Abstract
Three novel deoxyribofuranosyl indole derivatives, FG050227 (1), FG050223 (2) and FG050204 (3), were identified as antimicrobial agents effective against Gram-positive bacterial strains and some fungi. The MIC values of (1), (2) and (3) against methicillin-resistant Staphylococcus aureus were 3.0, 6.0 and 13 μg ml−1, respectively. Compounds 1 and 3 had bactericidal activity against exponentially growing S. aureus and inhibited biosynthesis of peptidoglycan, protein, RNA and DNA. Compound 2 was bacteriostatic and inhibited the biosynthesis of protein and RNA. The results indicated that deoxyribofuranosyl indole derivatives could be potential lead compounds for the development of antimicrobial agents.
Similar content being viewed by others
Introduction
The emergence of resistant strains against antimicrobial agents is a serious problem in the clinical field. Staphylococcus aureus strains resistant to penicillin1 and methicillin2 were identified as early as 2 years after their initial use. Vancomycin-resistant enterococcus (VRE) emerged in 1987,3 a few years after vancomycin began to be used worldwide.4 S. aureus that is resistant to relatively intermediate concentrations of vancomycin was first reported in 1997 in Japan.5 Linezolide and daptomycin, the antibiotics effective against methicillin-resistant Staphylococcus aures (MRSA) and VRE, became ineffective within 16 and 27, 8 years after approval by Food and Drug Administration in the USA, respectively. Therefore, the discovery of novel antimicrobial agents effective against multi-drug resistant bacteria is crucial. Toward this aim, we screened chemical compounds against S. aureus. Here, we report the identification of three novel deoxyribofuranosyl indole compounds as potential antimicrobial agents against Gram-positive bacteria, including MRSA and VRE, and some fungi.
Materials and Methods
Microbial strains and culture conditions
The bacterial and fungal strains used in this study are summarized in Table 1. Bacterial cultures were prepared in either Luria Bertani (LB) medium (tryptone 10 g l−1, yeast extract 5 g l−1, NaCl 10 g l−1) or Muller–Hinton Broth (MHB) (DIFCO, Franklin Lakes, NJ, USA). For antimicrobial susceptibility tests, cation-adjusted MHB was used. Fungal suspensions were prepared in Roswell Park Memorial Institute medium (Sigma Aldrich, St Louis, MO, USA).
Chemicals and antibiotics
Vancomycin, daptomycin and norfloxacin were purchased from Wako Pure Chemicals (Osaka, Japan), Sequoia Research Products (Pangbourne, UK), and Sigma Aldrich, respectively. Rifampicin and chloramphenicol were obtained from Nacalai Tesque (Kyoto, Japan). Radiolabelled [3H]N-acetylglucosamine was obtained from GE Healthcare (Chalfont St Giles, UK), [methyl-3H]thymidine and [3H]uridine were obtained from Moravek Biochemical (Brea, CA, USA), and [35S]methionine was obtained from the Institute of Isotopes (Budapest, Hungary). All the chemicals used were of analytical grade.
Chemical library and screening method
Our chemical library contains intermediates of several natural product synthesis projects, (for example, K252a,9 ecteinascidin 743,10 kainic acid,11 etc). The compounds were screened based on MIC values against methicillin-susceptible S. aureus. MICs against bacteria and fungi were determined by broth dilution assay. 12, 13
Incorporation of [3H]N-acetylglucosamine into cell wall peptidoglycans
Measurement of incorporated [3H]N-acetylglucosamine was performed according to Maki et al.14 S. aureus NCTC8325 was used for the assay in the presence of 35 μCi [3H]N-acetyl glucosamine. Each of 1, 2 and 3 (10 × MIC), or vancomycin (100 μg ml−1), or norfloxacin (100 μg ml−1) was applied to see their effect on the incorporation. The radioactivity was counted with a liquid scintillation counter (LS6000SE, Beckman Coulter, Carlsbad, CA, USA).
Incorporation of radiolabeled thymidine, uridine and methionine
S. aureus RN4220 was grown in LB medium overnight at 37 °C. The culture was diluted 200-fold in the same medium and incubated at 37 °C until an OD600 of 0.3 was reached. Either 7 μCi [3H]uridine, 70 μCi [3H]thymidine or 20 μCi [35S]methionine was added to the culture and further incubated at 37 °C for 10 min. Rifampicin (100 μg ml−1), norfloxacin (100 μg ml−1) and chloramphenicol (100 μg ml−1) were used as inhibitors of RNA, DNA and protein synthesis, respectively, and vancomycin (10 μg ml−1) or ampicillin (100 μg ml−1) was used as negative control. Compounds (1), (2) and (3) (10 × MIC) were used for all the assays. Aliquots were collected at the indicated time, diluted 10 times by 5% TCA, and the acid insoluble fraction was obtained by filtration through glass fiber filters (Whatman, GE Healthcare, UK). Radioactivity retained on the filters was measured by a liquid scintillation counter.
Bactericidal assay
The bactericidal assay was performed as per NCCLS guidelines.15 Briefly, overnight culture of S. aureus MSSA1 at 37 °C in MHB was diluted 1000 times with the same medium and cultured for 2 h at 37 °C. For daptomycin, MHB was supplemented with 50 mg l−1 Ca2+. Compounds (1), (2) and (3) (5 × MIC) or daptomycin (5 μg ml−1) were added to 1 ml of the culture and incubated at 37 °C for 0 to 1 h. Culture aliquots were collected at indicated time, diluted and spread on LB agar plates and incubated at 37 °C for 24 h. Cell viability was determined by counting the CFU ml−1. The lower limit of detection was 104 CFU ml−1.
Results
Screening for antibacterial compounds
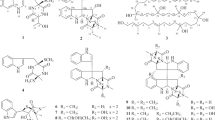
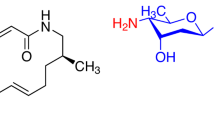
A total of 400 compounds synthesized for the purpose of total synthesis of natural products in the laboratory of one of the authors (TF) were screened for antibacterial activity against S. aureus. Of these 400 compounds, 47 were found to possess antibacterial activity against S. aureus. The MIC values against S. aureus (MSSA1) of these compounds were compared with select the best hit: FG050227 (1). Compounds with a structure similar to (1) (MIC: 3 μg ml−1) were selected from the chemical library (Figure 1: 1–11), from which two additional compounds FG050223 (2, MIC: 6 μg ml−1), and FG050204 (3, MIC: 13 μg ml−1) showed antibacterial activities, whereas (4–11) had no antibacterial activity at a concentration of 500 μM (Table 2).
Most of the compounds, except (9), had a deoxyribofuranosyl indole core. An additional indole ring is present next to deoxyribofuranosyl indole core in (1), (3), (7) and (8) with a sulfonyl group attached to it only in (1). The presence of iodine at the 2-position of the deoxyribofuranosyl indole core confers antibacterial activity (2), whereas an additional indole group in that position abolishes activity (8). Compound (7) similarly showed a loss of antimicrobial activity, whereas the addition of a sulfonyl group to the additional indole (1) restored activity. These findings suggest that the presence of an indole ring alone may reduce antibacterial activity, whereas the addition of a strong electron-withdrawing group, like a sulfonyl group, can recover the activity. Further comparison between (3), (7) and (8) revealed that even in the presence of the additional indole ring, modification of the upper group attached to deoxyribofuranosyl indole core can lead to antibacterial activity.
Antimicrobial spectrum of deoxyribofuranosyl indole compounds
Compounds (1), (2) and (3) were tested for antimicrobial activity against other microorganisms and were found to be effective against Gram-positive bacteria, including several clinical isolates of MRSA and VRE. The compounds were ineffective against Gram-negative bacteria (Table 2). Compounds (1) and (2) exerted antifungal activity against pathogenic true fungi like Candida albicans, Candida tropicalis and Cryptococcus neoformans with MIC values comparable to that against bacteria, whereas (3) showed much weaker antifungal activity compared with its antibacterial activity.
Effect on macromolecule biosynthesis in S. aureus
The incorporation of radiolabelled precursors into macromolecules peptidoglycan, DNA, RNA and protein was monitored on exponentially growing S. aureus with exposure to the compounds over a 3-h time period. Compounds (1) and (3) inhibited the incorporation of [3H]N-acetylglucosamine (Figure 2a), [3H]thymidine (Figure 2b), [3H]uridine (Figure 2c) and [35S]methionine (Figure 2d), suggesting that they inhibit the biosynthesis of peptidoglycan, DNA, RNA and protein. Compound (2) inhibited the incorporation of [3H]uridine (Figure 2c) and [35S]methionine (Figure 2d), whereas its inhibitory effect on the incorporation of [3H]N-acetylglucosamine (Figure 2a) and [3H]thymidine (Figure 2b) was not significant.
Effect of the antimicrobial agents and comparator antibiotics on the incorporation of radiolabelled precursors in S. aureus. Incorporation is shown as counts per minutes (CPM) measured by a liquid scintillation counter. Exponentially growing S. aureus cells were tested for the incorporation of radiolabelled precursors in the presence of the compounds and comparator antibiotics. At time zero, agents were added to exponentially growing cultures. Samples were collected at the indicated time intervals and radioactivity of acid insoluble fractions was measured. (a) [3H] N-acetylglucosamine (b) [3H] thymidine (c) [3H] uridine (d) [35S] methionine. Filled circles: 1 (10 × MIC), filled triangles: 2 (10 × MIC), filled squares: 3 (10 × MIC), open circles: negative controls, open triangles: positive controls, open squares: No drug.
Bactericidal activity against S. aureus
Bactericidal activities of the compounds were determined by killing assays performed on exponentially growing S. aureus over a period of 1 h. Compounds (1) and (3) exerted potent bactericidal activity immediately after the addition of these compounds (Figure 3). The bactericidal activity of these two compounds was stronger than that of daptomycin. Compound (2) did not kill bacteria, indicating that action of this compound is bacteriostatic (Figure 3).
Effect on viability of S. aureus. Exponentially growing S. aureus (∼108 CFU ml−1) was treated with compounds and a comparator antibiotic and cultured. Culture aliquots were collected at the indicated time, diluted and spread on agar plates, and incubated at 37 °C for 24 h. Cell viability was determined by counting the CFU ml−1. Open squares: 1 (5 × MIC), open triangles: 2 (5 × MIC), filled circles: 3 (5 × MIC), filled triangles: daptomycin (5 μg ml−1), filled squares: No drug, dotted line: limit of detection.
Discussion
Three novel dihydrofuranosyl indole compounds exerted antimicrobial activity against Gram-positive pathogens including MRSA and VRE, as well as some fungi. The molecular targets of these compounds are expected to be different from that of vancomycin and methicillin as demonstrated by their similar potency against strains resistant and sensitive to vancomycin and methicillin. Although compounds (1) and (2) showed similar potency against bacteria as well as fungi, (3) showed stronger activity against bacteria than fungi. On the other hand, compound (2) differed from compounds (1) and (3) in the inhibition pattern of macromolecule synthesis, suggesting that these compounds have different cellular targets. This was further verified by the bacteriostatic activity of (2) as opposed to the bactericidal activity of (1) and (3). As (1) and (3) exhibited bactericidal activity with inhibition of all macromolecules unlike (2), which showed bacteriostatic activity with inhibition of RNA and protein syntheses, presence of the indole moiety at the 2-position of deoxyribofuranosyl indole core seems to be important for conferring the bactericidal activity. Bacteriostatic agents, unlike bactericidal agents, inhibit bacterial growth and do not kill bacteria, demanding frequent introduction of the agent, and are thus not the appropriate choice for the treatment of serious, life threatening infections.16, 17 In this study, structural modifications led to changes from bacteriostatic to bactericidal activity. This is an important finding and opens the doors to other structural modifications aimed at converting bacteriostatic agents to bactericidal and non-active compounds to active. These findings suggest that these novel compounds can be used as lead compounds for the development of antimicrobial agents.
The potent inhibition of macromolecule synthesis and rapid bactericidal activity of compounds 1 and 3 suggested that they, like daptomycin18 and XF-73,19 might damage bacterial membranes. Further experiments are required to gain more insight into the exact mechanism of action of these novel compounds.
In conclusion, these compounds are potential lead compounds for the development of antimicrobial agents. Further structural modifications can be made as required with suitable combinations to expect antimicrobial properties and bactericidal activities.
References
Rammelkamp, C. H. & Maxon, T. Resistance of Staphylococcus aureus to the action of penicillin. Proc. Soc. Exp. Biol. Med. 51, 386–389 (1942).
Jevons, M. P. ‘Celbenin’ resistant Staphylococci. Br. Med. J. 14, 124–125 (1961).
Leclercq, R., Derlot, E., Duval, J. & Courvalin, P. Plasmid-mediated resistance to vancomycin and teicoplanin in Enterococcus faecium. New Eng. J. Med. 319, 157–161 (1988).
Moellering,, R. C. Jr. Vancomycin: A 50-year reassessment. Clin. Infect. Dis. 42, S3–S4 (2006).
Hiramatsu, K. et al. Methicillin-resistant Staphylococcus aureus clinical strain with reduced vancomycin susceptibility. J. Antimicrob. Chemother. 40, 135–136 (1997).
Tsiodras, S. et al. Linezolid resistance in a clinical isolate of Staphylococcus aureus. The Lancet 358, 207–208 (2001).
Mangili, A., Bica, I., Snydman, D. R. & Hamer, D. H. Daptomycin-resistant, methicillin-resistant Staphylococcus aureus bacteremia. Clin. Infect. Dis. 40, 1058–1060 (2005).
Marty, F. M. et al. Emergence of a clinical daptomycin-resistant Staphylococcus aureus isolate during treatment of methicillin-resistant Staphylococcus aureus bacteremia and osteomyelitis. J. Clin. Microbiol. 44, 595–597 (2006).
Kobayashi, Y., Fujimoto, T. & Fukuyama, T. Stereocontrolled total synthesis of (+)-K252a. J. Am. Chem. Soc. 121, 6501–6502 (1999).
Endo, A. et al. Total Synthesis of Ecteinascidin 743. J. Am. Chem. Soc. 124, 6552–6554 (2002).
Sakaguchi, H., Tokuyama, H. & Fukuyama, T. Stereocontrolled total synthesis of ()-Kainic Acid. Org. Lett. 9, 1635–1638 (2007).
Clinical and Laboratory Standards Institute. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically- Eighth Edition: Approved Standard M07-A, Clinical and Laboratory Standards Institute, Wayne, PA, (2009).
Clinical and Laboratory Standards Institute. Reference method for broth dilution antifungal susceptibility testing of yeasts: Approved Standard M27-A, Clinical and Laboratory Standards Institute, Wayne, PA, (1997).
Maki, H., Miura, K. & Yamano, Y. Katanosin B and plusbacin A3, inhibitors of peptidoglycan synthesis in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 45, 1823–1827 (2001).
National Committe for Clinical Laboratory Standards. Methods for determining bactericidal activity of antimicrtobial agents: Approved Guideline M-26-A, National Committe for Clinical Laboratory Standards, Wayne, PA, (1999).
Alder, J. & Eisenstein, B. The advance of bacterididal drugs in the treatment of infection. Curr. Infect. Dis. Reports 6, 251–253 (2004).
Stratton, C. W. Dead bugs don’t mutate: Susceptibility issues in the emergence of bacterial resistance. Emerg. Infect. Dis. 9, 10–16 (2003).
Hobbs, J. K., Miller, K., O’Neill, A. J. & Chopra, I. Consequences of daptomycin-mediated membrane damage in Staphylococcus aureus. J. Antimicrob. Chemother. 62, 1003–1008 (2008).
Ooi, N., Miller, K., Hobbs, J., Rhys-Williams, W., Love, W. & Chopra, I. XF-73, a novel antistaphylococcal membrane-active agent with rapid bactericidal activity. J. Antimicrob. Chemother. 64, 735–740 (2009).
Acknowledgements
We thank Ms Matsuzawa, Ms Noguchi (Yoshino) and Ms Kyougoku for technical assistance. This study was supported by the Program for Promotion of Fundamental Studies in Health Sciences of the National Institute of Biomedical Innovation (NIBIO).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Paudel, A., Hamamoto, H., Kobayashi, Y. et al. Identification of novel deoxyribofuranosyl indole antimicrobial agents. J Antibiot 65, 53–57 (2012). https://doi.org/10.1038/ja.2011.110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2011.110
Keywords
This article is cited by
-
One-Pot Multicomponent Synthesis, Antibacterial and Antiproliferative Evaluation of Indole Derivatives
Pharmaceutical Chemistry Journal (2023)
-
Targeting Staphylococcus aureus and its biofilms with novel antibacterial compounds produced by Lactiplantibacillus plantarum SJ33
Archives of Microbiology (2022)
-
Pharmacokinetic parameters explain the therapeutic activity of antimicrobial agents in a silkworm infection model
Scientific Reports (2018)
-
Andrographolide: antibacterial activity against common bacteria of human health concern and possible mechanism of action
Folia Microbiologica (2017)
-
Antibacterial activity of six medicinal Cameroonian plants against Gram-positive and Gram-negative multidrug resistant phenotypes
BMC Complementary and Alternative Medicine (2016)