Abstract
Paclitaxel is a potent and widely used antitumor agent. Considerable worldwide research efforts have been carried out on different production alternatives. Since the description of the first paclitaxel-producing fungi, more than 15 years ago, microorganisms have been investigated as potential alternatives for an environmentally acceptable, relatively simple and inexpensive method to produce paclitaxel. However, in spite of significant research on paclitaxel-producing microorganisms, no commercial fermentation process has been implemented up to now. The aim of this study is to review the present status of research on paclitaxel-producing microorganisms and the ongoing efforts to develop heterologous paclitaxel biosynthesis, and analyze the perspectives of microbial fermentation for paclitaxel production.
Similar content being viewed by others
Introduction
Paclitaxel is a diterpenic polyoxygenated pseudoalkaloid and a well-studied antitumor agent with application against a range of cancer types. The compound, considered a blockbuster anticancer drug, was originally isolated from Taxus brevifolia (Pacific yew), but is now known to be present in the approximately 11 Taxus species existing in the world.1 In the beginning, paclitaxel production depended on its extraction from the bark of yew trees. However, owing to the growing demand for paclitaxel in the 1980s and the shortage of mature trees, other alternative production methods were studied.2 Total synthesis was achieved by two research groups, but the large number of reaction steps required and low yield made the method impractical.3, 4 An important development was the semisynthesis from natural precursors extracted from needles of yew trees (including baccatin III and 10-deacetylbaccatin III). The technique has been used since the 1990s as a renewable source of commercial paclitaxel,5, 6, 7 but the process still relies on plant precursor compounds that are difficult to purify. Cell culture systems were successfully developed for large-scale paclitaxel production even though complex and specialized techniques, with long incubation times, are required.8, 9, 10, 11, 12 In spite of the remarkable progress in the different production alternatives during the past two decades, paclitaxel remains an expensive compound. Consequently, more economic supply is still required so that paclitaxel can be accessible to more people in the world.
Taxomyces andreanae was the first documented case of a paclitaxel-producing fungus.13, 14 Considerable attention was given to fermentation as potential source for paclitaxel. Microorganisms can be environmentally acceptable, relatively simple and inexpensive paclitaxel sources. Potential advantages of microorganisms include a fast growth in simple culture media, the possibility of culturing in large fermenters, resistance to shear stress, easy genetic manipulation, growth at high cell densities (bacteria), and dependable and unrestrained paclitaxel production; the latter two aspects being important requisites for industrial production. Soon after the key discovery of T. andreanae, Cytoclonal Pharmaceutics (Dallas, TX, USA) and Bristol-Myers-Squibb (New York, NY, USA) signed partnership agreements to further develop fungal fermentation of paclitaxel. However, in spite of the big expectations, outstanding advances have not been disclosed up to date. On the contrary, during the past few years it seems that a noticeable pause exists in the development efforts leading to paclitaxel fermentation. Thus, the aim of this study was to review the present status of paclitaxel-producing microorganisms, including those recently discovered, the advances in the search of paclitaxel biosynthetic genes of microbial origin and the ongoing efforts to establish heterologous paclitaxel biosynthesis, and discuss the perspectives of microbial fermentation technology for paclitaxel production in the future.
Paclitaxel-producing microorganisms
Table 1 shows representative microorganisms reported as paclitaxel producers, as well as their origin, the methods used to detect their producing capability or to quantify paclitaxel, as well as the concentrations reported in each case. A number of methods have been used to assign paclitaxel-producing capability and to quantify it, reflecting conceivably the difficulty in detecting paclitaxel at the low concentrations usually found in microbial cultures. The search for paclitaxel-producing microorganisms was primarily directed to the endophytes isolated from yew trees. A search among the endophytic fungi associated with T. brevifolia (Pacific yew) resulted in the discovery of the first fungal paclitaxel producer, the hyphomycetous fungus, T. andreanae.13, 15 Paclitaxel was detected in the extracts from fungal cultures grown in semisynthetic liquid media, by means of MS using FAB and electrospray ionization, chromatography, reactivity with monoclonal antibodies specific for paclitaxel and cytotoxicity studies. Feeding experiments with (14C) acetate and (14C) phenylalanine showed that both served as precursors of fungal (14C) paclitaxel.
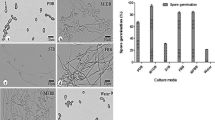
Subsequently, the research group of Strobel16 devoted considerable efforts in the search of paclitaxel-producing fungi from Asiatic, European and other American yews. Hundreds were screened and a number, mostly belonging to ascomycetes and imperfect fungi, were reported as paclitaxel producers. The genera included Pestalotiopsis, Pestalotia, Sporormia, Trichothecium, Tubercularia, Seimatoantlerium, Alternaria, Pithomyces, Monochaetia, Penicillium, Truncatella and Fusarium among others. One endophyte found commonly in yews is Pestalotiopsis sp., sometimes in combination with Pestalotia sp.1, 17 Pestalotiopsis microspora isolated from Taxus wallichiana, was shown to produce optimal yields of paclitaxel in still cultures.17 Other endophytes of T. wallichiana reported as paclitaxel producers included Sporormia minima,18 Trichothecium sp.18 and Seimatoantlerium nepalense, a coelomycetous species.19 Pestalotia heterocornis, a fungus isolated from soil collected in a yew forest, was reported to produce paclitaxel.20 Fungal populations from Taxus baccata were isolated in Italy.21 The isolates were primarily screened for taxane production using a commercial immunoassay. Taxanes were detected in 15 out of 150 fungal isolates, including the genera Alternaria, Aspergillus, Beauveria, Epicoccum, Fusarium, Gelasinospora, Geotrichum, Phoma and Phomopsis. More recently other groups of fungi have been isolated from other yew species. A Tubercularia sp. was isolated from Taxus mairei found in China.22 The fungus was identified through the mechanism of spore production and the characteristics of the spores. A fungus identified preliminary as Taxomyces sp., isolated from Taxus sp., was reported as a paclitaxel producer.23 Notable paclitaxel production was reported for Nodulisporium sylviforme.24, 25 Another paclitaxel-producing strain isolated from the bark of Taxus chinensis was reported as Fusarium solani based on the phylogenetic analysis of its ITS sequence and morphological characteristics.26
The coding sequence for taxadiene synthase (TS), the enzyme involved in formation of the taxane skeleton, was proposed as a novel primary screening method to identify paclitaxel-producing fungi.27 Among 38 endophytic fungi isolated from T. chinensis var. mairei, 12 were ts-positive, but paclitaxel was detected in the culture extracts obtained from only three unidentified strains. Other authors have reported the use of this approach to Mucor rouxianus, an isolate of T. chinensis28 and three different strains of Phomopsis isolated from Taxus cuspidata, Ginkgo biloba and Larix leptolepis.29 Another study suggested that genes coding for 10-deacetylbaccatin III-10-O-acetyl transferase (dbat) and C-13 phenylpropanoyl side chain-CoA acyltransferase (bapt) are more diagnostic as molecular markers.30 Fungi (n=90) isolated from Taxus x media and Taxus yunnanensis were firstly screened for a 200 bp fragment of the dbat gene and subsequently for a 530 bp fragment of the bapt gene. The genes were successfully amplified from DNA of 15 and then 6 strains, respectively. The occurrence of both sequences was confirmed in only three cases. Extracts prepared from cultures of those three fungi, analyzed by electrospray ionization-MS, showed peaks at m/z 876, similar to the sodium adduct ([M+Na]+) obtained with authentic paclitaxel. In addition, the authors reported that fungal paclitaxel was identical to authentic paclitaxel when analyzed by UV and NMR spectroscopy.31 Two out of the three strains were identified as Cladosporium cladosporioides31 and Aspergillus candidus32 according to 18S rDNA sequences and morphological characteristics.
The P. microspora isolated from the bald cypress (Taxodium distichum) showed that plants other than yews can be host of paclitaxel-producing endophytic microorganisms.33 Pestalotiopsis guepinii was isolated from Wollemia nobilis (Wollemi pine), a non-paclitaxel-producing exotic plant.34 Four other fungi isolated from Corylus avellana gave positive immunoassays for paclitaxel.35 Periconia sp. was isolated from Torreya grandifolia (China), a relative of yews, but not a source of paclitaxel.36 Fungal endophytes isolated from other plants were also reported as paclitaxel producers. Seimatoantlerium tepuiense was isolated from Maguireothamnus speciosus an endemic plant of the Southwest of Venezuela.37 Stegolerium kukenani, a hyphomycetous species, was isolated from Stegolepis guianensis on the summits of the Roraima and Kukenan tepuis of Venezuela.38 Other paclitaxel-producing fungi isolated from unusual plants include Pestalotiopsis terminaliae and Chaetomella raphigera39 isolated from Terminalia arjuna, Bartalinia robillardoides40 from Aegle marmelos (an Indian medicinal plant), Phyllosticta tabernaemontanae41 from Wrightia tinctoria, Phyllosticta spinarum42 from Cupressus sp., Phyllosticta citricarpa43 from Citrus medica and Phyllosticta discoreae from Hibiscus rosa-sinensis.44 In summary, fungi reported as paclitaxel producers comprise the orders Botryosphaeriales, Capnodiales, Diaporthales, Eurotiales, Helotiales, Hypocreales, Microascales, Mucorales, Pleosporales, Saccharomycetales, Sordariales and Xylariales (Table 1).
Several genera of bacteria were reported as paclitaxel producers.45, 46 The strains include Bacillus cereus, Bacillus megaterium, Bacillus subtilis, Pantoea sp., Curtobacterium sp. and Sphingomonas sp. However, apart from the two patent documents, no subsequent follow-up scientific papers were published. On the other hand, a number of actinomycete strains were isolated from T. baccata in Italy.21 Micromonospora sp., Kitasatospora sp. and Streptomyces sp. strains were positive to a commercial taxanes immunoassay. In a subsequent study, one strain classified as Kitasatospora sp. on its morphological characteristics and 16S rDNA sequence was reported as paclitaxel producer following HPLC-MS analysis and feeding experiments with labeled precursors.47
In the past years, there has been much interest in finding paclitaxel-producing microorganisms, particularly in fungi. The microorganisms reported so far are considerable in number and variety, thus there is a substantial potential for the development of fermentation processes for commercial paclitaxel production.
Factors affecting paclitaxel production
Optimization of the culture medium is a critical step to maximize paclitaxel production. However, studies on the factors that control paclitaxel production in microorganisms are rare. A commercially available immunoassay kit for paclitaxel was used to evaluate the effect of various media components on paclitaxel production by P. microspora strain Ne-32.48 Low phosphate levels (1 μg ml−1 monosodium phosphate) increased paclitaxel production more than twofold (0.8 ng ml−1, control) while maintaining satisfactory growth. Supplementing the control with Mn, Mg and Zn separately, increased the paclitaxel titers 1.6-, 2.1- and 1.9-fold, respectively. Ergosterol synthesis can compete for precursors with paclitaxel synthesis, and so 12 different inhibitors of the sterol pathway were tested at different concentrations. Paclitaxel production was stimulated by triadimefon (eightfold) and tebuconazol (1.5-fold), but at the expense of reduced mycelial growth. To avoid this drawback but preserve the paclitaxel stimulating effects, the inhibitors were added after 8–10 days culture. Tebuconazol increased paclitaxel production more than 50 fold compared with the control, whereas a 25-fold increase was obtained with triadimefon. As paclitaxel contains a benzoate moiety, sodium benzoate was also tested at several concentrations in combination with low phosphate levels. Maximum production of 3.3 ng ml−1 occurred at 10 μg ml−1 sodium benzoate.
Genes of the paclitaxel biosynthetic pathway
In yew trees, paclitaxel is produced at the end of a long biosynthetic pathway requiring around 19 enzymatic steps from the universal diterpenoid precursor geranylgeranyl diphosphate (GGPP).49, 50, 51 The first committed step comprises the cyclization of GGPP to taxa-4(5), 11(12)-diene by TS.52 Taxadiene is transformed by eight cytochrome P450-mediated oxygenations and two CoA-dependent acylations, the oxetane ring formation and oxidation at C9 to yield 10-deacetyl baccatin III. Next, an acetate group is added at C10 and a side chain at C13 to generate paclitaxel.53, 54, 55, 56, 57 The molecular basis of paclitaxel synthesis in fungi, however, is still almost unknown. Only few studies have reported the screening of paclitaxel-producing fungi using the biosynthetic genes as molecular markers.27, 29, 30, 31, 32, 58, 59 A first partial fungal TS-coding sequence (ts) was reported for M. rouxianus, a fungus isolated from T. chinensis.28 The sequence (632 bp) showed 98% identity to that of T. brevifolia (U48796). The only complete fungal paclitaxel biosynthetic genes available at the GenBank are dbat sequences, belonging to C. cladosporioides and A. candidus, both isolated from T. x media.31, 32 In the first case, the 1546 bp sequence (EU375527) had 99% identity with that of T. x media (EF028093) and 97% identity with that of T. wallichiana var. mairei (AY931014). The dbat sequence (1545 bp) of A. candidus MD3 (EU883596) showed 99% identity with the corresponding gene of T. x media (EF028093) and 97% identity with that of T. wallichiana var. mairei (AY931014). Figure 1 shows the amino acid sequence alignment of the dbat genes from the two cited fungi with two from Taxus sp. PCR was recently used to partially amplify the TS- (EC 4.2.3.17) and BAPT (EC 2.3.1.)-coding sequences from T. andreanae.58 Similarities of 97 and 96% were reported for ts and bapt, respectively, relative to Taxus sp., but the corresponding sequences apparently have not been published.
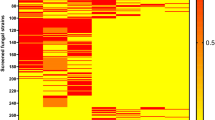
Alignment of deduced amino acid dbat gene sequences of C. cladosporioides (EU375527), A. candidus (EU883596), T. x media (EF028093) and T. wallichiana var. mairei (AY931014). The location of motif (HXXXD) is shown with asterisks (*)—box and the letters in bold show the variable sites. The alignment was created with the ClustalX 2.1 program (Larkin et al., 2007).88
Heterologous expression in bacteria and yeasts
The amount of paclitaxel produced by endophytic fungi is relatively small in comparison to that of Taxus sp. However, the assembly of the entire biosynthetic pathway in microbial heterologous hosts is an attractive alternative for paclitaxel production.60 Some innovative work has been carried out to engineer microbial heterologous hosts using the biosynthetic genes from different yew species. The genes encoding enzymes involved in several steps of the paclitaxel biosynthetic pathway have been isolated and functionally expressed in Escherichia coli and Saccharomyces cerevisiae.49, 51, 61, 62, 63, 64, 65, 66, 67, 68 The enzymes included taxane 2α-O-benzoyltransferase, taxa-4(5),11(12)-diene synthase responsible for cyclization of the universal diterpenoid precursor GGPP to taxa-4(5), 11(12)-diene, the cytochrome P450 taxoid 2α-, 5α-, 7β-, 10β- and 13α-hydroxylases responsible for regio-specific oxygenation on the taxadiene core, the acyl/aroyl CoA-dependent transferases responsible for esterification at the C2–O, C5–O and C10–O positions, and several steps of C13-side-chain assembly. In addition, the biosynthesis of taxadiene has been achieved in cell-free extracts of E. coli overexpressing the genes encoding for isopentenyl diphosphate isomerase, GGPP synthase and TS.61, 69, 70, 71, 72 In addition, considerable engineering of the paclitaxel pathway was carried out in S. cerevisiae.53 Using episomal vectors containing one or more gene cassettes, the independent functional expression of eight paclitaxel biosynthetic genes from T. canadensis was obtained. Furthermore, five sequential steps from isopentenyl diphosphate to taxadiene-5α-acetoxy-10β-ol were integrated in another recombinant yeast. The recombinant strain was able to produce taxadiene at 1000 ng ml−1, whereas taxadiene-5α-ol, a downstream intermediary, was produced in small amounts only (∼25 ng ml−1), thus indicating a pathway restriction at the first cytochrome P450 hydroxylation step. In another study, heterologous genes encoding for biosynthetic enzymes from the taxoid and isoprenoid biosynthetic pathways were introduced along with a regulatory factor inhibiting competitive pathways in S. cerevisiae.73 The yeast host containing a codon-optimized TS from T. chinensis, a truncated hydroxy methyl glutaryl-CoA reductase, UPC2-1 transcription factor gene and the Sulfolobus acidocaldarius GGPP synthase was constructed. The recombinant yeast gave taxadiene at 8700 ng ml−1; that is, a 40-fold increase compared with the host having the TS gene only. In addition, geranylgeraniol was detected at 33 100 ng ml−1. The production of paclitaxel from taxadiene still requires significant modifications of the molecule. Nevertheless, the above-mentioned fundamental study indicate that paclitaxel production in a heterologous host could be feasible.
Genome shuffling
Genome shuffling is a technique in which the genomes of two parent strains are recombined simultaneously at different sites; that is, multiple exchanges and multiple gene recombination. On fusion of parent protoplasts, the progeny may express characteristics of either parent or a hybrid expression.74, 75 Genome shuffling was applied for breeding the native paclitaxel-producing fungus, Nodulisporum sylviforme strain HQD33.76 First, N. sylviforme spores were treated in several mutagenesis rounds (UV, EMS 60Co and NTG) to obtain the strain NCEU-1.77 Next, different combinations of mutagens were applied to the preceding strain to get three hereditarily stable strains named HD100-23, UV40-19 and UL50-6. Protoplast of these three strains were fused randomly, regenerated and named F1-fused colonies. Three further rounds of genome shuffling applied to F1 colonies served to prepare the heterozygote colonies F2, F3 and F4. Fifty strains from the F3 generation and 88 strains from F4 generation were screened for paclitaxel production using TLC, HPLC and MS. The authors reported paclitaxel concentrations at 487.9, 516.4 and 496.7 ng ml−1 for the strains F4–17, F4–26 and F4–70, respectively, as determined by HPLC. These strains had a 31.5, 64.4 and 44.7% higher paclitaxel production compared to the starting strain NCEU-1 (314.1 ng ml−1) and considerably higher values than the native strain (51.1–125.7 ng ml−1). Mass spectra showing each a peak with m/z 855.2, assigned to paclitaxel, were reported in all three cases.
Discussion
The discovery of T. andreanae raised much interest in the prospects of microbial paclitaxel production. Recently, however, it was recognized that the development of paclitaxel production through fermentation has not come up to the initial expectations.58 Consequently, a commercial strain of T. andreanae was reanalyzed in detail using molecular methods (ts and bapt search) as well as metabolite profiling by means of chromatographic, spectroscopic and immunoenzymatic analyses. The partial sequences of ts and bapt genes were successfully amplified, but in spite of these two evidences, no confirmation of the paclitaxel production by the fungus was obtained by metabolite profiling, even after considerable efforts. However, these contradictory results might be due to the fact that fungi cultured on artificially defined media under laboratory conditions usually express a subset of the biosynthetic genes, which encode for biosynthesis of secondary metabolites.78 In fact, successive replating of paclitaxel-producing fungi in semisynthetic medium resulted in a decrease in paclitaxel production. Reactivation of paclitaxel production has been carried out with the use of some chemical compounds including p-hydroxybenzoic acid or serinol.48 More recently, small molecule epigenetic modifiers, such as DNA methyl transferase and histone deacetylase inhibitors, have been used to efficiently release the expression of secondary biosynthetic pathways in fungi.78
At first it appears that paclitaxel-producing fungi are widespread in the world and not confined to the endophytes of yew trees only. However, the study of Staniek et al.58 indicated fundamental aspects in paclitaxel determination that encourages a careful examination of the significance of different methods used either to quantify or assign the occurrence of paclitaxel in different microbial cultures. Monoclonal/polyclonal based immunoassays, TLC, HPLC, HPLC-MS, UV and NMR are frequently used. A common aspect in all these analyses is the requirement of culturing, extraction and purification of paclitaxel. The number of different techniques used to detect paclitaxel likely indicates the difficulty in analyzing the small amounts of paclitaxel found in microbial cultures. For that reason, many authors typically report data obtained with several techniques as evidence of microbial paclitaxel-producing capability. However, the risk that qualitative and quantitative paclitaxel data could be biased by the method and conditions of analysis should be recognized. Immunoassays for paclitaxel can result in a positive reaction if paclitaxel-like molecules are present; that is, other taxanes.79 Taxanes give similar UV spectra with a minimum at 210–215 nm and a maximum at 225–232 nm, and so most UV analyses are performed at 227–230 nm.80 Accurate quantitative HPLC analysis depends on complete resolution of paclitaxel from other taxanes with similar physicochemical properties; that is, separation can not be considered trivial as some taxanes have similar retention times. The HPLC-MS analysis, although very useful, can by no means be considered conclusive, as the characteristic paclitaxel m/z signals in MS spectra can also be signals of other similar taxanes, such as cephalomannine and 7-epitaxol (Figure 2, ZR Flores-Bustamante, unpublished data). The use of HPLC-tandem-mass spectrometry (HPLC-MS-MS) in multiple reaction-monitoring mode may be of great help to discriminate paclitaxel from other closely related taxanes. Recently, the transitions m/z 567.4 → 445.4, 609.3 → 549.5, 944.9 → 286.4, 812.6 → 286.1, 832.8 → 264.1, 854.4 → 286.1 and 812.6 → 286.1 for 10-deacetyl-baccatin III, baccatin III, 10-deacetyl-7-xylosyltaxol, 10-deacetyltaxol, cephalomannine, paclitaxel and 10-deacetyl-7-epitaxol, respectively, were used to analyze extracts of T. cuspidata, T. x media and T. chinensis var. mairei.81 There are no reports, however, on HPLC-MS-MS applications to identify paclitaxel or taxanes in microbial culture extracts. Other high-resolution techniques, such as FT ion cyclotron resonance multiple stages MS (FT-ICR-MSn), have not been used to detect paclitaxel in microbial cultures, but can be very useful for structural elucidation and to distinguish signals from ions with nearly identical m/z values.82 NMR, although more convincing than the techniques cited above, requires relatively large amounts of purified paclitaxel, difficult to obtain in microbial cultures.
(a) HPLC-ESI-MS chromatogram of taxane standard (Hauser Inc., Boulder, CO, USA) containing 10-deacetylbaccatin III (1), baccatin III (2), 10-deacetyl-7-xylosyltaxol B (3), 10-deacetyl-7-xylosyltaxol (4), taxinine M (5), 10-deacetyl-7-xylosyltaxol C (6), 10-deacetyl-taxol (7), 7-xylosyltaxol (8), cephalomannine (9), 10-deacetyl-7-epitaxol (10), taxol (11), taxol C (12) and 7-epitaxol (13). HPLC-ESI-MS mass spectra showing [M+Na]+ of (b) cephalomannine, (c) paclitaxel and (d) 7-epitaxol. System used Agilent 1100 series LC-MSD Trap equipped with binary pump, degasser, photodiode array detector, column oven, automatic sample injector and an ion-trap mass spectrometer (Agilent, Waldbronn, Germany). Separation performed on a Discovery HS F5 pentafluorophenyl propyl (150 mm × 2.1 mm, particle size 3 μm, Supelco, St Louis, MO, USA) at 21 °C. A linear gradient was used, starting at acetonitrile:ammonium acetate 10 mM (35:65) and changed to (50:50), (50:50), and finally (35:65) at 26, 30 and 32 min, respectively. The flow rate was 0.2 ml min−1 and 0.03 ml of sample solution was injected at 2500 ng ml−1 of each taxane.
Many fungi and some bacteria have been reported in the literature as paclitaxel producers with concentrations ranging from 0.024 to 800 ng ml−1 (Table 1). The multiple evidences given for several reported microorganisms can be considered strong enough to support paclitaxel production. Nevertheless, in some other cases the assigned paclitaxel production is most likely preliminary. More research using HPLC-MS-MS, FTMSn and NMR can help in such cases to establish more conclusive evidence about the capacity of such microorganisms to produce paclitaxel. The high variation in quantitative data also calls for some caution, in particular, as in most cases the highest paclitaxel amounts have been reported with HPLC as the quantifying method.
Methods based on the PCR amplification of paclitaxel biosynthetic sequences (ts, dbat and bapt) as molecular markers have been used recently as an effective tool for primary screening of paclitaxel-producing fungi. The occurrence of the first commitment step to produce paclitaxel in microorganisms can be identified by amplification of ts sequences, whereas amplification of dbat and bapt sequences can be indicative of paclitaxel biosynthetic steps more than 10 enzymatic steps downstream.30, 64 The preceding methods have been very useful, as they are not dependent on the production of paclitaxel in laboratory conditions. However, they point to the presence of only some of the coding sequences required for paclitaxel biosynthesis in the microbial molecular blueprint.
The study of paclitaxel-producing microorganisms holds important potential for basic and applied research. However, the lack of availability of a complete set of genes involved in paclitaxel biosynthesis is at present a limiting factor in both cases. The sequencing and analysis of the relatively small microbial genomes should contribute significantly to expand the number of known paclitaxel biosynthetic genes to set the basis for heterologous production. In addition, as over 400 natural taxanes are presently known, but only few genes involved in taxane biosynthesis have been isolated, an enormous field for discovery lies likely ahead with the study of taxane-producing microorganisms. Because of the above, we can consider that research related to taxane-producing microorganisms is still in its early days.
References
Strobel, G. A., Hess, W. M., Ford, E., Sidhu, R. S. & Yang, X. Taxol from fungal endophytes and the issue of biodiversity. J. Ind. Microbiol. Biotechnol. 17, 417–423 (1996b).
Suffness, M. Overview of paclitaxel research: progress on many fronts. Taxane anticancer agents. ACS Symp. Ser. 583, 1–17 (1995).
Nicolaou, K. C. et al. Total synthesis of taxol. Nature 367, 630–634 (1994).
Holton, R. A. et al. First total synthesis of taxol. J. Am. Chem. Soc. 116, 1597–1598 (1994).
Baloglu, E. & Kingston, D. G. I. The taxane diterpenoids. J. Nat. Prod. 62, 1448–1472 (1999).
Patel, R. Tour de paclitaxel. Biocatalysis for semisynthesis. Annu. Rev. Microbiol. 52, 361–395 (1998).
Frense, D. Mini-review, taxanes: perspectives for biotechnological production. Appl. Microbiol. Biotechnol. 73, 1233–1240 (2006).
Cragg, G. M. & Snader, K. M. Taxol: the supply issue. Cancer Cells 3, 233–235 (1991).
Ketchum, R. E. B., Gibson, D. M., Croteau, R. B. & Shuler, M. L. The kinetics of taxoid accumulation in cell suspension cultures of Taxus following elicitation with methyl jasmonate. Biotechnol. Bioeng. 62, 97–105 (1999).
Zhong, J. J. Plant cell culture for production of paclitaxel and other taxanes. Review. J. Biosci. Bioeng. 94, 591–599 (2002).
Wink, M. et al. Sustainable bioproduction of phytochemicals by plant in vitro cultures: anticancer agents. Gen. Res. 3, 90–100 (2005).
Nguyen, T., Eshraghi, J., Gonyea, G., Ream, R. & Smith, R. Studies on factors influencing stability and recovery of paclitaxel from suspension media and cultures of Taxus cuspidata cv. densiformis by high-performance liquid chromatography. J. Chromatogra. 911, 55–61 (2001).
Stierle, A., Strobel, G. A. & Stierle, D. Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of pacific yew. Science 260, 214–216 (1993).
Strobel, G. A., Stierle, A., Stierle, D. & Hess, W. M. Taxomyces andreanae a proposed new taxon for a bulbilliferous hyphomycete associated with Pacific yew. Mycotaxon 47, 71–80 (1993).
Stierle, A., Strobel, G., Stierle, D., Grothaus, P. & Bignami, G. The search for a taxol-producing microorgansim among the endophytic fungi of the pacific yew, Taxus brevifolia. J. Nat. Prod. LLoydia 58, 1315–1324 (1995).
Strobel, G. Rainforest endophytes and bioactive products. Crit. Rev. Biotechnol. 22, 315–333 (2002).
Strobel, G. et al. Taxol from Pestalotiopsis microspora, an endophytic fungus of Taxus wallichiana. Microbiology 142, 435–440 (1996a).
Shrestha, K., Strobel, G. A., Shrivastava, S. P. & Gewali, M. B. Evidence for paclitaxel from tree new endophytic fungi of Himalayan yew of Nepal. Planta Med. 67, 374–376 (2001).
Bashyal, B., Li, J. Y., Strobel, G. A. & Hess, W. M. Seimatoantlerium nepalense, an endophytic taxol producing coelomycete from Himalayan yew (Taxus wallachiana). Mycotaxon. 72, 33–42 (1999).
Noh, M. J. et al. Isolation of a novel microorganism, Pestalotia heterocornis, producing paclitaxel. Biotechnol. Bioeng. 64, 620–623 (1999).
Caruso, M. et al. Isolation of endophytic fungi and actinomycetes taxane producers. Annals Microbiol. 50, 3–13 (2000b).
Wang, J. F. et al. Taxol from Tubercularia sp. strain TF5, an endophytic fungus of Taxus mairei. FEMS Microbiol. Lett. 193, 249–253 (2000).
Wan, B., Li, A. M. & Wang, X. L. Separation of a fungus producing taxol. Sci. China Ser C. 44, 156–160 (2001).
Zhou, D. et al. Study on the mutagenesis of protoplasts from taxol-producing fungus Nodulisporium sylviforme. The J. Am. Sci. 1, 55–62 (2005).
Zhao, K., Zhou, D. P., Ping, W. X. & Jingping, G. Study on the preparation and regeneration of protoplast from taxol-producing fungus Nodulisporium sylviforme. Nat. Sci. 2, 52–59 (2004).
Deng, B. W., Liu, K. H., Chen, W. Q., Ding, X. W. & Xie, X. C. Fusarium solani, Tax-3, a new endophytic taxol-producing fungus from Taxus chinensis. World J. Microbiol. Biotechnol. 25, 139–143 (2009).
Zhou, X. et al. Screening of taxol-producing endophytic fungi from Taxus chinensis var. mairei. Appl. Biochem. Microbiol. 43, 439–443 (2007).
Miao, Z., Wang, Y., Yu, X., Guo, A. & Tang, K. New endophytic taxane production fungus from Taxus chinensi. Appl. Biochem. Microbiol. 45, 81–86 (2009).
Kumaran, R. S. & Hur, B. K. Screening of species of the endophytic fungus Phomopsis for the production of the anticancer drug taxol. Biotechnol. Appl. Biochem. 54, 21–30 (2009).
Zhang, P., Zhou, P. P., Jiang, C., Yu, H. & Yu, L. J. Screening of taxol-producing fungi based on PCR amplification from Taxus. Biotechnol. Lett. 30, 2119–2123 (2008).
Zhang, P., Zhou, P. P. & Yu, L. J. An endophytic Taxol-producing fungus from Taxus x media, Cladosporium cladosporioides MD2. Curr. Microbiol. 59, 227–232 (2009a).
Zhang, P., Zhou, P. P. & Yu, L. J. An edophytic Taxol-producing fungus from Taxus x media, Aspergillus candidus MD3. FEMS Microbiol. Lett. 293, 55–159 (2009b).
Li, J. Y., Strobel, G., Sidhu, R., Hess, W. M. & Ford, E. J. Endophytic taxol producing fungi from bald cypress Taxodium distichum. Microbiology 142, 2223–2226 (1996).
Strobel, G. A. et al. Pestalotiopsis guepinii, a taxol producing endophyte of the Wollemi Pine, Wollemia nobilis. Aust. J. Bot. 4586, 1073–1082 (1997).
Hoffman, A. et al. Bioprospecting for taxol in angiosperm plant extracts—using high performance liquid chromatography thermospray mass spectroscopy to detect the anticancer agent and its related metabolites in Filbert trees. Spectroscopy 13, 22–32 (1998).
Li, J. Y. et al. The induction of taxol production in the endophytic fungus Periconia sp. from Torreya grandifolia. J. Ind. Microbiol. Biotechnol. 20, 259–264 (1998b).
Strobel, G. A. et al. Seimatoantlerium tepuiense gen. nov., a unique epiphytic fungus producing taxol from the Venezuelan-Guayana. System Appl. Microbiol. 22, 426–433 (1999).
Strobel, G. et al. Stegolerium kukenani gen, et sp nov., an endophytic taxol producing fungus from the Roraima and Kukenan tepuis of Venezuela. Mycotaxon 78, 353–361 (2001).
Gangadevi, V. & Muthumary, J. Taxol production by Pestalotiopsis terminaliae, an endophytic fungus of Terminalia arjuna (arjun tree). Biotech. Appl. Biochem. 52, 9–15 (2009b).
Gangadevi, V. & Muthumary, J. Taxol, an anticancer drug produced by an endophytic fungus Bartalinia robillardoides Tassi, isolated from a medicinal plant, Aegle marmelos Correa ex Roxb. World J. Microbiol. Biotechnol. 24, 717–724 (2008).
Kumaran, R. S., Muthumary, J. & Hur, B. K. Isolation and identification of an anticancer drug, taxol from Phyllosticta tabernaemontanae, a leaf spot fungus of an angiosperm, Wrightia tinctoria. J. Microbiol. 47, 40–49 (2009a).
Kumaran, R. S., Muthumary, J. & Hur, B. K. Production of Taxol from Phyllosticta spinarum, an endophytic fungus of Cupressus sp. Eng. Life Sci. 8, 438–446 (2008b).
Kumaran, R. S., Muthumary, J. & Hur, B. K. Taxol from Phyllosticta citricarpa, a leaf spot fungus of the angiosperm Citrus medica. J. Biosci. Bioeng. 106, 103–106 (2008c).
Kumaran, R., Muthumary, J., Kim, E. K. & Hur, B. K. Production of taxol from Phyllosticta dioscoreae, a leaf spot fungus isolated from Hibiscus rosa-sinensis. Biotechnol. Bioprocess Eng. 14, 76–83 (2009b).
Page, M. & Landry, N. Bacterial mass production of taxanes with Erwinia. U.S. 5,561,055 (1996).
Page, M. et al. Bacterial mass production of taxanes and paclitaxel. U.S. 6,030,818 (2000).
Caruso, M. et al. Studies on a strain of Kitasatospora sp. paclitaxel producer. Annals. Microbiol. 50, 89–102 (2000a).
Li, J. Y., Sidhu, R. S., Bollon, A. & Strobel, G. A. Stimulation of taxol production in liquid cultures of Pestalotiopsis microspora. Mycol. Res. 102, 461–464 (1998a).
Hezari, M. & Croteau, R. Taxol biosynthesis: an update. Planta Med. 63, 291–295 (1997).
Jennewein, S. & Croteau, R. Taxol biosynthesis: molecular genetics, and biotechnological applications. Appl. Microbiol. Biotechnol. 57, 13–19 (2001).
Walker, K. & Croteau, R. Taxol biosynthetic genes. Phytochemistry 58, 1–7 (2001).
Hezari, M., Lewis, N. G. & Croteau, R. Purification and characterization of taxa-4(5),11(12)-diene synthase from Pacific yew (Taxus brevifolia) that catalyzes the first committed step of taxol biosynthesis. Arch. Biochem. Biophys. 322, 437–444 (1995).
DeJong, J. M. et al. Genetic engineering of Taxol biosynthetic genes in Saccharomyces cerevisiae. Biotechnol. Bioeng. 93, 212–224 (2006).
Walker, K., Long, R. & Croteau, R. The final acylation step in Taxol biosynthesis: cloning of the taxoid C13-side-chain N-benzoyltransferase from Taxus. Proc. Natl Acad. Sci. (USA) 99, 9166–9171 (2001).
Croteau, R. B., Ketchum, R. E. B., Long, R., Kaspera, R. & Wildung, M. Taxol biosynthesis and molecular genetics. Phytochem. Rev. 5, 75–97 (2006).
Jennewein, S., Rithner, C. D., Williams, R. M. & Croteau, R. B. Taxol biosynthesis: taxane 13 alpha-hydroxylase is a cytochrome P450-dependent monooxygenase. Proc. Natl Acad. Sci. (USA) 98, 13595–13600 (2001).
Schoendorf, A., Rithner, C. D., Williams, R. M. & Croteau, R. B. Molecular cloning of a cytochrome P450 taxane 10 beta-hydroxylase cDNA from Taxus and functional expression in yeast. Proc. Natl Acad. Sci. USA 98, 1501–1506 (2001).
Staniek, A., Woerdenbag, H. J. & Kayser, O. Taxomyces andreanae: a presumed paclitaxel producer demystified? Planta Med. 75, 1561–1566 (2009).
Zhao, K. et al. Aspergillus niger var. taxi, a new species variant of taxol-producing fungus isolated from Taxus cuspidata in China. J. Appl. Microbiol. 107, 1202–1207 (2009).
Withers, S. T. & Keasling, J. D. Biosynthesis and engineering of isoprenoid small molecules. Appl. Microbiol. Biotechnol. 73, 980–990 (2007).
Wildung, M. R. & Croteau, R. A cDNA clone for taxadiene synthase, the diterpene cyclase that catalyzes the committed step of taxol biosynthesis. J. Biol. Chem. 271, 9201–9204 (1996).
Walker, K., Schoendorf, A. & Croteau, R. Molecular cloning of a taxa-4(20), 11(12)-dien-5 alpha-ol-O-acetyl transferase cDNA from Taxus and functional expression in Escherichia coli. Arch. Biochem. Biophys. 374, 371–380 (2000).
Walker, K., Fujisaki, S., Long, R. & Croteau, R. Molecular cloning and heterologous expression of the C-13 phenylpropanoid side chain-CoA acyltransferase that functions in taxol biosynthesis. Proc. Natl Acad. Sci. (USA) 99, 12715–12720 (2002).
Jennewein, S., Wildung, M. R., Chau, M., Walker, K. & Croteau, R. Random sequencing of an induced Taxus cell cDNA library for identification of clones involved in taxol biosynthesis. Proc. Natl Acad. Sci. (USA) 101, 9149–9154 (2004a).
Jennewein, S., Long, R. M., Williams, R. M. & Croteau, R. Cytochrome P450 taxadiene 5a-hydroxylase, a mechanistically unusual monooxygenase catalyzing the first oxygenation step of taxol biosynthesis. Chem. Biol. 11, 379–387 (2004b).
Walker, K. & Croteau, R. Molecular cloning of a 10-deacetylbaccatin III-10-O-acetyl transferase cDNA from Taxus and functional expression in Escherichia coli. Proc. Natl Acad. Sci. (USA) 97, 583–587 (2000a).
Walker, K. & Croteau, R. Taxol biosynthesis: molecular cloning of a benzoyl-CoA: taxane 2 alpha-O-benzoyltransferase cDNA from Taxus and functional expression in Escherichia coli. Proc. Natl Acad. Sci. (USA) 97, 13591–13596 (2000b).
Williams, D. C. et al. Heterologous expression and characterization of a ‘pseudomature’ form of taxadiene synthase involved in paclitaxel (Taxol) biosynthesis and evaluation of a potential intermediate and inhibitors of the multistep diterpene cyclization reaction. Arch. Biochem. Biophys. 379, 137–146 (2000).
Hahn, F. M. & Poulter, C. D. Isolation of Schizosaccharomyces pombe isopentenyl diphosphate isomerase cDNA clones by complementation and synthesis of the enzyme in Escherichia coli. J. Biol. Chem. 270, 11298–11303 (1995).
Hefner, J., Ketchum, R. E. B. & Croteau, R. Cloning and functional expression of a cDNA encoding geranylgeranyl diphosphate synthase from Taxus canadensis and assessment of the role of this prenyltransferase in cells induced for Taxol production. Arch. Biochem. Biophys. 360, 62–74 (1998).
Huang, K. X., Huang, Q. L., Wildung, M. R., Croteau, R. & Scott, A. I. Overproduction, in Escherichia coli, of soluble taxadiene synthase, a key enzyme in the Taxol biosynthetic pathway. Protein Expr. Purif. 13, 90–96 (1998).
Huang, Q. L., Roessner, C. A., Croteau, R. & Scott, A. I. Engineering Escherichia coli for the synthesis of taxadiene, a key intermediate in the biosynthesis of taxol. Bioorganic Med. Chem. 9, 2237–2242 (2001).
Engels, B., Dahm, P. & Jennewein, S. Metabolic engineering of taxadiene biosynthesis in yeast as a first step towards Taxol (paclitaxel) production. Metabolic Eng. 10, 201–206 (2008).
Muralidhar, R. V. & Panda, T. Fungal protoplast fusion—a revisit. Bioproc. Biosyt. Eng. 22, 429–431 (2000).
Zhao, K., Ping, W. X. Ma, X. Liu, J. & Zhou, D. P. Breeding of high-yield strain of taxol by mutagenesis of protoplast and primary discussion of genetic differences between mutants and their parent strain. Acta. Microbiol. Sin. 45, 355–358 (2005).
Zhao, K. et al. Screening and breeding of high taxol producing fungi by genome suffling. Sci. China Ser. C-Life Sci. 5, 222–231 (2008).
Zhou, D. P., Sun, J. Q., Yu, H. Y., Zheng, X. Q. & Ping, W. X. Nodulisporium, a genus new to China. Mycosystema 20, 277–278 (2001).
Williams, R. B., Henrikson, J. C., Hoover, A. R., Lee, A. E. & Cichewicz, R. H. Epigenetic remodeling of the fungal secondary metabolome. Org. Biomol. Chem. 6, 1895–1897 (2008).
Grothaus, P. G., Raybould, T. J. G., Bignami, G. S., Lazo, C. B. & Byrnes, J. B. An enzyme immunosassay for the determination of taxol and taxanes in Taxus sp. tissues and human plasma. J. Immunol. Meth. 158, 5–15 (1993).
Theodoridis, G. & Verpoorte, R. Taxol analysis by high performance liquid chromatography: a review. Phytochem. Anal. 7, 169–184 (1996).
Li, S. et al. Determination of paclitaxel and other six taxoids in Taxus species by high-performance liquid chromatography–tandem mass spectrometry. J. Pharma. Biomed. Anal. 49, 81–89 (2009).
McDonald, L. A. et al. FTMS structure elucidation of natural products: application to muraymycin antibiotics using ESI multi-CHEF SORI-CID FTMSn, the top-down/bottom-up approach, and HPLC ESI capillary-skimmer CID FTMS. Anal. Chem. 75, 2730–2739 (2003).
Guo, B. H. et al. An endophytic taxol-producing fungus BT2 isolated from Taxus chiniensis var. mairei. African J. Biotechnol. 5, 875–877 (2006).
Xu, F., Tao, W., Cheng, L. & Guo, L. Strain improvement and optimization of the media of taxol-producing fungus Fusarium maire. Biochem. Eng. J. 31, 67–73 (2006).
Chi, Y., Zhao, D. L. & Zhou, D. P. Identification of taxol biosynthesis stage-enriched transcripts in Nodulisporium sylviforme, using suppression subtractive hybridization. World J. Microbiol. Biotechnol. 24, 2601–2605 (2008).
Kumaran, R. S., Muthumary, J. & Hur, B. K. Isolation and identification of taxol, an anticancer drug from Phyllosticta melochiae Yates, an endophytic fungus of Melochia corchorifolia L. Food Sci. Biotechnol. 17, 1246–1253 (2008a).
Gangadevi, V. & Muthumary, J. A novel endophytic Taxol-producing fungus Chaetomella raphigera isolated from a medicinal plant, Terminalia arjuna. Appl. Biochem. Biotechnol. 158, 675–684 (2009a).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Acknowledgements
We want to express our appreciation to Dr Demain for his stimulating and outstanding scientific contributions, and for his kindness and human qualities. Also, we thank the editors of this special issue for inviting us to participate in celebrating Dr Demain's 60 years of distinguished career as scientist and educator. Funding was provided by Semarnat-Conacyt (Project 0368), Fomix-Conacyt-Hidalgo (project 2006-01-49004) and ICyTDF (project PIFUTP08-90). AM-C received grant-aided support from CONACYT.
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper is contributed to the special issue of the Journal of Antibiotics celebrating Dr Arnold Demain's 60 years of scientific career.
Rights and permissions
About this article
Cite this article
Flores-Bustamante, Z., Rivera-Orduña, F., Martínez-Cárdenas, A. et al. Microbial paclitaxel: advances and perspectives. J Antibiot 63, 460–467 (2010). https://doi.org/10.1038/ja.2010.83
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2010.83