Abstract
A moderate growing strain designated as MPKL 26T was isolated from a soil sample of Bidar Fort, Karnataka, India. The strain MPKL 26T was Gram positive, bent rod in shape. The optimum pH and temperature for growth was 7.0 and 30 °C, respectively. The 16S ribosomal RNA gene sequence analysis revealed that strain MPKL 26T was closely related to S. atrocyanea DSM 20127T (98.09%), S. flava CW 108T (98.04%), S. soli CW 59T (97.99%) and S. notoginsengisoli SYP-B575T (97.0%) and showed DNA–DNA hybridization relatedness (46.05±1.2, 33.56±2.55, 32.56±1.7 and 26.79±2.5, respectively, between these strains) less than the threshold value for the delineation of genomic species. The peptidoglycon type was A3α type with glycine, alanine, lysine and glutamic acid as the amino acids. The whole-cell sugars were fructose, ribose, mannose, glucose and galactose. The polar lipids were diphosphatidylglycerol, phosphatidylglycerol and phosphatidylinositol along with three unknown polar lipids. The fatty acid profile contained C14:0, C16:0, iso-C14:0, iso-C15:0, iso-C16:0, iso-C17:0, anteiso-C15:0, anteiso-C17:0 and summed feature 4 (17:1 iso I/anteiso B). The predominant respiratory quinine was MK-9(H2) with MK-10(H2), MK-8(H2) and MK-8(H4) as minor respiratory quinines. The G+C content of the genomic DNA was 68.8 mol%. On the basis of phenotypic, chemotaxonomic and molecular characteristics, the strain MPKL 26T represents a novel species of the genus Sinomonas, for which the name Sinomonas mesophila sp. nov. is proposed with MPKL 26T as the type strain (=NCIM 5552T= JCM 30094T).
Similar content being viewed by others
Introduction
The genus Sinomonas was first proposed by Zhou et al.1 with the newly isolated strain S. flava CW 108T (the type species of the genus) and S. atrocyanea DSM 20127T (previously classified as Arthrobacter atrocyaneus). Soon after the genus published, another two species Arthrobacter echigonensis and Arthrobacter albidus were reclassified to the genus Sinomonas as S. echigonense and S. albida.2 At the time writing, one more species of this genus, S. notoginsengisoli was proposed by Zhang et al.3 The characteristic features of this genus are: the cells are bent rod in shape and has a high G+C content (66.6–71.8 mol%).1, 2, 3, 4 During the investigation on the biodiversity of microorganisms from soils of Bidar Fort (Karnataka, India), one strain designated as MPLK 26T was isolated; the 16S ribosomal RNA (rRNA) gene sequence analysis revealed that strain MPKL 26T was closely related to S. atrocyanea DSM 20127T (98.09%), S. flava CW 108T (98.04%), Si. soli CW 59T (97.99%) and S. notoginsengisoli SYP-B575T (97.0%). The low 16S rRNA gene sequence relatedness (<98.5%) encouraged us to carry out the phenotypic, chemotaxonomic and molecular characterization in order to classify the taxonomic position of the strain MPLK 26T. On the basis of these results, it was found that the strain MPKL 26T represents a novel species of the genus Sinomonas, for which the name S. mesophila sp. nov. is proposed.
Materials and methods
Strain and culture conditions
Strain MPKL 26T was isolated from the soil sample collected from Bidar Fort (17°55′19′′N 77°31′24′′E), Karnataka, India, by serial dilution plating method using International Streptomyces project (ISP) 4 medium.5 The purified strain was maintained on yeast extract–malt extract agar (ISP 2 medium)5 slants at 4 °C for short-term preservation and as glycerol suspensions (20%, v/v) at −80 °C for long-term preservation. The reference strains S. atrocyanea DSM 20127T, S. flava CW 108T, S. soli CW 59T and S. notoginsengisoli SYP-B575T were selected for the comparison of phenotypic characterization, DNA–DNA relatedness evaluation and chemotaxonomic determination.
Phenotypic characteristics
The morphological, physiological and biochemical characters were observed on YDC and TYB or PYES agar media at 30 °C unless mentioned.6, 7 Gram staining was carried out by using the standard Gram reaction. The morphological properties of strain MPKL 26T were observed with the aid of light microscopy (BH-2; Olympus optical co. Ltd., Tokyo, Japan) and scanning electron microscopy (QUANTA 200; FEI company, Hillsboro, OR, USA). For scanning electron microscopy, cultured cells were harvested by centrifugation, washed and suspended in 20 mm phosphate buffer (pH 7.0). The suspended cells were fixed with 2.5% glutaraldehyde. The cells were dehydrated in an ethanol series (v/v) ranging from 30, 60, 90 and 100%. Cells were dried to a critical drying point. Further, samples were coated with gold and observed under a scanning electron microscope. Growth at various temperature range (5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55 and 60 °C) and pH range 4.0–10.0 (at intervals of 1.0 pH unit) was performed using the buffer system as described by Xu et al.8 The sodium chloride tolerance at various concentrations (0, 0.5, 1, 1.5, 2, 3 and 5.0% w/v) was observed. Catalase activity was determined based on the production of bubbles after the addition of a drop of 3% (v/v) H2O2. Oxidase activity was determined based on oxidation of tetramethyl p-phenylenediamine.9 Cellulose, gelatin, starch; Tweens (20, 40, 60 and 80) hydrolysis; milk coagulation and peptonization were performed as described by Gonzalez et al.10 and other biochemical test was performed by standard methods.11 The enzymatic activities were determined by the API ZYM stripe (bioMérieux, France) according to the manufacturer’s instruction. Utilization of various substrates as sole carbon sources and chemical sensitivity assays was determined by Biolog GN III (Biolog Inc., Hayward, CA, USA) microplates according to the manufacturer’s instruction.
Chemotaxonomy
The isomer of amino acids in purified cell wall and whole-cell sugar hydrolysates were determined according to the procedures described by Hasegawa et al.,12 Lechevalier and Lechevalier13 and Tang et al. (a, b).14,15 Polar lipids were extracted as described by Minnikin et al.16 and identified by two-dimensional TLC.17 Menaquinones were extracted and analyzed using HPLC.18, 19 Cellular fatty acid analysis was performed by using the Microbial Identification System (Sherlock Version 6.1; MIDI database: TSBA6; Sasser 1990). Biomass for fatty acid analysis was obtained from cell grown on tryptose soy agar (Difco, Sparks, MD, USA) at 30 °C for 4 days.
Molecular analysis
Extraction of genomic DNA and PCR amplification of the 16S rRNA gene of the strain MPKL 26T was performed by our earlier standard protocol.20 The sequence obtained was compared with available 16S rRNA gene sequences of cultured species from the EzTaxon-e server (http://eztaxon-e.ezbiocloud.net/).21 Phylogenetic analysis was performed using the software package MEGA version 5.022 after multiple alignment of the sequences using CLUSTAL_X program.23 Distances (using distance options according to Kimura’s two-parameter model; Kimura)24 were calculated and clustering was performed with the neighbor-joining method.25 To determine the support of each clade, bootstrap analysis was performed with 1000 replications.26 The validity of the neighbor-joining tree was evaluated with maximum-likelihood tree using MEGA 5.0. (Arizona State University, Phoenix, AZ, USA).27, 28 The G+C content of the genomic DNA was determined by using reversed phase HPLC using Escherichia coli DH5α as the reference strain.29 The DNA–DNA hybridizations with MPKL 26T and its four reference strains (S. atrocyanea DSM 20127T, S. flava CW 108T, S. soli CW 59T and S. notoginsengisoli SYP-B575T) were carried out by using optical renaturation methods, using eight replications for each hybridization reaction.30
Results and Discussion
Phenotypic characteristics
Strain MPKL 26T was found to be Gram positive, aerobic and non-motile. The scanning electron microscope image (Figure 1) revealed that the cells were bent rod in shape, which is the peculiar character of the genus Sinomonas.1 Temperature for growth ranged from 20 to 40 °C with the optimum growth at 30 °C. The pH for growth ranged from 6 to 8 with the optimum at pH 7. The tolerance to sodium chloride was found to be up to 4% (w/v). The strain MPKL 26T grew well on YDC, PYES or TYB with no dark blue color on YDC media, this characteristic feature differentiates it from the strain S. atrocyanea DSM 20127T. Catalase and Voges–Proskauer test were positive, but oxidase, H2S and indole were negative. Milk coagulation was positive, whereas milk peptonization was negative. The strain hydrolyzed Tween 40 weakly, but not for the other Tweens (20, 60 and 80); this feature differentiated the strain from the other reference type strains (DSM 20127T, CW59T and CW108T). The strain MPKL 26T could utilize dextrin, D-trehalose, D-fructose, 3-methyl glucose, D-glucose-6-PO4, D-fructose-6-PO4, D-aspartic acid, L-aspartic acid, L-glutamic acid, L-histidine, D-galacturonic acid, D-glucuronic acid, α-keto-glutaric acid, D-malic acid and L-malic acid, whereas pectin and methyl pyruvate were weakly utilized; these sources of utilization found consistence in reference type strains (DSM 20127T, CW59T and CW108T). In contrast to the above, D-fucose was only utilized by the strain MPKL 26T. In addition to the above, the production of valine arylamidase and sensitivity to fusidic acid and vancomycin were differential characteristic of the strain MPKL 26T. A details characteristic features of the strain MPKL 26T and its type strains were mentioned in Tables 1 and 2.
Chemotaxonomic characteristics
The peptidoglycon type was A3α type with glycine, alanine, lysine and glutamic acid as the amino acids. The strain MPKL 26T contains fructose, ribose, mannose, glucose and galactose as whole-cell sugars; in comparison with the strain MPKL 26T, the whole-cell sugar, fructose was devoid in the other type strains (S. atrocyanea DSM 20127T, S. flava CW 108T, S. soli CW 59T and S. notoginsengisoli SYP-B575T). The polar lipids were diphosphatidylglycerol, phosphatidylglycerol and phosphatidylinositol along with three unknown polar lipids (Supplementary Figure S1). The cellular fatty acid compositions (%) of strain MPKL 26T and its reference type strains were mentioned in Table 3. The predominant respiratory quinine was MK-9(H2) with MK-10(H2), MK-8(H2) and MK-8(H4) as minor respiratory quinines.
Phylogenetic analysis and DNA–DNA relatedness
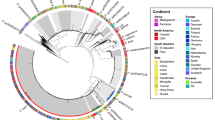
An almost complete 16S rRNA gene sequence (1528 bp) was obtained. The blast result indicated that the strain MPKL 26T showed high similarities to S. atrocyanea DSM 20127T (98.09%), S. flava CW 108T (98.04%), S. soli CW 59T (97.99%) and S. notoginsengisoli SYP-B575T (97.0%). The obtained sequence was submitted to GenBank under the accession number (KJ809567). The neighbor-joining tree (Figure 2) showed that the strain MPKL 26T clustered closely with the members of the genus Sinomonas. The cluster further found stable when the tree constructed by using maximum-likelihood method (Supplementary Figure S2). On the basis of phylogenetic analysis, the new isolate MPKL 26T should be affiliated to the genus Sinomonas.
The G+C content of strain MPKL 26T was determined to be 68.8 mol%, which was within the range of the members of the genus Sinomonas (66.6–71.8 mol%).1, 2, 3, 4 DNA–DNA hybridization values between strain MPKL 26T and its closest phylogenetic neighbors, S. atrocyanea DSM 20127T, S. flava CW 108T, S. soli CW 59T and S. notoginsengisoli SYP-B575T were 46.05±1.2, 33.56±2.55, 32.56±1.7, 26.79±2.5, respectively. The DNA–DNA hybridization values were less than the cutoff point (70%), which was considered to be the threshold value for the delineation of genomic species.31 On the basis of morphological, biochemical and molecular characters, the strain MPKL 26T represents a novel species of the genus Sinomonas, for which we propose the name S. mesophila sp. nov.
Description of S. mesophila sp. nov.
S. mesophila (me.so’phi.la. N.L. fem. adj. mesophila; refers to temperature-loving character)
Cells are Gram positive, non-motile, bent rods and aerobic in nature. Colonies are creamy white, circular and convex after 24 h cultivation at 30 °C on TYB, YDC or PYES agar, respectively. Mesophilic type of growth occurs with a temperature range (20–40 °C) and pH range (6–8) with the optimum growth at 30 °C and pH 7. Growth occurs up to 4% (w/v) sodium chloride. Catalase and Voges–Proskauer test are positive, but oxidase, H2S and indole are negative. Milk coagulation is positive, whereas milk peptonization is negative. Tween 40 is weakly hydrolyzed, whereas starch, cellulose, gelatin and Tweens (20, 60 and 80) are not. Dextrin, D-trehalose, D-turanose, D-raffinose, D-melibiose, D-salicin, α-D-glucose, D-fructose, D-galactose, 3-methyl glucose, L-rhamnose, D-sorbitol, D-mannitol, glycerol, D-glucose-6PO4, D-fructose-6-PO4, D-aspartic acid, glycyl-L-proline, L-alanine, L-arginine, L-aspartic acid, L-glutamic acid, L-histidine, L-pyroglutamic acid, L-serine, D-galacturonic acid, L-galactonic acid lactone, D-gluconic acid, D-glucuronic acid, mucic acid, quinic acid, D-saccharic acid, D-lactic acid methyl ester, α-keto-glutaric acid, D-malic acid, L-malic acid, γ-amino-butryric acid β-hydroxy-D, L-butyric acid, acetoacetic acid, propionic acid and acetic acid are utilized, whereas N-acetyl neuraminic acid and bromo-succinic acid are not utilized. Activity for esterase (C-4), esterase lipase (C-8), leucine arylamidase, valine arylamidase, acid phosphatase, phosphohydrolase, α-glucosidase and β-glucosidase are positive. Sensitive to guanidine hydrochloride, fusidic acid, lithium chloride, tetrazolium blue and vancomycin. The peptidoglycan type is A3α with glycine, alanine, lysine and glutamic acid as the amino acids. The whole-cell sugars are fructose, ribose, mannose, glucose and galactose. The polar lipids are diphosphatidylglycerol, phosphatidylglycerol and phosphatidylinositol along with three unknown polar lipids. The fatty acid profile contains C14:0, C16:0, iso-C14:0, iso-C15:0, iso-C16:0, iso-C17:0, anteiso-C15:0, anteiso-C17:0 and Summed feature 4 (17:1 iso I/anteiso B). The predominant respiratory quinine is MK-9(H2) with MK-10(H2), MK-8(H2) and MK-8(H4) as a minor respiratory quinine. The G+C content of strain MPKL 26T is 68.8 mol%.
The type strain is MPKL 26T (=NCIM 5552T= JCM 30094T), which was isolated from the soil of Bidar fort, Karnataka, India.
Accession codes
References
Zhou, Y . et al. Proposal of Sinomonas flava gen. nov., sp. nov., and description of Sinomonas atrocyanea comb. nov. to accommodate Arthrobacter atrocyaneus. Int. J. Syst. Evol. Microbiol. 59, 259–263 (2009).
Zhou, Y . et al. Description of Sinomonas soli sp. nov., reclassification of Arthrobacter echigonensis and Arthrobacter albidus (Ding et al. 2009) as Sinomonas echigonensis comb. nov. and Sinomonas albida comb. nov., respectively, and emended description of the genus Sinomonas. Int. J. Syst. Evol. Microbiol. 62, 764–769 (2012).
Zhang, M. Y . et al. Sinomonas notoginsengisoli sp. nov., isolated from the rhizosphere of Panax notoginseng. Antonie van Leeuwenhoek 106, 827–835 (2014).
Ding, L., Hirose, T . & Yokota, A. Four novel Arthrobacter species isolated from filtration substrate. Int. J. Syst. Evol. Microbiol. 59, 856–862 (2009).
Shirling, E. B . & Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 16, 313–340 (1966).
Kuhn, D. A . & Starr, M. P. Arthrobacter atrocyaneus, nov. sp., and its blue pigment. Arch. Microbiol. 36, 175–181 (1960).
Wieser, M . et al. Emended descriptions of the genus Micrococcus Micrococcus luteus (Cohn 1872) and Micrococcus lylae (Kloos et al. 1974). Int. J. Syst. Evol. Microbiol. 52, 629–637 (2002).
Xu, P . et al. Naxibacter alkalitolerans gen. nov., sp. nov., a novel member of the family Oxalobacteraceae isolated from China. Int. J. Syst. Evol. Microbiol. 55, 1149–1153 (2005).
Kovacs, N. Identification of Pseudomonas pyocyanea by the oxidase reaction. Nature 178, 703–704 (1956).
Gonzalez, C., Gutierrez, C . & Ramirez, C. Halobacterium vallismortis sp. nov., an amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can. J. Microbiol. 24, 710–715 (1978).
MacFaddin, J. F. in Biochemical tests for identification of medical bacteria, Williams & Wilkins: Baltimore, MD, USA, (1980).
Hasegawa, T., Takizawa, M . & Tanida, S. A rapid analysis for chemical grouping of aerobic actinomycetes. J. Gen. Microbiol. 29, 319–322 (1983).
Lechevalier, M. P . & Lechevalier, H. A. Chemical composition as a criterion in the classification of aerobic actinomycetes. Int. J. Syst. Bacteriol. 20, 435–443 (1970).
Tang, S. K . et al. Zhihengliuella alba sp. nov., and emended description of the genus Zhihengliuella. Int. J. Syst. Evol. Microbiol. 59, 2025–2032 (2009a).
Tang, S. K . et al. Kocuria halotolerans sp. nov., an actinobacterium isolated from a saline soil in China. Int. J. Syst. Evol. Microbiol. 59, 1316–1320 (2009b).
Minnikin, D. E., Collins, M. D . & Goodfellow, M. Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J. Appl. Bacteriol. 47, 87–95 (1979).
Collins, M. D . & Jones, D. Lipids in the classification and identification of coryneform bacteria containing peptidoglycan based on 2, 4-diaminobutyric acid. Appl. Bacteriol. 48, 459–470 (1980).
Collins, M. D., Pirouz, T., Goodfellow, M . & Minnikin, D. E. Distribution of menaquinones in actinomycetes and corynebacteria. J. Gen. Microbiol. 100, 221–230 (1977).
Kroppenstedt, R. M. Separation of bacterial menaquinones by HPLC using reverse phase (RP18) and a silver loaded ion exchanger as stationary phases. J. Liq. Chromatogr. 5, 2359–2367 (1982).
Li, W. J . et al. Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China) and emended description of the genus Georgenia. Int. J. Syst. Evol. Microbiol. 57, 1424–1428 (2007).
Kim, O. S . et al. Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 62, 716–721 (2012).
Tamura, K . et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Thompson, J. D . et al. The CLUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Saitou, N . & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–79 (1985).
Fitch, W. M. Toward defining the course of evolution: minimum change for a specific tree topology. Syst. Zool. 20, 406–416 (1971).
Felsenstein, J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Mesbah, M., Premachandran, U . & Whitman, W. B. Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int. J. Syst. Bacteriol. 39, 159–167 (1989).
Ezaki, T., Hashimoto, Y . & Yabuuchi, E. Fluorometric deoxyribonucleic acid-deoxyribonucleic acid hybridization in microdilution wells as an alternative to membrane filter hybridization in which radioisotopes are used to determine genetic relatedness among bacterial strains. Int. J. Syst. Bacteriol. 39, 224–229 (1989).
Wayne, L. G . et al. International committee on systematic bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int. J. Syst. Bacteriol. 37, 463–464 (1987).
Acknowledgements
We are grateful to Professor Dr Hans-Peter Klenk (DSMZ, Germany) and Dr Yu Zhou (Institute of Quality and Standard for Agro-products, Zhejiang Academy of Agricultural Sciences, China) for their kind providing the reference type strains, and Dr Syed G Dastager (CSIR-National Chemical Laboratory, Pune) for his help in performing API ZYM experiment. This work was funded jointly by projects of China tobacco Yunnan industrial (Nos. 2012JC07 and 2012FL02). WH and WJL extend their appreciation to the Deanship of Scientific Research at King Saud University for funding this work through the research group no RGP-205. W-J Li was also supported by Guangdong Province Higher Vocational Colleges & Schools Pearl River Scholar Funded Scheme (2014).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Prabhu, D., Quadri, S., Cheng, J. et al. Sinomonas mesophila sp. nov., isolated from ancient fort soil. J Antibiot 68, 318–321 (2015). https://doi.org/10.1038/ja.2014.161
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2014.161
This article is cited by
-
Sinomonas cellulolyticus sp. nov., isolated from Loktak lake
Antonie van Leeuwenhoek (2023)
-
First report of Sinomonas halotolerans from Parkinsonia aculeata rhizosphere
Biologia (2023)
-
Amycolatopsis alkalitolerans sp. nov., isolated from Gastrodia elata Blume
The Journal of Antibiotics (2020)
-
Sinomonas halotolerans sp. nov., an actinobacterium isolated from a soil sample
Antonie van Leeuwenhoek (2015)