Abstract
We investigated mutations of the iduronate-2-sulfatase (I2S) gene and structural characteristics of I2S to clarify genotype/phenotype relationships in 18 Japanese patients with mucopolysaccharidosis type II. The I2S gene was analyzed in five patients with a severe phenotype and in 13 patients with an attenuated phenotype. The tertiary structural model of the human I2S was constructed by homology modeling using the arylsulfatase structure as a template. We identified four missense mutations and a nonsense mutation in the severe phenotype; four missense, two nonsense, three frame shifts, and one each of splice and amino acid deletion in the attenuated phenotype. Seven of them (L73del, Q75X, G140R, C171R, V401 fs, C422 fs, and H441 fs) were novel mutations. Structural analysis indicated that the residues of the mutations found in the severe phenotype would have direct interactions with the active site residues or should break the hydrophobic core domain of I2S, whereas residues of the missense mutations found in the attenuated phenotype were located in the peripheral region. In addition, effects by deletion or frameshift mutations could also be interpreted by the structure. Structural analysis of mutant proteins would help in understanding the genotype/phenotype relationships of Hunter disease.
Similar content being viewed by others
Introduction
Mucopolysaccharidosis (MPS) type II (Hunter disease) is the most prevalent subtype among Japanese MPS patients; about half of patients with MPS in Japan have Hunter disease (Yamada et al. 1993). Hunter disease is characterized by systemic manifestations including short stature, joint contracture, dysostosis multiplex, typical facial appearance, thick skin and hirsutism, conductive and sensorineural hearing difficulty, ophthalmic problems such as retinitis pigmentosa, respiratory problems due to obstructive airways, congestive heart failure due to valvular diseases and cardiomyopathy, umbilical hernia, and hepatosplenomegaly (Neufeld and Muenzer 2001). Mutations in the iduronate-2-sulfatase (I2S) gene residing in Xq28 result in the primary cause of the disease, and the enzyme defect leads to an accumulation of dermatan sulfate and heparan sulfate in the body and increased excretion of these substances into urine (Neufeld and Muenzer 2001).
Patients with a severe clinical phenotype manifest clinical symptoms from infancy and develop central nervous system disorders. These severe patients usually do not survive their second decade of life, whereas those with an attenuated form gradually develop clinical manifestations later in childhood, show normal intelligence, and can survive up to middle age, although the attenuated form includes patients with a wide range of severity. Residual activity of I2S or the amount of excretion of glycosaminoglycans cannot predict the severity of the disease.
Gene mutations in Hunter disease were first reported in 1990 (Wilson et al. 1990). Since then, a broad spectrum of mutations has been reported (Sukegawa et al. 1992; Kim et al. 2003). However, the molecular mechanisms of genotype/phenotypes have not been elucidated. Here, we have clarified I2S gene mutations in 18 Japanese patients with Hunter disease, seven of which were novel mutations. We also discuss genotype/phenotype relationships in Hunter disease from results of the structural analysis of the I2S gene and the effects of several mutations on the three-dimensional (3D) structure of I2S.
Materials and methods
Subjects and enzyme activity
Clinical findings and enzyme activity of I2S are shown in Table 1. The I2S activity was measured by the procedure of Voznyi et al. (2001) with minor modification. Briefly, homogenates were prepared by sonication of cell material in water, and the concentration was adjusted by dilution with pH- and heat-denatured bovine serum albumin (BSA) solution (0.2%). Ten microliters of the homogenate (10 μg protein for fibroblasts or 15 μg for leukocytes) was mixed with 20 μl 1.25 mmol/l 4-methylumbelliferyl-[alpha]-iduronate 2-sulphate (MU-[alpha]Ido-2S, Moscerdam Substrates, Rotterdam, The Netherlands) in 0.1 mol/l sodium acetate buffer, pH 5.0, containing 10 mmol/l lead acetate. The reaction mixtures were incubated for 4 h 37°C, whereafter 40 μl concentrated McIlvain’s buffer, pH 4.5 (0.4 mol/l Na-phosphate/0.2 mol/l citrate), and 10 μl of LEBT solution (purified lysosomal enzymes from bovine testis) were added, and a second incubation of 24 h at 37°C was carried out. Reactions were terminated by the addition of 200 μl 0.5 mol/l Na2CO3/NaHCO3, pH 10.7, and 0.025% Triton X-100. The fluorescence of 4-methylumbelliferone was then measured.
DNA preparation and sequencing
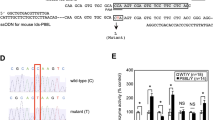
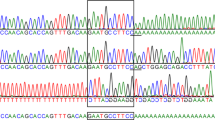
Using the GFX Genomic Blood DNA Purification Kit (Amersham Biosciences, Uppsala, Sweden), genomic DNA from peripheral leukocytes was extracted from 18 Japanese patients with Hunter disease (five patients with the severe phenotype and 13 patients with the attenuated phenotype) after obtaining their informed consent. Ten fragments of the I2S gene, exons and exon/intron boundaries, were amplified by polymerase chain reaction (PCR) using primers listed in Table 2 (Bunge et al. 1993; Rathmann et al. 1996; Isogai et al. 1998; Gort et al. 1998). The PCR reaction was carried out in a total volume of 50 μl containing 1.25 units of ExTaq polymerase, 10 nmol dNTPs, genomic DNA (100 ng), and the respective primers (0.5–2.0 μM). The PCR conditions were 10 cycles of 94°C for 30 s, 65–55°C for 30 s (with 1°C step-down by one cycle), and 72°C for 1 min, followed by 25 cycles of 94°C for 30 s, 55°C for 30 s, and 72°C for 1 min. Amplified products were precipitated, electrophoresed, extracted by RECOCHIP (Takara Bio, Otsu, Japan), and directly sequenced using the ABI PRISM 3100-Avant Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). Human Genomic DNA (Clontech, Palo Alto, CA, USA) was used as control.
Structural analysis
To investigate the effects of mutations determined in this study on clinical phenotypes, the tertiary structure of the human I2S was constructed using the arylsulfatase structure as a template by homology modeling (Shi et al. 2001), and effects of mutations on the tertiary structural model of the human I2S protein were assessed.
Results
Enzyme activity
The I2S enzyme activities of patients were all under detection limits, and there was no relationship between severity of the disease and residual activity of I2S (Table 1).
Genetic analysis of I2S gene
We performed mutational analysis by PCR but did not perform Southern blot analysis in this study. However, the results indicated that the causative mutations should be substitutions, small deletions, or small insertions, but not large genomic rearrangements.
Mutations found in the severe phenotype
Results of mutation analyses are summarized in Table 3. Four missense mutations (G224E, G336 V, R468 W, and R468L) and a nonsense (Q75X) mutation were identified in patients with the severe phenotype. The Q75X nonsense mutation was found in the most severe patient (case 4), who had manifested severe mental retardation from infancy, hydrocephalus at the age of 7 years (which led to gait and visual disturbances), and a large hepatomegaly.
Mutations found in the attenuated phenotype
Four missense mutations (G140R, C171R, K227 M, and D308 N), two nonsense mutations (W12X, Q531X), three frameshift mutations (V401 fs, C422 fs, and H441 fs), and one each of splice site mutation (G374G) and an amino acid deletion (L73del) were found in patients with the attenuated clinical phenotype. Case 18, the oldest patient with the highest IQ, had a nonsense mutation Q531X near the C-terminus of I2S. Three frameshift mutations, V401 fs, C422 fs, and H441 fs, also resided near the C-terminus. These termination and frameshift mutations were presumed not to affect the catalytic active site of I2S. However, as with case 7, a nonsense mutation W12X, which resided very close to the N-terminus, was found in patients with the attenuated form. Seven of the mutations identified in this report (L73del, Q75X, G140R, C171R, V401 fs, C422 fs, and H441 fs) were novel mutations. Six of the seven novel mutations were identified in the attenuated form.
3D structural analysis of I2S
The 3D structure of I2S protein revealed that putative active site residues were Asp45, Asn46, Cys84, Arg88, Lys135, His138, Asp334, His335, and Lys347, which corresponded to highly conserved residues in the sulfatase family (Fig. 1, orange spheres). Eukaryotic sulfatases contain a unique posttranslational modification in their active site, a formylglycine residue generated from a cysteine, and the active site is characterized by the presence of a cluster of conserved residues surrounding the Cys84 residue family (Fig. 1, yellow sphere) (Sukegawa et al. 2000). Positions of mutations in the 3D structure of I2S are indicated in Figs. 1 and 2, and effects on the I2S protein are summarized in Table 3.
Mutation in the I2S structure. The red and cyan spheres represent the mutated residues in missense mutations or one amino acid deletion. The active site center, C84 residue, is shown by a yellow sphere. Putative active site residues are represented by an orange sphere. Red spheres indicate residues (1) G224, (2) G336, and (3) R468 related to the severe phenotype. Cyan spheres indicate residues (4) G140, (5) C171, (6) D308, (7) L73, and (8) K227 related to the attenuated phenotype
Mutations found in the severe phenotype
The G224 is a part of the hydrophobic core of the protein structure (Fig. 1, no. 1), and the change from a hydrophobic residue, glycine, to a highly negatively charged large residue, glutamate, is extremely nonconservative. So, mutation G224E should result in the disruption of the tertiary structure itself due to impairment in the hydrophobic core formation. G336 is adjacent to the active site residues, D334 and H335. The G336 V, that is, the change from a small hydrophobic residue, glycine, to a large hydrophobic residue, valine, should result in modification of the active site geometry and reduce enzymatic activity (Fig. 1, no. 2). The R468 is adjacent to the positively charged active site residue, K347, forming a favorable electrostatic field for substrate entrance into the active site cavity. So, R468L or R468 W, that is, the changes from the positively charged residue, arginine, to large hydrophobic residues, leucine or tryptophan, should result in modification of the active site geometry and also in a significant change in substrate affinity (Fig. 1, no. 3). G75X results in early termination of enzyme protein, leading to loss of enzyme activity. These mutations found in severe patients should have strong effects on the tertiary structure of I2S represented by the loss of enzymatic activity.
Mutations found in the attenuated phenotype
The G140, C171, and D308 are surface-exposed residues (Fig. 1, nos. 4–6, respectively) and are distant from the active site. The G140R and C171R, that is, the changes into the hydrophilic residue, arginine, and D308 N, the conservative change, should not alter the overall structure, and the effects of mutations on the enzymatic activity can be minimized. The L73del mutation involves the loss of one amino acid of the beta sheets, and it occurs on the peripheral part of the IDS protein (Fig. 1, no. 7), so changes could preserve the protein structure and activity.
The K227 is one residue on a loop region between beta strands near the active site, but the side chain of K227 stretches toward the opposite side of the active site (Fig. 1, no. 8). The lysine residue has a positively-charged nitrogen atom on the top of the side chain, but it has a long hydrophobic root with similar size to the side chain of methionine. The change K227 M should not alter the overall structure or active site geometry significantly, but partial structure of the protein could be more influenced than that observed after changes in surface-exposed residues. This could explain why case 12 with this mutation had a relatively severe mental retardation (IQ of 55) and severe dysostosis multiplex compared with other patients with the attenuated phenotype.
The deletion mutant, Q531X, would produce polypeptides with a loss of 20 amino acids in the C-terminus, but the deleted part contains only two short helices and short loops in the subdomain of I2S (Fig. 2a). The mutant protein should have most of its catalytic domain and subdomain preserving a residual enzymatic activity. The W12X would produce only a short N-terminal part of the I2S structure, resulting in no active enzyme formation.
The frameshift mutation H441 fs looses the C-terminal 110 amino acids of I2S and has an additional four residues. The C422 fs looses the C-terminal 129 amino acids of I2S and has an additional 17 residues, but the sequence is very similar to the corresponding part of the native protein. The V401 fs looses the C-terminal 150 amino acids of I2S and has an additional 28 residues. Structural analysis indicated that these frameshift mutants loose most of the subdomain structure but preserve the main catalytic domain of I2S (Fig. 2b–d). In addition, modeling analyses of mutants with additional polypeptides caused by frameshifts indicated that the additional polypeptides do not alter overall protein structure (data not shown). Previous study of the mutation R443X found in an attenuated form of Hunter disease also suggested the milder effect on the protein by the deletion of the subdomain (Sukegawa et al. 1992).
The G374G does not change amino acids; however, it is considered to activate a cryptic splice site resulting in an in-frame deletion of 20 amino acids, E375–G394 (Bunge et al. 1993). The authors suggested that the mutant with this deletion should preserve its enzymatic activity because the deletion occurred in the long nonconserved region among the sulfatase family (Bunge et al. 1993). In the 3D structure, the deletion would have occurred in a surface loop region between the beta strands, which are distant from the active site of the main catalytic domain. Therefore, the remaining part of the enzyme may have sufficient I2S activity to result in the observed mild clinical phenotype.
Discussion
Since 1990, a number of mutations in the I2S gene have been identified in patients with MPS-II. Some of these mutations are related to the severity of the disease, while genotype/phenotype relationships still remain unclear because of the presence of many complex factors between them. Recently, the 3D structure of N-acetylgalactosamine-6-sulfate sulfatase has been used for analysis of genotype/phenotype correlations in patients with MPS IVA (Sukegawa et al. 2000). Structural analysis of the enzyme could clearly indicate molecular effects on protein structure and function of mutations such as those observed in the present study.
Previous genetic studies of the I2S gene showed some relationships between common mutations and phenotypes. Missense mutations at R88 (R88C, R88G, R88H, R88L, and R88P) have been commonly reported in Europe and Russia (Rathmann et al. 1996; Froissart et al. 1998; Karsten et al. 1998; Balzano et al. 1998; Filocamo et al. 2001). These R88 mutations related to the severe phenotype. According to structural analysis, the R88 residue itself is one of the most important active site residues. This finding suggested that any type of amino acid substitution would affect enzymatic activity in the severe phenotype as observed in patients.
The P86L was identified in Japan, Europe, and South Africa and was related to the severe phenotype (Isogai et al. 1998; Froissart et al. 1998; Vafiadaki et al. 1998; Balzano et al. 1998). One of the authors speculated that this missense mutation activated a cryptic splice acceptor site and produced a truncated mRNA. However, this substitution could directly alter the geometry of the active site residues. This observation indicated that effects of the mutation should also be considered from structural aspects.
The S333L is one of the most common mutations among various ethnicities, and it relates to the severe form (Sukegawa et al. 1995; Olsen et al. 1996; Isogai et al. 1998; Vafiadaki et al. 1998; Froissart et al. 1998; Karsten et al. 1998; Li et al. 1999). The S333 residue is adjacent to the active site residues, D334 and H335. Substitution from serine to leucine at this position should alter the active site geometry as seen in G336 V mutation observed in this study.
In addition to these missense mutations, a unique point mutation, c.1122C>T (G374G), has been found in many other ethnicities, European countries, Russia, and Korea (Bunge et al. 1993; Rathmann et al. 1996; Gort et al. 1998; Balzano et al. 1998; Vafiadaki et al. 1998; Karsten et al. 1998; Hartog et al. 1999; Filocamo et al. 2001). This common mutation appears to relate mainly to the attenuated phenotype, as indicated in our structural analysis, but some patients showed severe manifestations. It was previously indicated that a small amount of correctly spliced mRNA was also produced (Bunge et al. 1993). The synthesis of the small amount of normal I2S could explain the differences between clinical phenotypes.
In contrast to the above mutations, W12X found in this study, which was also reported in Italy (Filocamo et al. 2001), could not be explained by structural analysis. This mutation and many other mutations in exon I were nonsense or frameshift mutations (R8X, R8 fs, W12X, A23 fs, and L24 fs), and they should result in nonactive short polypeptides. However, they were related to the attenuated form. These findings indicate the limitation of structural analysis of the protein and also the necessity for further genetic or biochemical studies.
In addition to the above genetic and structural aspects, precise correlation between genotype and phenotype should be reassessed about the definitions of the clinical phenotypes. For example, the mutation A85T is widely found in the world, but there has been no clear phenotype/genotype relationship (Rathmann et al. 1996; Gort et al. 1998; Froissart et al. 1998; Li et al. 1999). Reasons for the discrepancy would be partly due to the manner of classifying the severity of Hunter disease. The conventional classification has been defined mainly according to the severity regarding intelligence (Young et al. 1982). However, some patients with severe mental disability have relatively mild somatic manifestations, and vice versa (Table 1). In addition, the natural history of Hunter disease is still not well delineated. Previous study using the classification according to the severity regarding intelligence and the skeletal findings could suggest the more precise phenotype classification, including an intermediate type (Hiraizumi et al. 1989), but it required an experienced examiner. Recently, new disability scores such as the Functional Independence Measure, which can be easily applied by the general physician, have been used to delineate the complex phenotype classification in genetic diseases (Colvin et al. 2003; Chung et al. 2004). Further molecular investigations with a reliable disability score should be performed to better understand genotype-phenotype correlation for future therapeutic approaches.
References
Balzano N, Villani GR, Grosso M, Izzo P, Di Natale P (1998) Detection of four novel mutations in the iduronate-2-sulfatase gene. Mutations in brief no. 123. Hum Mutat 11:333
Bunge S, Steglich C, Zuther C, Beck M, Morris CP, Schwinger E, Schinzel A, Hopwood JJ, Gal A (1993) Iduronate-2-sulfatase gene mutations in 16 patients with mucopolysaccharidosis type II (Hunter syndrome). Hum Mol Genet 2:1871–1875
Chung BH, Wong VC, Ip P (2004) Spinal muscular atrophy: survival pattern and functional status. Pediatrics 114:e548–e553
Colvin L, Fyfe S, Leonard S, Schiavello T, Ellaway C, De Klerk N, Christodoulou J, Msall M, Leonard H (2003) Describing the phenotype in Rett syndrome using a population database. Arch Dis Child 88:38–43
Filocamo M, Bonuccelli G, Corsolini F, Mazzotti R, Cusano R, Gatti R (2001) Molecular analysis of 40 Italian patients with mucopolysaccharidosis type II: new mutations in the iduronidase-2-sulfatase (IDS) gene. Hum Mutat 18:164–165
Froissart R, Maire I, Millat G, Cudry S, Birot AM, Bonnet V, Bouton O, Bozon D (1998) Identification of iduronate sulfatase gene alterations in 70 unrelated Hunter patients. Clin Genet 53:362–368
Gort L, Coll MJ, Chabas A (1998) Mutations in the iduronidase-2-sulfatase gene in 12 Spanish patients with Hunter disease. Hum Mutat (Suppl 1):S66–S68
Hartog C, Flyer A, Upadhyaya M (1999) Mutation analysis if iduronate-2-sulphatase gene in 24 patients with Hunter syndrome: characterization of 6 novel mutations. Hum Mutat 14:87
Hiraizumi Y (1989) A clinical and biochemical study of Hunter’s syndrome. Acta Sch Med Univ Gifu 37:343–377
Isogai k, Sukegawa K, Tomatsu S, Fukao T, Song X-Q, Yamada Y, Fukuda S, Orii T, Kondo N (1998) Mutation analysis in the iduronidase-2-sulfatase gene in 43 Japanese patients with mucopolysaccharidosis type II (Hunter disease). J Inherit Metab Dis 21:60–70
Karsten S, Voskoboeva E, Tishkanina S, Pettersson U, Krasnopolskaja X, Bondeson ML (1998) Mutational spectrum of the iduronidase-2-sulfatase (IDS) gene in 36 unrelated Russian MPS II patients. Hum Genet 103:732–735
Kim CH, Hwang HZ, Song SM, Paik KH, Kwon EK, Moon KB, Yoon JH, Han CK, Jin DK (2003) Mutational spectrum of the iduronate 2 sulfatase gene in 25 unrelated Korean Hunter syndrome patients: identification of 13 novel mutations. Hum Mutat 21:449–450
Li P, Bellows AB, Thompson JN (1999) Molecular basis of iduronate-2-sulphatase gene mutations in patients with mucopolysaccharidosis type II (Hunter syndrome). J Med Genet 36:21–27
Neufeld F, Muenzer J (2001) The mucopolysaccharidoses. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic and molecular bases of inherited disease, 8th edn. McGraw Hill, New York, pp 3421–3452
Olsen TC, Eiken HG, Knappskog PM, Kase BF, Mansson JE, Boman H, Apold J (1996) Mutations in the iduronate-2-sulfatase gene in five Norwegians with Hunter syndrome. Hum Genet 97:198–203
Rathmann M, Bunge S, Beck M, Kresse H, Tylki-Szymanska A, Gal A (1996) Mucopolysaccharidosis type II (Hunter syndrome): mutation “hot spots” in the iduronate-2-sulfatase gene. Am J Hum Genet 59:1202–1209
Shi J, Blundell TL, Mizuguchi K (2001) FUGUE: sequence-structure homology recognition using environment-specific substitution tables and structure-dependent gap penalties. J Mol Biol 310:243–257
Sukegawa K, Tomatsu S, Tamai K, Ikeda M, Sasaki T, Masue M, Fukuda S, Yamada Y, Orii T (1992) Intermediate form of mucopolysaccharidosis type II (Hunter disease): a C1327 to T substitution in the iduronate sulfatase gene. Biochem Biophys Res Commun 183:809–813
Sukegawa K, Tomatsu S, Fukao T, Iwata H, Song XQ, Yamada Y, Fukuda S, Isogai K, Orii T (1995) Mucopolysaccharidosis type II (Hunter disease): identification and characterization of eight point mutations in the iduronate-2-sulfatase gene in Japanese patients. Hum Mutat 6:136–143
Sukegawa K, Nakamura H, Kato Z, Tomatsu S, Montano AM, Fukao T, Toietta G, Tortora P, Orii T, Kondo N (2000) Biochemical and structural analysis of missense mutations in N-acetylgalactosamine-6-sulfate sulfatase causing mucopolysaccharidosis IVA phenotypes. Hum Mol Genet 9:1283–1290
Vafiadaki E, Cooper A, Heptinstall LE, Hatton CE, Thornley M, Wraith JE (1998) Mutation analysis in 57 unrelated patients with MPS II (Hunter disease). Arch Dis Child 79:237–241
Voznyi YV, Keulemans JL, van Diggelen OP (2001) A fluorimetric enzyme assay for the diagnosis of MPS II (Hunter disease). J Inherit Metab Dis 24:675–80
Wilson PJ, Morris CP, Anson DS, Occhiodoro T, Bielicki J, Clements PR, Hopewood JJ (1990) Hunter syndrome: isolation of an iduronate-2-sulfatase cDNA clone and analysis of patient DNA. Proc Natl Acad Sci USA 87:8531–8535
Yamada Y, Tomatsu S, Sukegawa K, Suzuki Y, Kondo N, Hopewood JJ, Orii T (1993) Mucopolysaccharidosis type II (Hunter disease): 13 gene mutations in 52 Japanese patients and carrier detection in four families. Hum Genet 92:110–111
Young ID, Harper PS, Newcombe RG, Archer IM (1982) A clinical and genetic study of Hunter’s syndrome. 2. Differences between the mild and severe forms. J Med Genet 19:408–411
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kato, T., Kato, Z., Kuratsubo, I. et al. Mutational and structural analysis of Japanese patients with mucopolysaccharidosis type II. J Hum Genet 50, 395–402 (2005). https://doi.org/10.1007/s10038-005-0266-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-005-0266-4
Keywords
This article is cited by
-
Convergent molecular mechanisms underlying cognitive impairment in mucopolysaccharidosis type II
Metabolic Brain Disease (2022)
-
Clinical, biochemical and molecular characteristics of Filipino patients with mucopolysaccharidosis type II - Hunter syndrome
Orphanet Journal of Rare Diseases (2017)
-
Decreasing Activity and Altered Protein Processing of Human Iduronate-2-sulfatase Mutations Demonstrated by Expression in COS7 Cells
Biochemical Genetics (2012)
-
Molecular analysis of the iduronate‐2‐sulfatase gene in Thai patients with Hunter syndrome
Journal of Inherited Metabolic Disease (2008)
-
Effect of Hunter disease (mucopolysaccharidosis type II) mutations on molecular phenotypes of iduronate‐2‐sulfatase: Enzymatic activity, protein processing and structural analysis
Journal of Inherited Metabolic Disease (2006)