Abstract
In diploid populations of size N, there will be 2 Nμ mutations per nucleotide (nt) site (or per locus) per generation (μ stands for mutation rate). If either the population or the coding genome double in size, one expects 4 Nμ mutations. What is important is not the population size per se but the number of genes (coding sites), the two being often interconverted. Here we compared the total physical length of protein-coding genomes (n) with the corresponding absolute rates of synonymous substitution (KS), an empirical neutral reference. In the classical occupancy problem and in the coupons collector (CC) problem, n was expressed as the mean rate of change (KCC). Despite inherently very low power of the approaches involving averaging of rates, the mode of molecular evolution of the total size phenotype of the coding genome could be evidenced through differences between the genomic estimates of KCC [KCC=1/(ln n + 0.57721) n] and rate of molecular evolution, KS. We found that (1) the estimates of n and KS are reciprocally correlated across taxa (r=0.812; p≪ 0.001); (2) the gamete-cell division hypothesis (Chang et al. Proc Natl Acad Sci USA 91:827–831, 1994) can be confirmed independently in terms of KCC/KS ratios; (3) the time scale of molecular evolution changes with change in mutation rate, as previously shown by Takahata (Proc Natl Acad Sci USA 87:2419–2423, 1990), Takahata et al. (Genetics 130:925–938, 1992), and Vekemans and Slatkin (Genetics 137:1157–1165, 1994); (4) the generation time and population size (Lynch and Conery, Science 302:1401–1404, 2003) effects left their “signatures” at the level of the size phenotype of the protein-coding genome.
Similar content being viewed by others
Introduction
Molecular evolution scaled up to the total length of protein-coding sequences in the genome is probably highly variable. Different species can have dramatically different rates of molecular evolution and dramatically different total sizes of protein-coding genomes. Human immunodeficiency virus has a rate of molecular evolution that is a million times faster, and the coding genome size that is million times smaller, than that of mammals (Fitch 1996). The spontaneous rate of change in RNA viruses tends to go down as the size of the target, or complexity, increases (Drake 1969; Drake and Holland 1999). While some component of genome size evolution takes place within genes, because genome size may be correlated with intron size across the broad phylogenetic sweep (Deutsch and Long 1999; Vinogradov 1999; McLysaght et al. 2000), here we study the relationship between the genomic content of protein-coding DNA (n) and the absolute rate of substitution at synonymous sites, i.e., molecular evolution by point (substitution) mutation (KS).
We make use of the coupon collector (CC) problem-related rate of average change (mutation) per nucleotide site across the length of protein-coding genome in order to suitably express n and compare it with the absolute rate of silent (assumed neutral) substitution, KS. The “CC-mutation rate” [KCC=1/(ln n + 0.57721) n] depends only on the total number of protein-coding nucleotide sites, n, in the given genome. It excludes any prior assumptions about which sites could be more important to the evolution of n (see Methods). An implication is that n-related rate of point substitution, KCC, analogous to KA, might be used to explore mode of selection on total size of the coding genome in phyletic evolution. Consequently, the notion of the KCC/KS ratio is qualitatively comparable to the traditional ratio of rates of substitution at amino acid replacement sites (KA) and at synonymous sites (KA/KS). If the KCC estimate is numerically similar to the mean absolute estimate of KS (expressed on the per-generation basis) in coding DNA, this would hint (within evolutionary and sampling error) at the overall neutrality of evolution of the size phenotype of the coding genome and, by implication, the operation of the generation time effect (GTE) at a level of n. If the KCC value fits a putatively neutral empirical control, KS (expressed on the per-year basis), this would rather suggest a nearly neutral mode of evolution of n with absolute time (rather than a generation length) as an evolutionary timeframe. In each case, n would seem to change on the same time scale as molecular evolution. It is expected that KCC/KS ≤ 1 under the neutral mutation theory (Kimura and Ohta 1974; Kimura 1983), reflecting zero constraint on coding-genome size. This is analogous to pseudogenes in which most amino acid variation is neutral and the apparent KA/KS ratio converges toward 1 (Li et al. 1981). It is true that most proteins are slow-evolving (relative to KS) despite the fact that many may be evolving entirely by positive selection, but we are concerned here only with the total size of protein-coding genome, the individuality of single protein genes being ignored. Any significant deviation of the KCC/KS value from unity can be interpreted as indicating that the n phenotype is under selective pressure and thus likely to be functional. The genomes with KCC/KS > 1 are formally defined as being subject to positive selection; that is, the n-related mutations are accumulating faster than would be expected given the underlying rate of silent substitution. The KCC/KS < 1 would indicate that relatively strong purifying (negative) selection operates against the putative n-related mutations, consistent with the neutral theory of molecular evolution in the present context of coding-genome size. However, the genomes with KCC/KS < 1 may still contain many sites under positive selection on n, but the contribution of those sites to the KCC/KS ratio for the entire protein-coding genome is offset by purifying selection at other sites (the KCC/KS quotient is further interpreted in the last paragraph of the coupon collecting analogy).
Data
The estimates of the haploid genome size (the C-value) and the absolute size of protein-coding genome (n, in nt number) in different species, listed in Table 1, were adduced from http://www.ncbi.nlm.nih.gov/Entrez/Genome/org.htlm, http://www.cbs.dtu.dk/services/GenomeAtlas/index.php, and the Genomemine site http://www.genomics.ceh.ac.uk/cgi-bin/gmine/gminemenu.cgi. Information on individual, well-characterized, viral and retroviral genomes was accessed via the NCBI Refseq number at http://www.ncbi.nlm.nih.gov/PMGifs/Genomes/viruses/html and http://www.ncbi.nlm.hih.gov/retroviruses/. The estimate of n was either obtained as a product of the protein-coding gene number and the median protein length for a given genome or inferred from the percentage of total genomic DNA (C-value) coding for protein (http://www.genomics.ceh.ac.uk/cgi-bin/gmine/gminemenu.cgi). The median lengths (nt) of protein-coding genes of all complete genomes available indicate the following orderings: archaea (median range: 690–750) < bacteria (750–885) < eukaryotes (1038–1158). The same orderings hold when restricted to protein-coding genes of size ≥600 nt (archaea 993–1020; bacteria 1020–1131; eukaryotes 1299–1419). The percent of protein-coding genes ≥600 nt relative to all protein-coding genes of the genome is 52–67% in archaea, 51–74% in bacteria, and 76–80% in eukaryotes (Karlin et al. 2002). Thirty-three nuclear and five organellar protein-coding genomes (Table 1) were examined with the median n of 3.43×105 nt [range, 2.2×103 (maize streak virus) to 4.36×107 (humans)]. The median KCC value was 2.19×10−7 (range, 10−5 to 1.27×10−9).
The absolute KS estimates (per synonymous nt site per year) in coding nuclear and organelle DNA (median, 8×10−9 ; range, 2×10−10 to 7×10−3), obtained from either sequence comparisons (using the fossil record of a divergence time and various genome sequences as outgroups) or experimentally, are credited to contributions by Nei and Gojobori (1986), Wolfe et al. (1987), Gojobori et al. (1990), Lei and Graur (1991), Drake (1991, 1993), Laroche et al. (1997), Ohta (1995), Li (1997), Drake et al. (1998), Martin et al. (1998, 2002), Clark et al. (1999), Eyre-Walker and Keightley (1999), Gianelli et al. (1999), Itoh et al. (1999), Kearney et al. (1999), Provan et al. (1999), Cavelier et al. (2000), Denver et al. (2000), Nachman and Crowell (2000), Palmer et al. (2000), Sigurðardóttir et al. (2000), Suzuki et al. (2000), Birky (2001), Lander et al. (2001), Venter et al. (2001), Chen and Li (2001), McVean and Vieira (2001), Heyer et al. (2001), Yang et al. (2001), Akman et al. (2002), Hughes et al. (2002), Itoh et al. (2002), Kondrashov (2003), Kumar and Subramanian (2002), Umemura et al. (2002), Yi et al. (2002), Britten et al. (2003), Hellmann et al. (2003), Howell et al. (2003), Matsuzaki et al. (2004), and Hanada et al. (2004). For the 38 genomes studied (Table 1), the median KCC/KS=3.2 (range, 0.001–2234). We stress that plastid genomes (NC 001807: Homo sapiens mtDNA; NC 000932: A. thaliana cpDNA; NC 001328: C. elegans mtDNA) were not excluded from the analysis to confirm the broad representability of the data.
Materials and methods
The coupon collecting (CC) analogy
Laplace (1812) introduced the original CC problem, and it has been since discussed as a classical mathematical occupancy problem in several texts on probability, for example Feller (1968). Our random experiment was to sample repeatedly, with replacement, from the population D={1, 2,...,N}. This generates a sequence of independent random variables, each uniformly distributed on D: X1, X2, X3... We shall interpret the sampling in terms of CC: each time the collector buys a certain product she (!) receives a coupon (a baseball card or a toy, for example) which is equally likely to be any one of N types. Thus, in this setting, Xi is the coupon type received on the ith purchase. Let the random variable VN,n denote the number of distinct values in the first n selections. Our interest is the sample size needed to get k distinct sample values: WN,k=min {n:VN,k=k},k=1, 2,..., N. In terms of the CC, this random variable gives the number of products required to get k distinct coupon types. Note that the possible values of WN,k are k,k + 1, k + 2,... We will be particularly interested in WN,N, the sample size needed to get the entire population—the number of products required to get the entire set of coupons.
To give the CC problem a distinctly urn-problem flavor (akin to the ménage problem or the birthday problem), recall that CC is equivalent to placing m balls into N bins (viewed as nucleotide sites) so that no bin is empty. At each step, we sample one of N nucleotide sites with uniform probability. At the moment when all nucleotide sites sustain at least one mutation, most sites will have sustained multiple mutational hits, and only a few will have been mutated just once—a mutation not resulting always in a “substitution.” This study was intended to explore only a total extent of coding sites and was not concerned with classification of sites into “degeneracy classes.” The number of mutation events required such that each site experiences at least one hit is given by Euler’s approximation for the partial sum of the harmonic series, D [D=(ln n + Cn)]. The Euler’s constant C ≈ 0.57721. Note that D is essentially the solution of the CC problem. The sum of contributing mutations until all nucleotide sites have been hit at least once is a biologically meaningful stochastic measure of the physical size of the coding genome viewed in terms of site substitution. For example (Table 1), humans have ~32,500 protein-coding genes equivalent to ~4.36×107 nt (the average protein gene is ~1340 nt long). Because D=7.92×108 {4.36×107 [(ln 4.36×107) + 0.55721]}, its inverse, 1/D, is the harmonic mean of probabilities of substitution, 1/D=KCC=1.26×10−9 per nucleotide site. Typically, the harmonic mean is appropriate (because the substitution rates fluctuate) for situations where the average for rates, their general trend over time, is desired. The main contribution comes from small values. The KCC value is numerically similar to the absolute KS estimate in human exons [1.28×10−9 site×year−1 ; Nachman and Crowell (2000), and see Table 1], so KCC/KS ≈ 1. This sets a provisional scale of comparison for interpreting the relationship between n (or, equivalently, KCC) and the average genomic KS estimate in coding genomes of a broad range of species. That 1/D ≈ KS is to be expected if D is viewed as the rate of accumulation of new neutral mutants (r). Generally, r is 1/u, the reciprocal of the forward mutation rate u. Since presently D=r, it follows that 1/D ≈ KS if genome size is neutral to selection. Because the physical size of the protein-coding DNA (n) of extant genomes has been unchanged for millions of years, the KCC/KS ratio is reflective of n-related substitution rate relative to KS. The KCC estimate is time independent in the sense of being unrelated to any real chronology and depends only on the total number of nucleotides currently coding for proteins. The value of KCC increases as the length of the coding genome decreases for substitutions to have occurred until, arbitrarily, each single site has been mutated. Dimensionally, the KCC/KS ratio yields time, scaled either in years (nt substitutions × site−1)/(nt substitutions × site−1 × years−1) or generations (if KS is × generations−1). This approach utilizes the comparison of two distinct mutation rates (KCC and KS) over a large number of nucleotide sites and mutational accumulation over long evolutionary times.
The estimates of KCC and KS in protein-coding genomes (expressed on the per-year basis) and their ratio across taxa are given in Table 1. The KCC/KS analysis may help explore the influence of selection. Also, the population size effect and the GTE may be expected to leave their signatures on the KCC/KS ratio as the KS value and its timeframe change. Analogous to the ratio of rates of nonsynonymous/synonymous substitution (KA/KS), if either the integral size of the protein coding genome (n) evolves in a neutral manner or an averaging of sites under positive and negative selective pressure takes place, KCC/KS would be expected to be close to one. If selection on n were positive, we would expect increasing deviations in favor of KCC with the KCC/KS ratio significantly greater than one, but if selection were purifying (pressure to conserve n), we would expect the opposite trend (for concrete examples see below). Fully distinguishing the effects of random evolutionary force of genetic drift, relaxed selection, and increased mutation pressure on the KCC/KS ratio is precluded by the similar effects of these forces, which also may act simultaneously.
The KCC and absolute rates of synonymous substitution
The putatively neutral mutation (substitution) rate, KS, is often approximated as a substitution rate at the third bases of codons. The KS vary across loci but have a surprisingly constant range among four major clades—plants, animals, bacteria, and fungi—in spite of enormous differences in cellular organization, body size, generation time, genome size, and ecology of these organisms. We contrasted the KCC values with absolute KS estimates intraspecifically (the latter providing neutral control and reference in real time) because natural selection does not strongly influence the fixation probability at synonymous sites and it, therefore, approximates the spontaneous mutation rate. In contrast to KS, the KCC may reflect the influence of selection. The KCC/KS patterns might, therefore, give helpful indications on the selective forces acting on the same phenotype, integral size of the coding genome, in a different way, depending on the species examined. Importantly, synonymous and nonsynonymous sites are interspersed in the totality of coding DNA (represented by the KCC estimate), and factors such as population size and genomic mutation rate will operate on both of them. For example, that the time scale (i.e., the GTE), or some other effect, operates should become evident from differences between KCC and KS (expressed on a per-year basis versus per-generation basis). For example, the bacterial microbes, with smaller genomes, should have correspondingly slower replication rates or operate at a faster time scale to “compensate” for the difference between the KCC rate and the empirical KS rate expressed on the per-year basis (see Table 1, Numerical examples, and The time scales change with change in rate of evolution).
The generation time effect
Since mistakes in DNA copying (replication fidelity + efficiency of DNA repair) contribute to mutation rate, we should expect that for any given rate of copy error, the more frequently DNA is copied, the more errors will accumulate (Britten 1986). This is known as the GTE. However, note that the frequency of DNA replication is a function of both generation time and the number of cell divisions per generation. For higher animals, the generation-time theory predicts that taxa with shorter reproduction times evolve at a higher rate at selectively neutral DNA sites because they have a greater number of germ-line cell divisions and, therefore, replication-induced mutations per unit time (Laird et al. 1969; Ohta 1993; Esteal and Collet 1994; Wu and Li 1985; Li 1997; Weinreich 2001). This explanation assumes that the higher number of cell divisions per unit time in shorter-generation taxa results from a larger number of gonadal generations per unit time that is not canceled by a possibly greater number of gonadal cell divisions per generation in larger-generation taxa. Assuming approximate neutrality of synonymous sites, the rate of divergence should be proportional to mutation rate, as a reflection of increase in the number of mutations per unit time (Li et al. 1987; Ohta 1993; Eyre-Walker and Gaut 1997) and, therefore, proportional to organismal generation span. We confirmed the germ-cell division hypothesis independently by using the KCC/KS ratios (see “The KCC/KS ratios corroborate the germ-cell division hypothesis”). The GTE is more emphatic for KS than for KA substitutions since the former more faithfully reflect the mutation rate (Ohta 1993). Thus, it seemed fairer to compare KCC with KS rather than with KA. We observed that in species with smaller coding genomes KCC ≈ KS if mutation time was scaled on the per-generation rather than on the per-year basis. If this were a genuine consequence of the GTE, it would argue for neutral evolution of the n phenotype. We observed that in species with long generation times and large n,KCC ≈ KS (on the per-year basis); on the contrary, in frequently replicating species with small n,KCC ≈KS (on the per-generation time scale). The KCC and KS, being measures of substitution at a site, seem to reflect the influence of the number of DNA replications per unit time. Independent confirmation of the germ-cell division hypothesis by using only the KCC/KS ratios would strengthen this contention (see The germ-cell division hypothesis and the KCC/KS ratio in mammals), suggesting that the correlation of KCC and KS across taxa is causative rather than a merely correlative phenomenon (see below).
Species with long generation times and large n generally have a small effective population size, Ne. The slightly deleterious mutations (which would be effectively selected against in large populations) will behave by drifting like effectively neutral mutations. Thus, as generation time increases, its effect on clock rate will be compensated by an increase in the rate of effectively neutral mutations. Also, large n implies low KCC value such that the “molecular clock” for evolution of n should roughly match the KS estimate expressed in absolute time better than generation time. The KCC ≈ KS, if KS is expressed on the per-generation scale rather in absolute time, would be expected in species with small n and large Ne. Indeed, as we observed with real data, KCC ≈ KS (per year) in species with large n and small Ne and KCC ≈ KS (per generation) in species with small n and large Ne (see illustrative Numerical examples).
Results and discussion
We found a strong negative correlation (p≪ 0.0001; Figs. 1 and 2) between the protein-coding genome size (n, or its statistical measure KCC) and rate of protein evolution across a broad range of species. It was more convenient to look for signatures of the GTE and the population size effect of fixation probability at a level of the n phenotype described as a substitution rate KCC. We looked for these signatures by comparing the KCC and KS estimates at different time scales. Note that the KCC–KS test may lose some of its meaning, with some conclusions remaining vague, as a consequence of a difficulty to deal with pitfalls in the KCC–KS test at the moment.
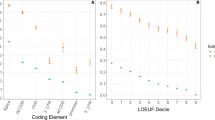
Correlation of the protein-coding genome size (n) and average rate of synonymous substitution (KS) among multiple species. Regression analysis of log-transformed data from 33 species and five organelle genomes (Table 1). Each point specifies the total genomic coding DNA (n, along the abscissa) and the absolute KS estimate (along the ordinate) characteristic of a species. Fluctuation in the data suggests that logarithmic scales are the “natural” scales for these data. There is a highly significant correlation (the Pearson’s correlation coefficient is r=0.811; p≪0.0001)
Correlation of the coupon-collector-related rate of substitution (KCC) and the KS estimate among multiple species. Regression analysis of complete log-transformed data (Table 1). Each point specifies the KCC estimate (along the abscissa) and the absolute KS estimate (along the ordinate) for a given species. There is a highly significant correlation (Pearson’s correlation coefficient is r=0.812; p≪0.0001)
The population size bottleneck versus genome size
At the time of speciation, the protein-coding genome size, being often more ancient than the speciation event itself, rarely changes substantially, if at all, whereas Ne goes through a bottleneck. For a small Ne, the proportion of nondeleterious mutations, which have some chance of spreading, is much larger than for a large Ne (see Eyre-Walker et al. 2002 for the significance of Ne in the nearly neutral mutation model). However, in genomes with large n, as seen in species with small Ne (e.g., mammals), more proteins are expected to be important, being more or less deep in the complex protein-regulatory networks (Hirsh and Fraser 2001; Fraser et al. 2002, and see the following section). The genomes with a large n imply capacious (and more complex) gene-regulating networks and, from the perspective that stronger selection leads to a lower substitution rate, one might expect that the protein genes would be expected to tolerate very few mutations. On the contrary, small protein-coding genomes can tolerate a higher fraction of slightly deleterious mutations and undergo more pronounced effects at population bottlenecks due to neutral drift. Therefore—although in species with small Ne, drift rather than selection predominate, and in those with large Ne, selection is stronger than drift—in species with large n and more complex “interactedness” of proteins, selection may be strong, even at times of population bottleneck such that drift effects may fail to manifest. It is possible that a high number of genetic/metabolic routes constrains genes’ evolutionary rate because mutations in genes involved in multiple pathways decrease flux through many metabolic routes (Kitami and Nadeau 2002). In both humans and mice, the KCC/KS=1 (this is not strictly true, see Numerical examples), suggesting that the extent of n itself does not seem to fulfill a function or play an important role in determining the higher rate of evolution in rodents than in humans (Gu and Li 1992). Our results allow the interpretation that fixation of the n-related point mutations may occur despite their slightly deleterious effects. These mutations are not sufficiently deleterious to be eliminated by purifying selection and can be fixed in the population by random drift, which is affected both by Ne (Lynch and Conery 2003) and n.
The KCC/KS ratios corroborate the germ-cell division hypothesis
We compared the KCC/KS ratios between humans and mice in a quest for a separate confirmation of the contribution of DNA replication errors to the operation of the GTE. Chang et al. (1994) showed that the sex ratio of mutation rates, α, (Myata et al. 1987) is approximately equal to the sex ratio (c) of the number of replications in the germline per generation in males and females. This suggested that errors during doubling rounds of DNA division in the gonadal germinal tissue are the primary source of mutations that are responsible for lineage effects (the germ-cell division hypothesis). The germ-cell division hypothesis is intimately related to the KS (employed in the KCC–KS test) because it predicts a higher KS in organisms with short rather than a long generation time because the number of gametic divisions in males per unit time is expected to be higher for short-lived organisms than for long-lived ones. Because KCC and KS are evolutionarily highly correlated, the ratios of c values and the KCC/KS comparisons in humans and mice should be equal under neutrality. This is to be expected because the historically accumulated numbers of germ-cell divisions, and division-related errors, should affect KS and KCC to a similar extent under neutrality. Consequently, validity of the germ-cell division hypothesis would be falsified at a level of the n phenotype if the quotient of c estimates in humans (h) and mice (m) [c(h)/c(m)] and the quotient of KCC/KS ratios in humans and mice [KCC(h)/KS(h)]/[KCC(m)/KS(m)] would match. The neutral evolution of n (i.e., KCC), in step with KS, would also be supported should the quotient of the KCC/KS values of two mammalian species match the quotient of their respective c estimates. The following shows that these quotients are indeed fairly similar.
The human gametogenesis data suggest the c(h) estimate to be ~6 if the male’s age is 20 and ~10 if the male’s age is 25. In mice, the c(m) value was estimated to be s ~ 2 if the male is 5 months at the time of fertilization. The ratio c(h)/c(m)=2/6 ≈ 0.3. Note that in order to be comparable with available c values, the KS estimates are expressed on the per-generation basis. Now, using the KS(h) estimate of ~10−8 per site per generation and KS(m) ~3×10−9 per site per generation, the KCC(h) and KCC(m) being 1.27×10−9 and 1.34×10−9, respectively (Table 1), we obtain the ratio of 0.284 {[KCC(h)/KS(h)]/[KCC(m)/KS(m)]=(1.27×10−9)×(3×10− 9)/(1×10−8)×(1.34×10−9)]}. A fair agreement between the c(h)/c(m) quotient (~0.3) and the [KCC(h)/KS(h)]/[KCC(m)/KS(m)] quotient (~0.284), despite a considerable difference in generation times between humans (~20 years) and mice (~5 months), and despite very rough estimates of c,KS, and KCC used, could be tentatively interpreted as an independent support for the germ-cell division hypothesis. Because the KCC values do not confound, but reinforce an expected equivalence between the c(h)/c(m) and [KCC(h)/KS(h)]/[KCC(m)/KS(m)] quotients, it also reflects the operation of the GTE at the level of the n phenotype. By inference, a potential influence of the GTE on n is also implied. Because a connection between the rate of point mutation (substitution) and the gamete-cell division is presently suggested, errors in DNA replication in the germline are implied as major determinants of both KS and KCC.
The time scales change with change in rate of evolution
We set the provisional time scale by having KCC/KS (per year) ≈ 1 for large protein-coding genomes (human and mouse, but see Numerical examples). Equivalently, we may have set the time scale by using the small protein-coding genomes to demarcate neutrality. This would require a change in unit scale to start with, from a year (appropriate for large n) to a generation (appropriate for small n), hence KCC/KS (per generation) ≈ 1 for small n. The larger the absolute number of protein genes (or n, which we preferred not to view in terms of Ne) in a genome the lower its KS value (Table 1). We explain this correlation (Figs. 1 and 2) by assuming that as n increases, the proteins have a larger number of protein interactors and a greater effect on organism fitness, which slows the evolutionary rate across the protein-coding genome. Genes that evolve more quickly have less effect on fitness when mutated than do genes that evolve more slowly (Hirsh and Fraser 2001). This is in keeping with the evidence of Fraser et al. (2002) who showed that the connectivity (“interactedness”) of well-conserved proteins is negatively correlated with their rate of evolution. Indeed, the unexpectedly small number of genes discovered in the human genome suggests that complexity of genetic/biological networks may have an important role in vertebrate evolution. The absolute number of new mutations is higher for, and rare mutations are more likely to occur in, larger genomes (or polyploidy), the rate of mutation and Ne being fixed. The fixation of new mutations is the inverse of Ne but it may also be influenced by n. Only in a strictly neutral case is the mutation rate independent of Ne. Consequently, the clustering of KS around 10−9 and KCC around 10−8 (partially explainable if n were “sensed” logarithmically in evolution), may imply that these rates decrease “compensatorily” as a consequence of increase in n. If this inference is correct, n should be strongly correlated to the rate of molecular evolution, as demonstrated in Figs. 1 and 2. Another way to explain rate constancy observed across lineages is that decline in Ne is counterbalanced by and in proportion to the build-up of n, thus providing an opportunity to generate more complex organisms. This possibility is in agreement with a significant correlation between the composite parameter Neμ and genome size recently demonstrated by Lynch and Conery (2003).
If the protein evolution is due in large part to slightly deleterious substitutions (Ohta 1973, 1992; Charlesworth and Charlesworth 1997), the KS should be depressed in large genomes because of the higher likelihood for multiple-protein interactions. Kaufman’s (1993) generalized landscape model, the NK model, also implies that the substitution rate decreases, that is, selective constraints become stronger, as the number of amino acids making the protein increases. In balance, reduction in Ne diminishes the efficiency of selection against mildly deleterious mutations in coding regions, leading to an expansion in coding genome size, as previously proposed by Ohta (1973) and recently by Lynch and Conery (2003). We interpret a general negative correlation between n and KS as suggestive of evolution of n in large genomes by mildly deleterious substitutions, KCC/KS (per year) ≈ 1, and its evolution in a neutral mode, KCC/KS (per generation) ≈ 1, in small genomes.
A slightly different way of looking at the numerical equality between KCC and KS (per year) in the genomes with large n is as follows: As Ohta (1972b) has pointed out, it is not necessary that the fitness of a molecule (in our case, the fitness of the length phenotype of protein-coding DNA) remains precisely the same under a given mutation for that mutation to be considered neutral. A mutation, which produces a change in fitness, will cause the population size of the original and mutant strains to diverge exponentially from one another over time. However, the time constant of this exponential is inversely proportional to the change in fitness. Thus, if the change in fitness is small, its effects will be felt only on very long time scales (corresponding to time scales of planetary development). In effect, the KCC/KS (per year) ≈ 1 implies that the effect of change in fitness imparted by changes in n is very small in large genomes and will be felt only on a very long time scale. Effectively, if not precisely, changes in n are neutral even if there is a genome-wide selection for n.
The correlation between the KCC and KS further suggests that lineage effects affect similarly both the KCC and KS, implying the same cause(s) for both. The cause of lineage effect is most probably the difference in the rate of mutation among taxa due to various factors such as the GTE and metabolic rate, but there are other possibilities. For example, if slightly deleterious mutations segregate at both synonymous and the coding genome size-relevant sites, then differences in Ne would generate correlated differences in rate along lineages and between KCC and KS.
Faster evolution (in absolute time) of coding DNA size in lower organisms than mammals may suggest that the rate of molecular evolution of n is related to generation length rather than absolute time, thus strengthening the random drift hypothesis. There is an apparent equivalence between this suggestion for the lower organisms and the germ-cell division hypothesis in mammals (see above). The fact that KCC ≠ KS (on the per-year basis) in organisms with faster KS and small n, whereas in these same organisms KCC ≈ KS (on the per-replication basis), suggests the operation of GTE, equivalently viewed as a change in evolutionary time scale across the species (see Numerical examples). It should be noted that short genomes that replicate very fast do not have all protein-coding positions fixed in a single species (pseudospecies). Also, the observation in lower organisms that KCC ≈ KS (per generation), while KCC ≠ KS (per year), may be a consequence of fluctuations, at different levels, that occur more rapidly and drastically in smaller populations (especially in vitro). However, it is not clear why these fluctuations would cause the KCC/KS to approach unity exactly when the KS estimate is expressed on the per-replication basis. They would rather be expected to affect the time scale of molecular evolution in a random fashion. Both the large genomes (on the per-year basis) and the bacterial genomes (on a much shorter time scale of replication) have similar KS values, which are on the same order of magnitude as their KCC values. Obviously, as KCC varies in magnitude across species, KS occurs on separate time scales for mutation rate, slow and fast. This indicates the divergence of some scale parameter governing the change in the KCC/KS ratio. The tendency of the KCC/KS (per year) ratio to approximate 1 in the limit of large values of n (or low KCC) implies, as mentioned above, that as the protein-coding genome becomes lengthier, the limit is placed more strongly on the resolution with which selection can detect changes in fitness imposed by increase in n. Equivalently, the change in time scale for short (viral) genomes yields KCC/KS (per replication) ≈ 1.
The change in time scale that we observed has some precedent in evolutionary genetics. It has been studied earlier by Takahata (1990), Takahata et al. (1992), and Vekemans and Slatkin (1994), albeit in an entirely different context of the topology of an allelic genealogy under balancing selection. This topology is similar to that of a neutral allele genealogy but with a different time scale, which (for the coalescent) is equivalent to a change in Ne. We observed that the time scale of molecular evolution of n increases with decreasing values of KS. As n increases, the KCC decreases and assumes a numerical value of a magnitude similar to KS expressed in absolute time. As n reduces, the KCC increases and assumes a value, which is of a magnitude similar to that of corresponding KS but now with a generation as a time unit. That KCC (reflecting the size of the entire coding genome) should equal the absolute empirical KS estimates (obtained on ~1/3 of all coding sites) is to be expected because KS reflects the spontaneous substitution rate, which is unaffected by the type of the site being hit by mutation.
Vekemans and Slatkin (1994) showed by simulation and numerical analysis that the time scales of the gene genealogies are increasing with the number of gene copies. This observation is qualitatively analogous to our evidence that the time scale of evolution of n increases as it becomes larger from a single generation (for small n) to that of approximately a year (for large n). In other words, with suitable change of time scale, the KCC value approximates the empirical KS value for most values of n.
Although we can compare the estimate of KCC against the neutral expectation (KS), we cannot take into account the fluctuations in Ne that may well be important. However, it appears that the time scales of KS are more sensitive to changes in mutation rate than to changes in Ne, the case being similar with allelic genealogies (Vekemans and Slatkin 1994). The number of alleles, unlike the coalescence times, is more sensitive to changes in Ne than to changes in mutation rate. Our gathered data on n and the absolute KS estimates across the species demonstrate essentially the same phenomenon, i.e., the time scale changes in key with change in mutation rate, as shown earlier by Takahata (1990), Takahata et al. (1992), and Vekemans and Slatkin (1994).
Numerical examples
The fact that KCC ≈ KS implies a low level of constraint on n. This could be caused by fixation of slightly deleterious n-related mutations (expected in species with small long-term Ne) from the relaxation of selection on mutations affecting n or from a high rate of adaptive point substitutions affecting n. The first of these explanations seems the most plausible because Ne in hominids is expected to be atypically low. We observed that KCC ≈ KS (per year) in human and mice genomes with large n (~4×107 nt). This might suggest that both KCC and KS are independent of the GTE and the metabolic rate effect and that KCC evolves in a nearly neutral fashion. Therefore, for nearly neutral mutations, the GTE of mutation rate is partially canceled with the population size effect of fixation probability, resulting in a molecular clock, i.e., the KCC ≈ KS (on per-year basis). It is immediately evident that KCC/KS ratio is not proportional to the inverse of Ne, reflecting the irrelevance of n for the faster KS in rodents, as stated above. A more conservative estimate of n in the mouse, and a larger KCC, with KS kept at 1.33×10−9 (Table 1), does not result in the KCC/KS ratio >1 (as might be expected since the Ne for Mus domesticus is ≈10-fold greater than that of humans for both nuclear and mitochondrial genes), which again argues for a very weak selection (i.e., a nearly neutral model of evolution) on the absolute size of a functional stretch of the genome. However, the KS in the human and mouse lineages since the split of primates and rodents (75 MYR), have been recently estimated as 2.2×10−9 and 4.5×10−9, respectively (Waterston et al. 2002). Note that these KS estimates are the averages since the time of divergence and that current KS estimates may differ even more as the difference in generation times between humans and most rodents should be more significant now than shortly after divergence (assuming the GTE on KS). Consequently, the mouse KCC/KS ratio should be <0.3 (1.34×10−9/4.5×10−9) instead of ~1 (as given in Table 1), entirely the consequence of a higher mutation rate in the rodent. The mouse KCC/KS < 0.3 and human KCC/KS ~1 translate to the fact that rodent KA/KS value < primate KA/KS value, the average KA/KS ratio between human and rodent being ~0.2 (Wolfe and Sharp 1993). This would support the GTE hypothesis (shorter generation time driving a higher mutation rate) independently at the level of the n phenotype.
The silent substitution rates are largely a function of n for the RNA viruses (Drake 1969), and the longer the RNA virus genome, the lower its substitution rate and its KCC estimate. For small coding genomes (e.g., viruses), KCC is generally considerably smaller than KS (per year), with KCC/KS < 1. We explain this as a consequence of large Ne, resulting in more effective purifying selection (pressure to conserve n). However, KCC is numerically similar to KS (per generation). One example is the Moloney murine leukemia virus (NC 001501; total genome size, 8332 nt; n, 5217 nt; KCC=2.1×10−5) in which the KS estimate is >3.5×10−6 per replication (Drake 1993; Drake and Holland 1999), which gives KCC/KS < 6.0, probably closer to ~3. The total mutation rate (TMR) in this virus is 2×10−5 (per replication), which gives KCC/TMR ≈ 1. The KS (per year) estimate in this virus is ~1.16×10−3 (Gojobori et al. 1990) and KCC/KS ≈ 0.012. This implies <500 (6/0.012) replications/year, roughly similar to ~331(1.16×10−3/3.5×10−6) replications/year if the coding genome size were not factored in. This similarity suggests the neutrality of evolution of n because factoring for n (using the KCC/KS ratio) does not affect the estimated number of replications per year. Another example is the Rous sarcoma virus (NC 001407; n ~8.06×103 nt) with KS >1.54×10−3 per site per year (Gojobori and Yokoyama 1987; Suzuki et al. 2000) or ~4.6×10−5 per site per replication, implying >33 replications per year (1.54×10−3/4.6×10−5). Since KCC=1.3×10−5, the KCC/KS=0.28 (~1) on the per-replication basis and <0.0087 on the per-year basis, implying >34 generations per year (0.28/0.0082), an agreement that again confirms the neutral evolution of viral n. Yet another example is the HIV-1 virus (NC 001802; n ~8.46×103 nt) with KS ~7.0×10−3 per site per year or ~2.4×10−5 per site per replication. This gives the KCC/KS (per replication)=0.5 (1.3×10−5/2.4×10−5) and KCC/KS (per year)=0.0018, with ~278 replications per year (0.5/0.0018), which matches 292 (7.0×10−3/2.4×10−5), implying again a lack of constraint on the viral n phenotype. These examples accord with the recent evidence (Hanada et al. 2004) that the main source of KS variation in RNA viruses, which may vary by five orders of magnitude (from 1.3×10−7 to 6.2×10−2 per synonymous site per year), was differences in the replication frequency. Further examples (Table 1) also conform to an inverse proportionality between nucleotide mutation rate per generation and n across species (Drake and Holland 1999; Keightley and Eyre-Walker 2000). The change in time scale of molecular evolution of n may reflect higher rates of fixation of slightly deleterious length mutations in organisms, which habitually pass the bottlenecks in Ne (Ohta 1972a), as Ne is negatively correlated to generation time (Chao and Carr 1993; Keightley and Eyre-Walker 2000). However, the magnitude of the effect of population and generation time on KS is not known for real populations, and so it may be that the GTE is not completely canceled out by Ne.
The KCC/KS ratio in endosymbionts versus enteric bacteria
The expected dependence of n on Ne and mutation rate has been supported by observation of reduced n in chronic pathogens and symbionts, which may experience small Ne due to bottlenecks during infection of hosts (Andersson and Andersson 1999; Andersson and Kurland 1998; Moran 1996; Zomorodipour and Andersson 1999) and higher per-site mutation rates (Ochman et al. 1999). Thus, in bacteria and viruses with small Ne and high mutation rates, the selection required to maintain a given genome size increases and should become visible since the genome size and organization are more evolutionarily labile than gene sequences (Huyen and Bork 1998). The KA/KS in E. coli averages about 0.05 whereas its KCC/KS ~3.44 (Table 1), implying a faster (more neutral) rate of evolution of n than the rate of gene evolution in this organism.
Compared with their free-living relatives, endosymbionts feature higher KCC/KS ratios (ranging between 11.8 and 16.9; Table 1), which parallels higher KA/KS ratios observed in bacterial endosymbionts. Moran’s (1996) explanation, that rates of accumulation of mildly deleterious mutations (observed as nonsynonymous changes) are accelerated in the endosymbiotic species, may also serve to explain the larger KCC/KS value for the Buchnera, 16.22, which is about 4.7-fold that for E. coli (16.22/3.44; Table 1) when KS is expressed on an absolute time scale. Since the Buchnera KS is about twice that for low-coding-bias genes of E. coli–S. typhimurium in absolute time (Clark et al. 1999), the difference between 4.7 and 2 reflects a considerable difference in n between these microbes. Therefore, we can make the following ratios: KS(Buchnera)/KS(E.coli) ~2; the ratio of n in Buchnera to E. coli is 0.14 (544,000 nt/4,080,000 nt), and [KCC/KS(Buchnera)]/[KCC/KS(E.coli)] ≈ 4.7. These ratios are roughly similar: 2/0.14 ≈ 16.22 − 2 and (4.7/0.14)/2=16.78. Equivalently, the higher median KCC/KS ratio in endosymbionts (14.8) versus enterics (3.49) parallels the much smaller KS/KA ratios in Buchnera. Clark et al. (1999) have shown this to be consistent with a reduced effect of purifying selection either because of their smaller Ne causing more drift or because of relaxation of selection. As Buchnera shows an average mutation rate that is approximately four-fold higher per generation than in E. coli (Clark et al. 1999), the consistently approximately four-fold (14.8/3.6) higher KCC/KS ratio in endosymbionts versus enterics (Table 1) would parallel this observation, implying that reduced n of endosymbionts and enterics results from a factor other than selection.
Since the KCC/KS ratio varies ~100-fold when KS is expressed on the per-year basis (Table 1) but becomes more nearly constant when expressed per generation, we suggest that the operation of GTE provides a simple explanation for strongly correlated values of KCC and KS and for the difference between KCC and KS for small genomes rather than the difference in KS across species. Because the extent of n correlates with KS (essentially KCC/KS ≈ 1), n should be expected to evolve at a temporal mode similar to that of the protein-coding genes in a given species. Strictly, the KCC value is time-independent, and the KCC/KS ≈ 1 would indicate the appropriate time scale of evolution of n because, dimensionally, KCC/KS=year (or a generation).
The KCC/KS ratio in organellar genomes
The hugely higher KCC/KS ratio in organelle as opposed to that in nuclear protein-coding genomes strongly supports the notion that the difference between KCC and KS in the organelle genomes is real, due to a difference in natural selection rather than in mutation rate or accident (Table 1). High KCC/KS ratio in the asexual mitochondrial genome may reflect strong selective pressure for its survival related to coding-genome size in face of the Muller ratchet and maintenance of genetic conservation (more or less the same set of genes in different organisms) versus structural diversity (variability in size) across taxa. The small n of plastid genomes is perhaps witness to the elimination of mildly deleterious n-related mutations from the mtDNA, thereby retarding Muller’s ratchet. Consequently, a balancing or positive selection operates on n due to narrowed and very specialized functions of the plastids in a mutable environment on different habitats. Conversely, the nuclear coding genome data are compatible with the existence of the widespread neutral mode of size evolution in which size-related mutations may rise to fixation by random drift without significantly affecting the fitness. In genomes with smaller Ne, such as mtDNA and cpDNA, substitution rates of positively selected sites can depend on the total number of new mutations in the population per generation whereas neutral substitution rates depend only on the mutation rate per individual.
The protein-coding segment of the human mtDNA (NC 001807; n ≈ 10,000 nt, is only ~0.023% of the protein-coding nuclear DNA (104/4.36×107). Since the KS estimate for the nuclear protein-coding DNA is ~1.27×10−9 per nt per year, the KS value for the mtDNA coding equivalent should be ~43.5-fold (1/0.023) higher, or ~5.48×10−8 [(1.27×10−9)43.5]. This is indeed similar to the phylogenetic rate of mtDNA evolution in primates (~5×10−8 per nt per year). This value is an order of magnitude less than the pedigree-based estimate of the coding mtDNA mutation rate (1.5×10−7), suggesting a dominant role for purifying selection in the evolution of the mtDNA in natural populations even at the so-called silent sites (Howell et al. 2003). A large KCC/KS ratio (~167) in human protein-coding mtDNA would be one indicator of increased adaptive selection intensity operating on overall size, the miniscule n being strongly favored. We used the pedigree-derived (Denver et al. 2000; Cavelier et al. 2000; Howell et al. 2003), rather than phylogenetically derived, estimates of divergence in the human coding region mtDNA to obtain the KCC/KS ratio. Strikingly elevated KCC/KS value in most organelle DNA (excepting the C. elegans mtDNA) suggests that very strong positive selection plays a key role in the evolutionary conservation of very small n.
The KCC/KS ratio in viral genomes
The rate of substitution in viruses depends both on the rate of mutation per replication and on the “generation time” (replication cycle of the viral genome) of the virus (Li 1997). The fitness of rapidly evolving viruses is not affected by fixation of substitutions by drift. Rapid rates of evolution result from either lack of selective constraint with a consequent accumulation of neutral alleles or from positive Darwinian selection driving advantageous substitutions to fixation. We view increase in KCC in viruses as the n-phenotypic consequence of increase in Ne. With retrospect in absolute time, the median KCC/KS (per year) ≈ 0.053 for the viruses (Table 1) would seem to indicate that the coding-genome sizes may not have experienced substantial adaptive evolution, being under the historical action of negative selection to conserve n. Also, the KCC < KS might seem to suggest that, even for quickly evolving viruses, purifying selection operates, preventing the n-changing mutations from reaching fixation. This is because a smaller KCC/KS value implies a slower rate of evolution of genome size, and a higher value implies a faster rate of evolution. However, it should be recalled (see Numerical examples) that, on the per-replication basis, viral KCC/KS values converge toward unity. Consequently, a change in time scale, from absolute time to a generation length, is required for small genomes with high mutation rates (Takahata 1990; Takahata et al. 1992; Vekemans and Slatkin 1994) in order to analyze the KCC/KS ratios realistically, perhaps because higher mutation rates result in a stronger pressure to increase neutrality. Even if observed in absolute rather than in generational time, there is evidence of strong positive selection on the n phenotype in the human polyoma virus JC (NC 001699; KCC/KS=60.7) and in the HTVL-1 virus (NC 003977; KCC/KS=13), probably reflecting changing biological and ecological regimes, but a relaxation of negative selection on their size cannot be theoretically excluded. The KCC/KS (per year) ~1 (0.8) for the hepatitis B virus (NC 003977; HBV). This is a consequence of KS, which is ~100 times lower than those of retroviruses with similar KCC estimates. Despite the KCC/KS ≈ 1 in HBV on the per-year basis, suggesting the lack of constraint on n, it may also reflect the operation of GTE if the replication frequency of the HBV genome is not as high as that of retroviruses (Gojobori et al. 1990) and/or an independence of genome replication on reverse transcription. If KS is expressed on the per-replication basis, the KCC/KS ratio becomes ≫1, with an interpretation as given above for the human polyoma virus JC and the HTVL-1 virus with high KCC/KS values. Low replication frequency of the HBV genome and/or an independence of its replication on reverse transcription still remain valid explanations.
Conclusions
The conclusions that emerge from this study are: (1) the KCC rates strongly correlate with the absolute estimates of KS, probably as a passive response to differences in evolutionary rates; (2) the ratio KCC/KS (per year) ≈ 1 in large genomes suggests a nearly neutral evolution of the size of the coding genome where the increased generation time is “compensated” by an increase in the rate of effectively neutral mutations; (3i) the GTE (predicted by the neutral theory of evolution) reveals itself in small genomes by the KCC/KS (per year) ≠ 1 because in these same organisms, the ratio KCC/KS (per generation) ≈ 1; and (4) the time scale of evolution of absolute size of the protein-coding genome is keyed to the mutation rate (KS) across taxa. We emphasize that these conclusions are the first approximations being based on crude estimates of KCC and KS. They are valid only to the extent that KCC is considered as realistically reflecting the evolution of n. Also, the KCC estimation assumes the uniform substitution rate among sites whereby the sites particularly important for the evolution of protein-coding genome size may be misrepresented. This problem is, of course, more significant for very small genomes.
The notion of the broad-scale relationship between KCC and KS is unlikely as straightforward as the present demonstrative study might make it seem. The CC–KS test should be extended, and modified, for within-species and between-species studies on a larger sample size. A possibility should be mentioned that there might be an increased efficacy and/or strength of selection on short genome sizes with a stronger relative impact of mutation per site per single replication on the size phenotype of short coding genomes. On the other hand, the KS (per year) estimate is closer to KCC for large coding genomes, possibly because they sustain relatively less impact of selection on coding genome size per single coding site simply due to their large genome size. Obviously, there is ample room for circularity in interpreting KCC/KS ratios reflecting subtle complexity at the population, whole genome, and genic level, which influences the KCC/KS ratio. Putative effects of selection on n suggested by the KCC/KS ratio cannot be meaningfully summarized in a marginal fashion or with respect to a single variable. Future modifications of the KCC–KS test may be effective in compensating for any lack of realism present in the current simple model. They may provide a window on the GTE/population size effect on n and may uncover a possible effect of variation in content of coding DNA on molecular evolution.
References
Akman L, Yamashita A, Watanabe H, Oshima K, Shiba T, Hatori M, Aksoy S (2002) Genome sequence of the endocellular obligate symbiont of tsetse flies, Wigglesworthia glossinidia. Nat Genet 32:402–407
Andersson JO, Andersson SGE (1999) Genome degradation is an ongoing process in Rickettsia. Mol Biol Evol 16:1178–1191
Andersson SGE, Kurland CG (1998) Reductive evolution of resident genomes. Trends Microbiol 6:263–268
Birky CW (2001) The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Ann Rev Genet 35:125–148
Britten RJ (1986) Rates of DNA sequence evolution differ between taxonomic groups. Science 231:1393–1398
Britten RJ, Rowen L, Williams J, Cameron RA (2003) Majority of divergence between closely related DNA samples is due to indels. Proc Natl Acad Sci USA 100:4661–4665
Cavelier L, Jazin E, Jalonen P, Gyllensten U (2000) mtDNA substitution rate and segregation of heteroplasmy in coding and noncoding regions. Hum Genet 107:45–50
Chang BH, Shimmin LC, Shyue SK, Hewett-Emmett D, Li W-H (1994) Weak male-driven molecular evolution in rodents. Proc Natl Acad Sci USA 91:827–831
Chao L, Carr DE (1993) The molecular clock and the relationship between population size and generation time. Evolution 47:688–690
Charlesworth B, Charlesworth D (1997) Rapid fixation of deleterious alleles can be caused by Müller’s ratchet. Genet Res 70:63–73
Chen FC, Li W-H (2001) Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am J Hum Genet 68:444–456
Clark MA, Moran NA, Baumann P (1999) Sequence evolution in bacterial endosymbionts having extreme base composition. Mol Biol Evol 16:1586–1598
Denver DR, Morris K, Lynch M, Vassilieva LL, Thomas WK (2000) High direct estimate of the mutation rate in the mitochondrial genome of Caenorhabditis elegans. Science 289:2342–2344
Deutsch M, Long M (1999) Intron–exon structure of eukaryotic model organisms. Nucleic Acids Res 27:3219–3228
Drake JW (1969) Comparative rates of spontaneous mutation. Nature 221:1132
Drake JW (1991) A constant rate of spontaneous mutation in DNA-based microbes. Proc Natl Acad Sci USA 88:7160–7164
Drake JW (1993) Rates of spontaneous mutation among RNA viruses. Proc Natl Acad Sci USA 90:4171–4175
Drake JW, Holland JJ (1999) Mutation rates among RNA viruses. Proc Natl Acad Sci USA 96:13910–13913
Drake JW, Charlesworth B, Charlesworth D, Crow JF (1998) Rates of spontaneous mutation. Genetics 148:1667–1686
Esteal S, Collet C (1994) Consistent variation in amino-acid substitution rate, despite uniformity of mutation rate: protein evolution in mammals is not neutral. Mol Biol Evol 11:643–647
Eyre-Walker A, Gaut BS (1997) Correlated rates of synonymous site evolution across plant genomes. Mol Biol Evol 5:455–460
Eyre-Walker A, Keightley PD (1999) High genomic deleterious mutation rates in hominids. Nature 397:344–347
Eyre-Walker A, Keightley PD, Smith NGC, Gaffney D (2002) Quantifying slightly deleterious mutation model of molecular evolution. Mol Biol Evol 19:2142–2149
Feller W (1968) An introduction to probability theory and its applications, vol I. Wiley, New York
Fitch WM (1996) The variety of human virus evolution. Mol Phylogenet Evol 5:247–258
Fraser HB, Hirsh AE, Steinmetz LM, Sharfe C, Feldman MW (2002) Evolutionary rate in the protein interaction network. Science 296:750–752
Gianelli FT, Anagnostopoulos T, Green PM (1999) Mutation rates in humans. II. Sporadic mutation-specific rates and rate of detrimental human mutations inferred from hemophilia B. Am J Hum Genet 65:1580–1587
Gojobori T, Yokoyama S (1987) Molecular evolutionary rates of oncogenes. J Mol Evol 26:148–156
Gojobori T, Moriyama EN, Kimura M (1990) Molecular clock of viral evolution and the neutral theory. Proc Natl Acad Sci USA 87:10015–10018
Gu X, Li W-H (1992) Higher rates of amino acid substitution in rodents than in man. Mol Phylogenet Evol 1:211–214
Hanada K, Suzuki Y, Gojobori T (2004) A large variation in the rates of synonymous substitution for RNA viruses and its relationship to a diversity of viral infection and transmission modes. Mol Biol Evol 21:1074–1080
Hellmann I, Zollner S, Enard W, Ebersberger I, Nickel B, Pääbo S (2003) Selection on human genes as revealed by comparisons to chimpanzee cDNA. Genome Res 13:831–837
Heyer E, Zietkiewicz E, Rochowski A, Yotova V, Puymirat J, Labuda D (2001) Phylogenetic and familial estimates of mitochondrial substitution rates: study of control region mutations in deep-rooting pedigrees. Am J Hum Genet 69:1113–1126
Hirsh AE, Fraser HB (2001) Protein dispensability and the rate of evolution. Nature 411:1046–1049
Howell N, Smejkal CB, Mackey DA, Chinnery PF, Turnbull DM, Herrnstadt C (2003) The pedigree rate of sequence divergence in the human mitochondrial genome: there is a difference between phylogenetic and pedigree rates. Am J Hum Genet 72:659–670
Hughes AL, Friedman R, Murray M (2002) Genomewide pattern of synonymous nucleotide substitutions in two complete genomes of Mycobacterium tuberculosis. Emerg Infect Dis 8:1342–1346
Huynen MA, Bork P (1998) Measuring genome evolution. Proc Natl Acad Sci USA 95:5849–5856
Itoh T, Okayama T, Hashimoto H, Takeda J-I, Davis RW, Mori H, Gojobori T (1999) A low rate of nucleotide change in Escherichia coli K-12 estimated from a comparison of the genome sequences between two different substrains. FEBS Lett 450:72–76
Itoh T, Martin W, Nei M (2002) Acceleration of genomic evolution caused by enhanced mutation rate in endocellular symbionts. Proc Natl Acad Sci USA 99:12944–12948
Karlin S, Brocchieri L, Trent J, Blaisdel BE, Mrázek J (2002) Heterogeneity of genome and proteome content in bacteria, archaea, and eukaryotes. Theor Popul Biol 61:367–390
Kauffman SA (1993) The origins of order: self-organization and selection in evolution. Oxford University Press, New York
Kearney CM, Thomson MJ, Roland KE (1999) Genome evolution of tobacco mosaic virus populations during long-term passaging in a diverse range of hosts. Arch Virol 144:1513–1526
Keightley PD, Eyre-Walker A (2000) Deleterious mutations and the evolution of sex. Science 290:331–333
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge, UK
Kimura M, Ohta T (1974) On some principles governing molecular evolution. Proc Natl Acad Sci USA 71:2848–2852
Kitami T, Nadeau JH (2002) Biochemical networking contributes more to genetic buffering in human and mouse metabolic pathways than does gene duplication. Nat Genet 32:191–194
Kondrashov AS (2003) Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum Mutat 21:12–27
Kumar S, Subramanian S (2002) Mutation rates in mammalian genomes. Proc Natl Acad Sci USA 99:803–808
Laird CD, McConaughy BL, McCarthy BJ (1969) Rate of fixation of nucleotide substitutions in evolution. Nature 224:149–154
Lander ES, Linton LM, Birren B et al (255 co-authors) (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Laplace P (1812) Theorie analytique des probabilites, 3rd edn. Courcier, Paris, 1820pp (with supplements)
Laroche J, Li P, Maggia L, Bousquet J (1997) Molecular evolution of angiosperm mitochondrial introns and exons. Proc Natl Acad Sci USA 94:5722–5727
Li W-H (1997) Molecular evolution. Sinauer Assoc. Inc., Sunderland, MA, USA
Li W-H, Graur D (1991) Fundamentals of molecular evolution. Sinauer Assoc. Inc., Sunderland, MA, USA
Li W-H, Gojobori T, Nei M (1981) Pseudogenes as a paradigm of neutral evolution. Nature 292:237–239
Li W-H, Tanimura M, Sharp PM (1987) An evaluation of the molecular clock hypothesis using mammalian DNA sequences. J Mol Evol 25:330–342
Lynch M, Conery JS (2003) The origins of genome complexity. Science 302:1401–1404
Martin W, Stroebe B, Goremykin V, Hansmann S, Hasegawa M, Kowallik KV (1998) Gene transfer to the nucleus and the evolution of chloroplasts. Nature 393:162–165
Martin W, Rujan T, Richly E, Hansen A, Cornelsen S, Lins T, Leister D, Stroebe B, Hasegawa M, Penny D (2002) Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc Natl Acad Sci USA 99:12246–12251
Matsuzaki M, Misumi O, Shin-I T, Maruyama S, Takahara M, Miyagishima S-Y, Mori T, Nishida K, Yagisawa F, Nishida K, Yoshida Y et al (2004) Genome sequence of the ultrasmall unicellular alga Cyanidioschyzon merolae 10D. Nature 428:653–657
McLysaght A, Enright AJ, Skrabanek L, Wolfe KH (2000) Estimation of synteny conservation and genome compaction between pufferfish (fugu) and human. Yeast 17:22–36
McVean GAT, Vieira J (2001) Inferring parameters of mutation, selection and demography from patterns of synonymous site evolution in Drosophila. Genetics 157:245–257
Moran NA (1996) Accelerated evolution and Muller’s ratchet in endosymbiotic bacteria. Proc Natl Acad Sci USA 93:2873–2878
Myata T, Hayashida H, Kuma K, Mitsuyasu K, Yasunaga T (1987) Male-driven molecular evolution: a model and nucleotide sequence analysis. Cold Spring Harb Symp Quant Biol 52:863–867
Nachman MW, Crowell SL (2000) Estimate of the mutation rates per nucleotide in humans. Genetics 156:297–304
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Ochman H, Elwyn S, Moran NA (1999) Calibrating bacterial evolution. Proc Natl Acad Sci USA 96:12638–12643
Ohta T (1972a) Evolutionary rate of cistrons and DNA divergence. J Mol Evol 1:150–157
Ohta T (1972b) Population size and rate of evolution. J Mol Evol 1:305–314
Ohta T (1973) Slightly deleterious mutant substitution in evolution. Nature 246:96–98
Ohta T (1992) The nearly neutral theory of molecular evolution. Annu Rev Ecol Syst 23:263–286
Ohta T (1993) An examination of the generation time effect on molecular evolution. Proc Natl Acad Sci USA 90:10676–10680
Ohta T (1995) Synonymous and nonsynonymous substitutions in mammalian genes and the nearly neutral theory. J Mol Evol 40:56–63
Palmer JD, Adams KL, Cho Y, Parkinson CL, Qiu Y-L, Song K (2000) Dynamic evolution of plant mitochondrial genomes: mobile genes and introns and highly variable mutation rates. Proc Natl Acad Sci USA 97:6960–6966
Provan J, Soranzo N, Wilson NJ, Goldstein DB, Powell W (1999) A low mutation rate for chloroplast microsatellites. Genetics 153:943–947
Sigurðardóttir S, Helgason A, Gulcher JR, Stefansson K, Donnelly P (2000) The mutation rate in the human mtDNA control region. Am J Hum Genet 66:1599–1609
Suzuki Y, Yamaguchi-Kabata Y, Gojobori T (2000) Nucleotide substitution rates of HIV-1. AIDS Rev 2:39–47
Takahata N (1990) A simple genealogical structure of strongly balanced allelic lines and trans-species evolution of polymorphism. Proc Natl Acad Sci USA 87:2419–2423
Takahata N, Satta Y, Klein J (1992) Polymorphism and balancing selection at major histocompatibility complex loci. Genetics 130:925–938
Umemura T, Tanaka Y, Kiyosawa K, Alter HJ, Shih JW-K (2002) Observation of positive selection within hypervariable regions of newly identified DNA virus (SEN virus). FEBS Lett 510:171–174
Vekemans X, Slatkin M (1994) Gene and allelic genealogies at a gametophytic self-incompatibility locus. Genetics 137:1157–1165
Venter JC, Adams MD, Myers EW et al (274 co-authors) (2001) The sequence of the human genome. Science 291:1304–1351
Vinogradov AE (1999) Intron–genome size relationship on a large evolutionary scale. J Mol Evol 49:376–384
Waterston RH, Lindblat-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R et al (222 co-authors) (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562
Weinreich DM (2001) The rates of molecular evolution in rodent and primate mitochondrial DNA. J Mol Evol 52:40–50
Wolfe KH, Sharp PM (1993) Mammalian gene evolution: nucleotide sequence divergence between mouse and rat. J Mol Evol 37:441–456
Wolfe KH, Li W-H, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast and nuclear DNA. Proc Natl Acad Sci USA 84:9054–9058
Wu C-I, Li W-H (1985) Evidence for higher rates of nucleotide substitutions in rodents than man. Proc Natl Acad Sci USA 82:1741–1745
Yang H-P, Tanikawa AY, Kondrashov AS (2001) Molecular nature of 11 spontaneous de novo mutations in Drosophila melanogaster. Genetics 157:1285–1292
Yi S, Ellsworth DL, Li W-H (2002) Slow molecular clock in old world monkeys, apes and humans. Mol Biol Evol 19:2191–2198
Zomorodipour A, Andersson SGE (1999) Obligate intracellular parasites: Rickettsia prowazekii and Chlamydia trachomatis. FEBS Lett 452:11–15
Acknowledgements
We are indebted to two anonymous referees for insightful and constructive suggestions and for pointing out deficiencies in the original version of this paper. This work has been supported by Grant M1300 of the Serbian Ministry of Science.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rajic, Z.A., Jankovic, G.M., Vidovic, A. et al. Size of the protein-coding genome and rate of molecular evolution. J Hum Genet 50, 217–229 (2005). https://doi.org/10.1007/s10038-005-0242-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-005-0242-z