence of the H allele (0.30). Although the sample sizes of the other HGDP-CHEP populations were quite small (n⩽11), estimated H allele frequencies ranged from 0.00 (Lahu, Dai and Miao) to 0.22 (Hezhen), 0.33(Xibo), 0.35 (Yi) and 0.50 (Naxi).
Distribution of the Q241H variant in the Old World (a) and in Asia (b). Allele frequencies of the H allele in each population are indicated in black areas (directly genotyped populations) or gray areas (HGDP-CEPH populations) of the pie charts. GH: Ghanaians; XA: Xhosans; AE: South Africans of European descent; UG: Uyghurs; TB: Tibetans; MG: Mongolians; NP: Nepalis; SH: Sinhalese; TM: Tamils; BA: Bangladeshis; TH: Thai; HN: Han Chinese (northern); HS: Han Chinese (southern); KR: Koreans; JK: Japanese (Kanto); JO: Japanese (Okinawa); PL: Palauans; SA: Samoans; YK: Yakuts; OR: Oroqens; DR: Daurs; HZ: Hezhens; XB: Xibo; MO: Mongola; TU: Tu; NX: Naxi; YI: Yi; MI: Miao; TJ: Tujia; LH: Lahu; DA: Dai; and SE: She. The positions of SH, TM and YK in the map B do not reflect the geographic locations of the sampling sites. The H allele frequencies of the directly genotyped populations are listed in Supplementary Table S1.
Polymorphism patterns of MLXIPL and surrounding regions
Our sequencing analysis revealed that all eight chimpanzees tested harbored the Q residue at amino acid position 241 of MLXIPL. Additionally, the published genome sequence of a rhesus macaque also encodes Q at this position. Therefore, the major Q allele was assigned as the human ancestral allele of Q241H variant. The functional predictions for the Q-to-H substitution at position 241 were ‘possibly damaging’ and ‘tolerated’ according to Polyphen and SIFT, respectively. Unfortunately the LD status of the Q241H variant and surrounding SNPs in humans is not well characterized as the Q241H variant was neither genotyped in the HapMap populations nor implemented in any commercially available genome-wide association studies platform. We therefore analyzed the LD status of the Q241H variant with a total of 46 nearby SNPs in 47 randomly selected Mongolian individuals. None of the additional 46 SNPs genotyped deviated from the Hardy–Weinberg equilibrium in these individuals. The Q241H variant was located in a tight 191-kb LD block that involved the entire coding region of MLXIPL as well as four nearby genes (Figure 2). The rs17145738, which has been shown to be associated with triglycerides in populations of European descent,2, 4, 5, 6 was also located in the 191 kb LD block and was in absolute LD (r2=1.0) with the Q241H variant in the Mongolians tested. We further determined the rs17145738 genotypes of 95 Japanese, 121 Ghanaians and 100 South Africans of European descent. rs17145738 and the Q241H variant were in tight LD in the Japanese (r2=1.0) and the South Africans of European descent (r2=0.91), but no tight LD status was observed in the Ghanaians (r2=0.15).
Linkage disequilibrium status of the Q241H variant and nearby SNPs. LD status of the Q241H variant with 46 SNPs is illustrated by the magnitude of the r2 value. The location of the Q241H variant is highlighted. The genomic positions of the SNPs and nearby genes (black arrows) are based on the NCBI Build36 assembly (nucleotide position 72 360 917–72 759 283 of human chromosome 7). The range of the 191 kb LD in the genome is indicated by the thick gray line.
Discussion
The strong association between the Q241H variant of MLXIPL and low plasma triglyceride concentrations was first reported by Kooner et al.,3 and we successfully replicated this association in our previous study.14 In the present study, we found additional evidence for the association between the Q241H variant and low plasma triglyceride concentrations in 2373 Asian individuals of multiple ethnic origins. The association was also supported by regression models based on subsets of the individual populations including the Mongolian, Japanese and Chinese populations. We believe that the absence of significant associations in the models based on the Thai population was due only to the small sample sizes. Furthermore, our screening for the Q241H variant revealed that the triglyceride-lowering H allele was found at high frequencies in various ethnic groups in Central and East Asia. The Q241H variant of MLXIPL is considered to be an important protective factor against polygenic hypertriglyceridemia in Central and East Asians, as well as in populations of European descent.21, 22
It remains unclear whether the observed associations were due to a deleterious effect of the Q241H variant on MLXIPL function. The Q residue at position 241 is located in an evolutionarily conserved motif of MLXIPL, namely the Glucose-Sensing Module.7 The Polyphen algorithm predicted that the Q-to-H substitution might impair MLXIPL function, but the SIFT algorithm did not support any such effect. The functional consequences of the Q-to-H substitution should be confirmed experimentally in order to address this question. It is also possible that the observed associations simply reflected linkage disequilibrium with another variant. In the genome-wide association studiess conducted by Kathiresan et al. and Willer et al., the Q241H variant was not genotyped and rs17145738 was identified as the marker with the strongest evidence for association with low plasma triglyceride concentrations.2, 4 Our LD analyses showed that the Q241H variant and rs17145738 were in tight LD in East Asians and in South Africans of European descent (r2⩾0.90) and fell into a 191-kb LD block that included MLXIPL and four nearby genes. On the other hand, H241Q variant and rs17145738 did not show strong LD in Ghanaians (r2=0.15). We therefore propose further association studies on individuals of African origins to help substantiate the association between the Q241H variant and low plasma triglyceride concentrations.
It is intriguing that the ‘anti-hyperlipidemic’ H allele was found at higher frequencies in Central Asians. Mongolians and Tibetans apparently originated from nomadic tribes inhabiting northern China,23 and genetic studies have consistently supported a close ancestral relationship between Tibetans and Mongolians.24, 25, 26 The Uyghur are Turk-speaking people who live predominantly in the Xinjiang Uyghur Autonomous Region of China and can be traced back to the nomadic tribes of eastern Turkistan.27 Although modern Uyghur populations contain a substantial admixture of European genes,28 the close genetic relationship between the Uyghur and Mongolians was supported by a mitochondrial DNA study.29 We inferred the Q241H variant genotypes of the HGDP-CEPH populations from their rs17145738 genotypes and found that the H allele was also prevalent in Yakuts, who are Turk-speaking people living predominantly in Siberia. Additionally, several Chinese minority groups in HGDP-CEPH, including the Hezhen, Xibo, Naxi and Yi, showed a high prevalence of the H allele, although sample sizes of these populations were quite small. Linguistic and historical evidence suggests that these HDGP-CEPH groups may share ancestry with the three central Asian populations.27 The considerable genetic and historical evidence for relationships among these populations suggests that the frequency of the H allele was increased in their common ancestors in Central Asia, who probably lived a nomadic lifestyle dependent on livestock herding.
The worldwide distribution of the Q241H variant may reflect the history of the dietary adaptations of modern humans. MLXIPL function, which ultimately converts excess glucose to triglycerides, would be important for coping with severe fluctuations in the availability of food. The brain and erythrocytes depend largely on glycolysis rather than on the β-oxidation of fatty acids for the generation of energy. In a situation in which carbohydrates were scarce but fats were readily available, impaired conversion of glucose to triglycerides by the liver might promote survival by reserving glucose for these more dependent cell types. The traditional foods and dietary behaviors of Central Asians, such as Mongolians, strongly suggest that fat-rich livestock products, including meats, milk and butter, were important dietary energy sources for their nomadic ancestors. We believe that the rise of the H allele in Central Asians may be a genetic component of adaptation to a nomadic livestock herding system. We are now performing resequencing and population genetic analyses of MLXIPL in Mongolians and other Central Asian populations to attempt to detect signatures of such selection for the Q241H variant.
References
Hegele, R. A. Plasma lipoproteins: genetic influences and clinical implications. Nat. Rev. Genet. 10, 109–121 (2009).
Kathiresan, S., Melander, O., Guiducci, C., Surti, A., Burtt, N. P. et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nat. Genet. 40, 189–197 (2008).
Kooner, J. S., Chambers, J. C., Aguilar-Salinas, C. A., Hinds, D. A., Hyde, C. L., Warnes, G. R. et al. Genome-wide scan identifies variation in MLXIPL associated with plasma triglycerides. Nat. Genet. 40, 149–151 (2008).
Willer, C. J., Sanna, S., Jackson, A. U., Scuteri, A., Bonnycastle, L. L., Clarke, R. et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat. Genet. 40, 161–169 (2008).
Kathiresan, S., Willer, C. J., Peloso, G. M., Demissie, S., Musunuru, K., Schadt, E. E. et al. Common variants at 30 loci contribute to polygenic dyslipidemia. Nat. Genet. 41, 56–65 (2009).
Aulchenko, Y. S., Ripatti, S., Lindqvist, I., Boomsma, D., Heid, I. M., Pramstaller, P. P. et al. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nat. Genet. 41, 47–55 (2009).
Uyeda, K. & Repa, J. J. Carbohydrate response element binding protein, ChREBP, a transcription factor coupling hepatic glucose utilization and lipid synthesis. Cell Metab. 4, 107–110 (2006).
Yamashita, H., Takenoshita, M., Sakurai, M., Bruick, R. K., Henzel, W. J. & Shillinglaw, W. A glucose-responsive transcription factor that regulates carbohydrate metabolism in the liver. Proc. Natl Acad. Sci. U. S. A. 98, 9116–9121 (2001).
Kabashima, T., Kawaguchi, T., Wadzinski, B. E. & Uyeda, K. Xylulose 5-phosphate mediates glucose-induced lipogenesis by xylulose 5-phosphate-activated protein phosphatase in rat liver. Proc. Natl Acad. Sci. U. S. A. 100, 5107–5112 (2003).
Cairo, S., Merla, G., Urbinati, F., Ballabio, A. & Reymond, A. WBSCR14, a gene mapping to the Williams-Beuren syndrome deleted region, is a new member of the Mlx transcription factor network. Hum. Mol. Genet. 10, 617–627 (2001).
Iizuka, K., Bruick, R. K., Liang, G., Horton, J. D. & Uyeda, K. Deficiency of carbohydrate response element-binding protein (ChREBP) reduces lipogenesis as well as glycolysis. Proc. Natl Acad. Sci. U. S. A. 101, 7281–7286 (2004).
Stoeckman, A. K., Ma, L. & Towle, H. C. Mlx is the functional heteromeric partner of the carbohydrate response element-binding protein in glucose regulation of lipogenic enzyme genes. J. Biol. Chem. 279, 15662–15669 (2004).
Ma, L., Tsatsos, N. G. & Towle,, H. C. Direct role of ChREBP.Mlx in regulating hepatic glucose-responsive genes. J. Biol. Chem. 280, 12019–12027 (2005).
Nakayama, K., Bayasgalan, T., Yamanaka, K., Kumada, M., Gotoh, T., Utsumi, N. et al. Large scale replication analysis of loci associated with lipid concentrations in a Japanese population. J. Med. Genet. 46, 370–374 (2009).
Pan, L. A., Chen, Y. C., Huang, H., Zhang, L., Liu, R., Li, X. et al. G771C polymorphism in the MLXIPL gene is associated with a risk of coronary artery disease in the Chinese: a case-control study. Cardiology 114, 174–178 (2009).
Vrablik, M., Ceska, R., Adamkova, V., Peasey, A., Pikhart, H., Kubinova, R. et al. MLXIPL variant in individuals with low and high triglyceridemia in white population in Central Europe. Hum. Genet. 124, 553–555 (2008).
Cann, H. M., de Toma, C., Cazes, L., Legrand, M. F., Morel, V., Piouffre, L. et al. A human genome diversity cell line panel. Science 296, 261–262 (2002).
Li, J. Z., Absher, D. M., Tang, H., Southwick, A. M., Casto, A. M., Ramachandran, S. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
Liew, M., Pryor, R., Palais, R., Meadows, C., Erali, M., Lyon, E. et al. Genotyping of single-nucleotide polymorphisms by high-resolution melting of small amplicons. Clin. Chem. 50, 1156–1164 (2004).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Wang, J., Ban, M. R., Zou, G. Y., Cao, H., Lin, T., Kennedy, B. A. et al. Polygenic determinants of severe hypertriglyceridemia. Hum. Mol. Genet. 17, 2894–2899 (2008).
Hegele, R. A., Ban, M. R., Hsueh, N., Kennedy, B. A., Cao, H., Zou, G. Y. et al. A polygenic basis for four classical Fredrickson hyperlipoproteinemia phenotypes that are characterized by hypertriglyceridemia. Hum. Mol. Genet. 18, 4189–4194 (2009).
McNally, R. The Atlas of Mankind, Mitchell Beazley, London, 1982.
Nei, M. & Roychoudhury, A. K. Evolutionary relationships of human populations on a global scale. Mol. Biol. Evol. 10, 927–943 (1993).
Qian, Y., Qian, B., Su, B., Yu, J., Ke, Y., Chu, Z. et al. Multiple origins of Tibetan Y chromosomes. Hum. Genet. 106, 453–454 (2000).
Su, B., Xiao, C., Deka, R., Seielstad, M. T., Kangwanpong, D., Xiao, J. et al. Y chromosome haplotypes reveal prehistorical migrations to the Himalayas. Hum. Genet. 107, 582–590 (2000).
Olson,, J. S. An Ethnohistorical Dictionary of China, Greenwood Press, Westport, 1998.
Xu, S. & Jin, L. A genome-wide analysis of admixture in Uyghurs and a high-density admixture map for disease-gene discovery. Am. J. Hum. Genet. 83, 322–336 (2008).
Comas, D., Calafell, F., Mateu, E., Perez-Lezaun, A., Bosch, E., Martinez-Arias, R. et al. Trading genes along the silk road: mtDNA sequences and the origin of central Asian populations. Am. J. Hum. Genet. 63, 1824–1838 (1998).
Acknowledgements
This study was supported in part by the Ministry of Education, Culture, Sports, Science and Technology, grant-in-aid for young scientists (A) 21687021, JKA through its promotional funds from KEIRIN RACE.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Nakayama, K., Yanagisawa, Y., Ogawa, A. et al. High prevalence of an anti-hypertriglyceridemic variant of the MLXIPL gene in Central Asia. J Hum Genet 56, 828–833 (2011). https://doi.org/10.1038/jhg.2011.109
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.109
Keywords
This article is cited by
-
Genome-wide DNA methylation status of Mongolians exhibits signs of cellular stress response related to their nomadic lifestyle
Journal of Physiological Anthropology (2022)
-
Relationship of lipid regulatory gene polymorphisms and dyslipidemia in a pediatric population: the CASPIAN III study
Hormones (2018)
-
Replication analysis of genetic association of the NCAN-CILP2 region with plasma lipid levels and non-alcoholic fatty liver disease in Asian and Pacific ethnic groups
Lipids in Health and Disease (2016)
-
Association between the MLX Interacting Protein-Like, BUD13 Homolog and Zinc Finger Protein 259 Gene Polymorphisms and Serum Lipid Levels
Scientific Reports (2014)