Abstract
The mitochondrial DNA (mtDNA) G13513A mutation in the ND5 subunit gene has been recently reported as a common cause of some phenotypes of mitochondrial myopathy. Until now, the prevalence and characteristics of this mutation in Leigh syndrome (LS) has not been determined. We screened 84 patients with Leigh syndrome (LS) and found the mutation in six (7%) of them. The proportions of mutant mtDNA in muscles were relatively low (42–70%). The onset of symptoms for patients with this mutation was from 9 months to 5 years. It should be noted that five patients had cardiac conduction abnormalities, particularly Wolff-Parkinson-White (WPW) syndrome (three patients). This study suggests that G13513A mutation is a frequent cause of LS and that patients with this mutation may have a characteristic clinical course.
Similar content being viewed by others
Introduction
Leigh syndrome (LS) (MIM 256000) is a progressive neurodegenerative disorder characterized by bilaterally symmetrical lesions in the brainstem and/or basal ganglia in infancy and childhood (Leigh 1951; Pincus 1972; van Erven et al. 1987). LS is associated with defects in mitochondrial energy production. Though the protein subunits of the mitochondrial respiratory chain enzymes are predominantly encoded by nuclear genes, some mitochondrial DNA (mtDNA) mutations are pathogenic in LS, mainly T→G or T→C point mutations occurring at nucleotide position (np) 8993 (8993 mutation) (Tatuch et al. 1992; Santorelli et al. 1993; de Vries et al. 1993) and T→C or T→G at np9176 (Thyagarajan et al. 1995; Campos et al. 1997; Makino et al. 1998). Rare mtDNA mutations such as A3243G, A8344G, and C11777A have also been reported (Koga et al. 2000; Berkovic et al. 1991; Komaki et al. 2003).
The mtDNA G13513A mutation was first reported in patients with MELAS (mitochondrial encephalomyopathy, lactic acidosis, and stroke-like episodes) by Santorelli et al. (1997). This G13513A mutation was discovered to be a more frequent cause of MELAS than previously recognized (Pulkes et al. 1999; Penisson-Besnier et al. 2000; Corona et al. 2001). Moreover, the G13513A mutation was also seen in Leigh-like syndrome (Chol et al. 2003).
In this study, we screened 84 Japanese patients with LS for this mutation and detected it in six of them. We report on the LS patients with the G13513A mutation and discuss their phenotypes.
Patients and methods
We used the following diagnostic criteria for LS in this study, referring to the previous reports (Rahman S, et al. 1996; Makino M, et al. 2000; Arii J, et al. 2000): (1) progressive neurologic disease with motor and/or intellectual developmental delay, (2) elevated lactate levels ( >20 mg/dl) in the blood and/or CSF, and (3) characteristic features on neuroimaging ( i.e., hyperintense lesions on T2-weighted MRI or hypointense lesions on CT scans in the bilateral brainstem and/or basal ganglia). One hundred Japanese patients fulfilled these criteria, but 16 were excluded because of stroke-like episodes and the presence of ragged red fibers (RRF) or strongly succinate dehydrogenase-reactive blood vessels (SSV) in their muscles (Hasegawa et al. 1991).
Muscle samples were obtained by open biopsy. The samples were divided into three portions for histochemical, biochemical, and mtDNA analyses. Serial frozen sections were stained with various histochemical methods, including Gomori trichrome and cytochrome c oxidase (COX). Respiratory chain enzyme activities in skeletal muscles were assayed by slight modification of the published methods (Koga et al. 1988).
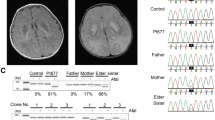
Total DNA was extracted from muscle biopsies and/or blood samples by conventional methods. We first screened deletions and well-known point mutations in mtDNA at np3243, 8344, 8993, and 9176 by the method previously described (Goto et al. 1990; Makino et al. 2000). To detect a G→A mutation at np13513, we amplified 170-bp fragments using oligonucleotide primers corresponding to np13369–13394 and np13514–13538 with a G-to-A mismatch at np13515 in 25 cycles of denaturation (94°C, 30 s), annealing (50°C, 30 s), and extension (72°C, 30 s). The products after digestion with MboI were electrophoresed on a 4% agarose gel and visualized with ethidium bromide under a UV transilluminator (Alpha Imager 2000; Alpha Innotech). The mismatch-containing primer introduced a restriction site for MboI only in the wild type mtDNA, which was cleaved into two fragments of 144 and 26 bp (Fig. 1). The amount of digested versus undigested DNA fragments were measured with a densitometer. To correctly determine the proportion of the mutant mtDNA, we constructed plasmids containing wild or mutant DNA using a TOPO TA cloning kit according to the manufacturer’s instructions. A mixture-template standard curve was used to revise the percentage of mutant mtDNA. We performed total mtDNA sequencing for selected cases, such as those with diffuse COX deficiency in their muscle biopsy, by the method previously described (Akanuma et al. 2000).
Restriction fragment length polymorphism analysis of the G13513A mutation. The PCR products after digestion with MboI were electrophoresed on a 4% agarose gel. The amplified 170-bp fragment was cleaved into two fragments 144 bp and 26 bp in size (the 26-bp fragment is not shown here). W 100% wild type, M 100% mutant, P1~P6 patients’ muscles
We obtained informed consent for muscle pathology and genetic analysis from all patients, and in the case of minors, their parents, using the form approved by the IRB of our institute.
Results
Of the 84 patients, 19 showed diffuse COX deficiency in their muscle biopsy, but their total sequences of mtDNA had no significant mutation (data not shown). These findings indicated that all these cases were caused by nuclear gene mutations such as SURF-1 or others (Zhu et al. 1998; Tiranti et al. 1998). Of the remaining 65 patients, point mutation in mtDNA was detected in 19. Six patients had the G13513A mutation (Fig. 1). The same number of patients had the 8993 mutation (four with T-to-G and two with T-to-C mutations). The T9176C, C11777A, and A8344G mutations were found in four, two, and one patients (Makino et al. 2000; Komaki et al. 2003). All patients with the G13513A mutation had neither other mutation at np3243, 8344, 8993, or 9176 nor deletion. None of the 150 healthy Japanese controls or 120 patients with the A3243G mutation harbored the G13513A mutation.
Clinical features and laboratory data of the six patients with G13513A mutation are summarized in 1. The onset of neurological symptoms other than developmental delay varied from 9 months to 5 years (mean±SD; 28.8±21.1 months). In comparison, onset for patients with the 8993 mutation was 4.6±4.6 months. Ptosis was present in five out of the six patients. It should also be noted that five patients had cardiac conduction abnormalities, especially Wolff-Parkinson-White (WPW) syndrome (three patients).
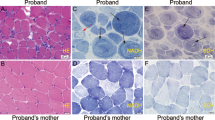
In muscle pathology, there were nonspecific myopathic changes, while one patient (patient 1) had an equivocal SSV (Table 2). The mean percentage of mutant mtDNA in muscles was 59±10% (range 42–70%). No clear correlation was found between the percentage of mutant mtDNA and onset or severity of the disease. The activities of the mitochondrial respiratory chain enzyme were within normal limits in all four patients examined.
Discussion
The G13513A mutation was first reported in patients with MELAS (Santorelli et al. 1997). Its pathogenicity has been confirmed with several lines of evidence as follows: The mutant was detected in heteroplasmic fashion in patients, resulted in amino acid substitution at a highly conserved position in the ND5 subunit, and was absent in healthy controls (Santorelli et al. 1997; Pulkes et al. 1999). Single-fiber polymerase chain reaction studies demonstrated a significantly higher amount of mutant mtDNA in RRFs compared with non-RRFs (Pulkes et al. 1999). Furthermore, Corona et al. (2001) reported two MELAS patients with a 13514 A→G mutation, which hits the same codon affected by the G13513A mutation. The report suggested that the amino acid position of this codon was crucial for the function of complex I. In mouse mitochondria, respiration was tightly controlled by the NADH dehydrogenase ND5 subunit gene (Bai et al. 2000). Therefore, ND5 might also play an important role in human mitochondria. Some recent reports have suggested the validity of this theory (Kirby et al. 2000; Taylor et al. 2002; Liolitsa et al. 2003; Crimi et al. 2003).
The lesions of LS affect brainstem and medulla oblongata, particularly midbrain and pons, and most commonly the tegmentum, the periaqueductal gray, and the posterior colliculi. Other lesions are found in the basal ganglia, spinal cord, cerebellum, and cerebral cortex (Leigh 1951; Pincus 1972; van Erven et al. 1987). Chol et al. (2003) reported that three patients with the G13513A mutation exhibited a selective brainstem involvement in whom they referred to Leigh-like syndrome. We think it unnecessary to use the term Leigh-like because the brainstem lesion is essential for LS. A recent report also used Leigh disease for three patients with this mutation, two of whom had no radiological involvement of basal ganglia (Kirby et al. 2003). Three of our six patients with the G13513A mutation showed involvement of both brainstem and basal ganglia, and the remaining three showed a selective brainstem involvement.
In this study, we found the G13513A mutation and confirmed its relatively high incidence (approximately 7%) in LS patients. However, the frequency was not as high as that of Chol’s report (2003). This mutation was found in the same number of patients as was 8993 in this study, but it seems to be the second most common mtDNA mutation in LS because the 8993 mutation has been reported to account for as many as 15–24% of cases (Santorelli et al. 1993; Rahman et al.1996).
We also found that onset in patients with this mutation was delayed compared with the typical infantile form, or 8993 mutation. Moreover, the clinical signs were characteristic because ptosis and cardiac conduction abnormalities were frequently seen (83%). Although these symptoms could often be recognized in Kearns-Sayre syndrome (KSS), our patients did not have apparent ophthalmoplegia, retinitis pigmentosa, or mtDNA deletion. Interestingly, three patients had WPW syndrome. There are few reports on the association between WPW syndrome and mtDNA mutation. In an examination of the surgical resected accessory atrioventricular pathway in a patient with WPW syndrome, highly abnormal cardiomyocytes with abnormal mitochondria were characterized (Peters et al. 1994). Further study on cardiac tissue specimens, especially the conduction system, using a combined method of morphology, histochemistry, and molecular genetics, is necessary to explain the association between this mutation and cardiac conduction abnormalities.
Contrary to the previous reports (Santorelli et al. 1997; Pulkes et al. 1999; Chol et al. 2003), our patients had no significant respiratory enzyme deficiency in their muscle biopsies; instead, they were affected with LS. One reason for this might be that relatively low heteroplasmy in muscles could not induce an enzyme deficiency in our patients. In mice experiments, it has been shown that tissue with mtDNA abnormality can show normal enzyme activity until the mtDNA abnormality accumulates to a critical level (Nakada et al. 2001). A severely damaged region may have more mutant mtDNA and hence profound enzyme deficiency. Very recently, Kirby et al. (2003) reported that mutant loads of approximately 50% or less induced the defect in complex I amount and activity in skeletal muscle, liver, and fibroblasts. We could not detect apparent enzyme deficiency in the skeletal muscles with the mutant proportion from 42 to 64%. Factors other than the average mutant load should be considered, such as uneven distribution of the individual cells with different mutant load resulting in normal enzyme activity.
It remains unclear how the 13513 mutation may be a cause of both MELAS and LS. Our preliminary results suggested that RRF and/or SSV were present in almost all MELAS patients with this mutation, but they were absent in the LS patients in this study (data not shown). We could not attribute variability of this phenotype solely to the proportion of mutant mtDNA; there must be other factors relating to cell or tissue specificity. Alternatively, we should suppose a possibility that our patients will show the symptoms compatible to MELAS or KSS later in life.
In conclusion, this mutation should be considered as possibly existing in LS patients. Understanding this mutation would provide us more insight into the complex genotype-phenotype relationships in mitochondrial diseases.
References
Akanuma J, Muraki K, Komaki H, Nonaka I, Goto Y (2000) Two pathogenic point mutations exist in the authentic mitochondrial genome, not in the nuclear pseudogene. J Hum Genet 45:337–341
Arii J, Tanabe Y (2000) Leigh syndrome: serial MR imaging and clinical follow-up. Am J Neuroradiol 21:1502–1509
Berkovic SF, Shoubridge EA, Andermann F, Andermann E, Carpenter S, Karpati G (1991) Clinical spectrum of mitochondrial DNA mutation at base pair 8344. Lancet 338:457
Bai Y, Shakeley RM, Attardi G (2000) Tight control of respiration by NADH dehydrogenase ND5 subunit gene expression in mouse mitochondria. Mol Cell Biol 20:905–815
Campos Y, Martin MA, Rubio JC, Solana LG, Garcia-Benayas C, Terradas JL, Arenas J (1997) Leigh syndrome associated with the T9176C mutation in the ATPase 6 gene of mitochondrial DNA. Neurology 49:595–597
Chol M, Lebon S, Benit P, Chretien D, Lonlay P, Goldenberg A, Odent S, Hertz-Pannier L, Vincent-Delorme C, Cormier-Daire V, Rustin P, Rötig A, Munnich A (2003) The mitochondrial DNA G13513A MELAS mutation in the NADH dehydrogenase 5 gene is a frequent cause of Leigh-like syndrome with isolated complex I deficiency. J Med Genet 40:188–191
Corona P, Antozzi C, Carrara F, D’Incerti L, Lama tea E, Tiranti V, Zeviani M (2001) A novel mtDNA mutation in the ND5 subunit of complex I in two MELAS patients. Ann Neurol 49:106–110
Crimi M, Galbiati S, Moroni I, Bordoni A, Perini MP, Lamantea E, Sciacco M, Zeviani M, Biunno I, Moggio M, Scarlato G, Comi GP (2003) A missense mutation in the mitochondrial ND5 gene associated with a Leigh-MELAS overlap syndrome. Neurology 60:1857–1861
Erven PMM van, Cillessen JPM, Eekhoff EMW, Gabreels FJM, Doesburg WAJG, Lemmens WAJG, Sloff JL, Renier WO, Ruitenbeek W (1987) Leigh syndrome, a mitochondrial encephalo(myo)pathy. Clin Neurol Neurosurg 89:217–230
Goto Y, Nonaka I, Horai S (1990) A mutation in the tRNA-Leu(UUR) gene association with the MELAS subgroup of mitochondrial encephalomyopathies. Nature 348:651–653
Hasegawa H, Matsuoka T, Goto Y, Nonaka I (1991) Strongly succinate dehydrogenase-reactive blood vessels in muscles from patients with mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. Ann Neurol 1991:601–605
Kirby DM, Kahler SG, Freckmann ML. Reddihough D, Thorburn DR (2000) Leigh disease caused by the mitochondrial DNA G14459A mutation in unrelated families. Ann Neurol 48:102–104
Kirby DM, Boneh A, Chow CW, Ohtake A, Ryan MT, Thyagarajan D, Thorborn DR (2003) Low mutant load of mitochondrial DNA G13513A mutation can cause Leigh’s disease. Ann Neurol 54:473–478
Koga Y, Nonaka I, Sunohara N, Yamanaka R, Kumagai K (1988) Variability in the activity of respiratory chain enzymes in mitochondrial myopathies. Acta Neuropathol 76:135–141
Koga Y, Akita Y, Takane N, Sato Y, Kato H (2000) Heterogenous presentation in A3243G mutation in the mitochondrial tRNA-Leu(UUR) gene. Arch Dis Child 82:407–411
Komaki H, Akanuma J, Iwata H, Takahashi T, Mashima Y, Nonaka I, Goto Y (2003) A novel mtDNA C11777A mutation in Leigh syndrome. Mitochondrion 2:293–304
Leigh D (1951) Subacute necrotizing encephalomyelopathy in an infant. J Neurol Neurosurg Psychiatry 14:216–221
Liolitsa D, Rahman S, Benton S, Carr LJ, Hanna MG (2003) Is the mitochondrial complex I ND5 gene a hot-spot for MELAS causing mutation? Ann Neurol 53:128–132
Makino M, Horai S, Goto Y, Nonaka I (1998) Confirmation that a T-to-C mutation at 9176 in mitochondrial DNA is an additional candidate for Leigh’s syndrome. Neuromuscul Disord 8:149–151
Makino M, Horai S, Goto Y, Nonaka I (2000) Mitochondrial DNA mutations in Leigh syndrome and their phylogenetic implications. J Hum Genet 45:69–75
Nakada K, Inoue K, Ono T, Isobe K, Ogura A, Goto YI, Nonaka I, Hayashi JI (2001) Inter-mitochondrial complementation: Mitochondria-specific system preventing mice from expression of disease phenotypes by mutant mtDNA. Nature Med 7:934–940
Penisson-Besnier I, Reynier P, Asfar P, PharmD OD, Sortais A, Dubas F, Emile J, Malthiery Y (2000) Recurrent brain hematomas in MELAS associated with an ND5 gene mitochondrial mutation. Neurology 55:317–318
Peters NS, Rowland E, Bennet JG, Green CR, Anderson RH, Severs NJ (1994) The Wolff-Parkinson-White syndrome: the cellular substrate for conduction in the accessory atrioventricular pathway. Eur Heart J 15:981–987
Pincus JH (1972) Subacute necrotizing encephalomyelopathy (Leigh’s disease): a consideration of clinical features and etiology. Dev Med Child Neurol 14:87–101
Pulkes T, Eunson L, Patterson V, Siddiqui A, Wood NW, Nelson IP, Morgan-Hughes JA, Hanna MG (1999) The mitochondrial DNA G13513A transition in ND5 is associated with a LHON/MELAS overlap syndrome and may be a frequent cause of MELAS. Ann Neurol 46:916–919
Rahman S, Blok RB, Dahl HHM, Danks DM, Kirby DM, Chow CW, Christodoulou J, Thorburn DR (1996) Leigh syndrome: clinical features and biochemical and DNA abnormalities. Ann Neurol 39:343–351
Santorelli FM, Shanske S, Macaya A, Devivo DC, DiMauro S (1993) The mutation at nt 8993 of mitochondrial DNA is a common cause of Leigh’s syndrome. Ann Neurol 34:827–834
Santorelli FM, Tanji K, Kulikova R, Shanske S, Vilarinho L, Hays AP, DiMauro S (1997) Identification of a novel mutation in the mtDNA ND5 gene associated with MELAS. Baiochem Biophys Res Commun 238:326–328
Tatuch Y, Christodoulou J, Feigenbaum A, Clarke JTR, Wherret J, Smith C, Rudd N, Petrova-Benedict R, Robinson BH (1992) Heteroplasmic mtDNA mutation (T-to-G) at 8993 can cause Leigh disease when the percentage of abnormal mtDNA is high. Am J Hum Genet 50:852–858
Taylor RW, Morris AAM, Hutchinson M, Turnbull DM (2002) Leigh disease associated with a novel mitochondrial DNA ND5 mutation. Eur J Hum Genet 10:141–144
Thyagarajan D, Shanske S, Vazquez-Memije M, De Vivo D, DiMauro S (1995) A novel mitochondrial ATPase 6 point mutation in familial bilateral striatal necrosis. Ann Neurol 38:468–472
Tiranti V, Hoertnagel K, Carrozzo R, Galimberti C, Munaro M, Grantiero M, Zelante L, Gasparini P, Marzella R, Rocchi M, Bayona-Bafaluy MP, Enriquez J-A, Uziel G, Bertini E, Dionisi-Vici C, Franco B, Meitinger T, Zeviani M (1998) Mutations of SURF-1 in Leigh disease associated with cytochrome c oxidase deficiency. Am J Hum Genet 63:1609–1621
Vries DD de, Engelen BGM van, Gabreels FJM, Ruitenbeek W, Oost BA van (1993) A second missense mutation in the mitochondrial ATPase 6 gene in Leigh’s syndrome. Ann Neurol 34:410–412
Zhu Z, Yao J, Johns T, Fu K, De Bie I, Macmillan C, Cuthbert AP, Newbold RF, Wang J-C, Chevrette M, Brown GK, Brown RM, Shoubridge EA (1998) SURF1, encoding a factor involved in the biogenesis of cytochrome c oxidase, is mutated in Leigh syndrome. Nat Genet 20:337–343
Acknowledgements
We are grateful to all the attending physicians for providing clinical information on the patients, to Drs. Mimaki, Komaki, and Akanuma for valuable discussion, and to Ms. Murayama and Kato for technical assistance. This study was supported in part by a Research Grant (13B-1) for Nervous and Mental Disorders from the Ministry of Health, Labor and Welfare of Japan (YG).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sudo, A., Honzawa, S., Nonaka, I. et al. Leigh syndrome caused by mitochondrial DNA G13513A mutation: frequency and clinical features in Japan. J Hum Genet 49, 92–96 (2004). https://doi.org/10.1007/s10038-003-0116-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-003-0116-1
Keywords
This article is cited by
-
Multisystem mitochondrial diseases due to mutations in mtDNA-encoded subunits of complex I
BMC Pediatrics (2020)
-
Molecular basis of Leigh syndrome: a current look
Orphanet Journal of Rare Diseases (2020)
-
The mitochondrial 13513G>A mutation is most frequent in Leigh syndrome combined with reduced complex I activity, optic atrophy and/or Wolff–Parkinson–White
European Journal of Human Genetics (2007)
-
Sequence polymorphisms of the mtDNA control region in a human isolate: the Georgians from Swanetia
Journal of Human Genetics (2006)