Abstract
Protoplast preparation, regeneration and fusion represent essential tools for those poorly studied biotechnologically valuable microorganisms inapplicable with the current molecular biology protocols. The protoplast production and regeneration method developed for Planobispora rosea and using the combination of hen egg-white lysozyme (HEWL) and Streptomyces globisporus mutanolysin was applied to a set of antibiotic-producing filamentous actinomycetes belonging to the Streptosporangiaceae, Micromonosporaceae and Streptomycetaceae. 107–109 protoplasts were obtained from 100 ml of culture, after incubation times in the digestion solution ranging from a few hours to 1 or 2 days depending on the strain. The efficiency of protoplast reversion to the normal filamentous growth varied from 0.1 to nearly 50%. Analysis of cell wall peptidoglycan in three representative strains (Nonomuraea sp. ATCC 39727, Actinoplanes teichomyceticus ATCC 31121 and Streptomyces coelicolor A3(2)) has evidenced structural variations in the glycan strand and in the peptide chain, which may account for the different response to cell digestion and protoplast regeneration treatments.
Similar content being viewed by others
Introduction
The so-called rare or uncommon actinomycetes include a group of filamentous actinomycetes other than Streptomyces spp., which are quite difficult to isolate, cultivate and genetically manipulate.1 Members of these poorly represented and difficult-to-handle group of microorganisms produce valuable antibiotics. However, the study and the following cost-effective exploitation of uncommon actinomycetes have been slow because of the lack of genetic tools, which hamper strain and product improvement. As mobile genetic elements and conjugation systems are poorly characterized in rare actinomycetes, a crucial methodology to achieve their transformation by exogenous DNA or to recombine whole genomes arising from different cell lines (Whole Genome Shuffling-WGS) is based on protoplast manipulation and fusion. Protoplast preparation and regeneration in Streptomyces spp. were originally reported by Okanishi and co-workers.2 The method developed for streptomycetes was then applied with uneven success to Micromonospora spp.3 and Brevibacillus spp.,4 but it soon became clear that the protocol used later was species- or even strain-specific. In other industrially valuable actinomycetes, ad hoc techniques have been developed in some cases,5, 6, 7, 8 but methods endowed with a broad applicability and robustness to be applied for DNA recombination and WGS are not available. A possible explanation for the different response to protocols of protoplast preparation and regeneration from taxonomically related species of filamentous actinomycetes should be searched for in the interspecific variation of their cell wall composition. Differences in the peptidoglycan composition and structure indeed have taxonomical implications and represent one of the chemotaxonomical traits on the basis of the polyphasic classification of actinomycetes.9, 10 Variation in the fine structure (composition and density) of peptidoglycan in response to growth and environmental conditions, aging, maturation and fine-tuning of bacterial cell wall structure to best suit changing conditions may further explain the observed intraspecific variability in susceptibility to cell wall digestion.11
We have recently applied a tailor-made methodology of cell wall digestion and regeneration to the uncommon actinomycete Planobispora rosea, with the aim of improving its production of the thiazolyl peptide antibiotic GE2270 by genome shuffling.12 In this paper, we have extended the use of this approach to actinomycete species, which produce valuable antibiotics already in clinical use or under development (teicoplanin,13 A40926-precursor of the semisynthetic dalbavancin-,14, 15 ramoplanin16 enduracidin17, 18) or potentially interesting molecules to be used per se or as scaffold for new drugs19, 20 (Table 1). The different response to protoplast treatment has been correlated with the diversity of peptidoglycan precursors in representative strains.
Materials and methods
Strains and cultural conditions
Strains listed in Table 1 were obtained from ATCC and NRRL public culture collections. They were maintained as a lyophilized Master Cell Bank (MCB). A Working Cell Bank (WCB) was prepared from the first generation slant originating from the MCB as described.21 Cryo-vials from the WCB were thawed at room temperature, and 2 ml were used to inoculate 100 ml of the growth medium (specified for each strain in Table 1) in 500-ml baffled flasks containing 20–30 glass beads. One percent agar was added to the growth medium as suggested by Hobbs et al.22 to obtain better dispersed growth. For protoplast preparation, 10% of the culture grown for 24–48 h at 28 °C and 200 r.p.m. was inoculated in 100 ml of the protoplast preparation medium (specified for each strain in Table 1), and growth was allowed for a further 24–48 h at the same temperature and agitation conditions. The composition of liquid media is described in Table 2.
Protoplast preparation
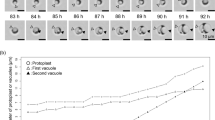
Protoplasts were prepared essentially as described for P. rosea.12 In brief, ca. 100 ml of each grown culture were centrifuged at 3250 g. The mycelium was washed once in P medium, and 10 g (fresh weight) were suspended in 50 ml of P medium. For cell wall digestion, hen egg-white lysozyme (HEWL) (Sigma-Aldrich, St Louis, MO, USA) and Streptomyces globisporus mutanolysin (Sigma-Aldrich) were dissolved in P medium at the final concentrations of 5 and 0.018 mg ml–1, respectively. The non-ionic detergent Pluronic (Sigma-Aldrich) was added at the final concentration of 100 mg l–1. Incubation with the digestion solution was at 28 °C with gentle shaking at 50 r.p.m., and lasted from 4 to 48 h, depending on the strain. Protoplasts were then detached from residual mycelium clumps by thoroughly pipetting up and down and separated from residual hyphal fragments by filtration through glass wool. Filtration through 5 or 1.2 μm durapore membrane filters (Millipore, Billerica, MA, USA) was further applied when required. Protoplasts were then centrifuged at 30 000 g and finally resuspended in fresh P medium. The formation of protoplasts was monitored by microscopic observation (Zeiss phase-contrast microscope at × 400, Carl Zeiss S.p.A., Arese, Italy). Total protoplast number was determined by using a Petroff–Hausser counting chamber.
Regeneration of mycelium from protoplasts
Regeneration of mycelium from protoplasts was performed using the overlay technique suggested by Shirahama et al.23 Plates were seeded by pouring 0.2 ml of protoplast suspension on hypertonic media (M3 or M103 or MP3), and the upper layer was medium VMS0.1.12 To assess residual hyphal contamination of protoplast suspensions, control plates, with medium V0.1 as the under layer and medium VM0.1 as the upper layer, were seeded. In these media devoid of sucrose, hyphal cells but not protoplasts were able to grow. Composition of solid media is described in Table 3.
Peptidoglycan precursor extraction and analysis
Peptidoglycan precursors were extracted from the mycelium of Actinoplanes teichomyceticus ATCC 31121, Nonomuraea sp. ATCC 39727 and Streptomyces coelicolor A3(2) by a method previously described.24 In brief, mycelium, grown in vegetative medium to the exponential phase, was incubated for 1 h with ramoplanin (approximately eightfold the ramoplanin MIC value for each strain) to block cell wall final assembly and thus to amplify the intracellular pool of uridinediphospho-linked precursors.24 Then, cells were harvested, suspended in water (0.1 g fresh weight per ml) and boiled for 20 min. After cooling first at room temperature and then in ice, the mycelium was centrifuged at 39 000 g. The supernatant was lyophilized and dissolved in 0.1 volume of water adjusted to pH 3 with formic acid. The samples were analyzed by reversed-phase HPLC and ESI-MS on an LCQ-Deca spectrometer equipped with an ion trap analyzer (Thermo Finnigan, San Josè, CA, USA) as described in Beltrametti.24
Results
Growth of the strains and protoplast production
As we have reported for P. rosea,12 many filamentous actinomycetes grow in liquid media as tough pellets consisting of filamentous hyphae. As a consequence of this peculiar growth, cell wall is poorly accessible to enzymatic hydrolysis, and mycelium is scarcely converted into protoplasts. We have observed that the first necessary condition for good protoplast production is to obtain a good dispersed growth of a strain. Therefore, media and growth conditions applied to P. rosea were tested with few modifications on the strains listed in Table 1. Asa rule, we had first screened media known to support biomass production of our strains (data not shown) added with 1 g l–1 agar, and then we supplemented the best medium for each strain with appropriate concentrations of sucrose and proline (the latter acting as an osmo-protectant25). The addition of sucrose and proline favored adaptation to the components of the hypertonic buffer (P medium) used for the subsequent cell wall digestion and for protoplast production.26, 27 Table 1 describes the selected strains, their antibiotic products and the media developed for their liquid growth before cell wall digestion and for protoplast production. As observed for P. rosea, subcultivation from the starch-based medium V (Table 2) to its variant containing 103 g l–1 of sucrose and 3.5 g l–1 of proline (protoplast production medium VM, Table 2) gave the best compromise between biomass production and protoplast yield in A. teichomyceticus and S. fungicidicus (Table 1). For most of the other strains, the addition of low amounts of proline and sucrose directly into the growth medium (medium VSP, Table 2) further improved growth and facilitated acclimatization to hypertonic conditions. In Actinoplanes sp. ATCC 33076, producing ramoplanin, the best medium to support biomass and protoplast production was devoid of starch and supplemented with additional amino acid sources (V6, and its sucrose and proline added version V6SP, Table 2).28
Classical procedures of protoplast preparation in streptomycetes are based on cell wall digestion by the commercially available HEWL, which is an N-acetyl-β-D-muramidase that cleaves the β-1,4-glycosidic bond between the amino sugars forming the glycan strands.29 When HEWL was used as the unique enzyme component of the digestion solution for our strains, protoplasts were efficiently formed only in Streptomyces fungicidicus (Table 1) and Nonomuraea sp. ATCC39727 (data not shown). The addition of mutanolysin (another muramidase, produced by Streptomyces globisporus) was instead a determinant for protoplast production in the rest of the strains belonging to non-Streptomyces actinomycete genera (Table 1).
The efficiency of protoplast formation was assayed by microscopic enumeration at different times of incubation in the digestion solution. In contrast to the fast protoplast formation observed in Streptomyces coelicolor (∼15 min),29 maximum protoplast yield (108/109 protoplasts from 100 ml of culture) was achieved after incubation times ranging from a few hours in the case of S. fungicidicus to 1 or 2 days for most of the non-Streptomyces actinomycetes belonging to the Streptosporangiaceae. Even after prolonging the incubation times with digestion solution, a 10-fold lower protoplast yield was generally achieved in Actinoplanes spp. (the Micromonosporaceae). Moreover, the obtained protoplasts from Actinoplanes spp. appeared to be extensively contaminated by short hyphal fragments. Thus, a filtration step using 1.2 μm membrane filters was introduced in their preparation. A high degree of resistance to digestion solution in Actinoplanes spp. may be explained by their modified peptidoglycan composition (see experiment reported hereafter).
Regeneration of mycelium from protoplasts
In P. rosea,12 we have shown the importance of designing an appropriate regeneration medium. Indeed, a significant increase (from 0.1 to 30%) of its protoplast regeneration efficiency was achieved by the proper balance of sucrose, proline and microelements. In an initial approach, protoplast regeneration of the antibiotic-producing actinomycetes listed in Table 1 was tested in the starch-based regeneration medium M3 developed for P. rosea (see Table 3 for medium composition). However, results obtained on M3 were satisfactory only for P. rosea ATCC 53773 and Nonomuraea sp. ATCC 39727. Therefore, we reasoned that many of the selected strains produce antibiotics (teicoplanin, A40926, ramoplanin, enduracidin and lantibiotics) whose mechanism of action relies on the inhibition of cell wall synthesis. Traces of these molecules in the producer cells may contribute to hamper cell wall regeneration. To improve regeneration, medium M3 was first enriched in sucrose, yielding medium M103 (regeneration data equivalent to those in M3, not reported), and then added with relatively high concentrations of glucose and potato starch (medium MP3, Table 3), with the intent to inhibit antibiotic formation by catabolite repression.30 In most of the strains except in M. corallina NRRL 30420, the regeneration efficiency in M3 was indeed nearly identical to the ones achieved in MP3 or M103, and ranged from 0.1 to nearly 50% depending on the strain (Table 1).
For Actinoplanes spp., the high number of colonies grown in non-permissive conditions (medium V0.1, Table 3) confirmed that the protoplast solutions were still heavily contaminated by residual hyphae. Furthermore, two types of their colonies became detectable on the regeneration plates: the faster-growing colonies, presumably originated from residual hyphae, inhibited the growth of late-stage colonies (small slow-growing colonies) arising from protoplast regeneration (data not shown), as previously reported for some Streptomyces strains.31
In S. fungicidicus, protoplasts prepared by the digestion solution containing mutanolysin failed to regenerate, but regenerated scarcely if only lysozyme was used. This was probably because of over-digestion of the cell wall, in the presence of mutanolysin, which likely acts in streptomycetes as an endogenous autolysin.10
Determination of the structure of peptidoglycan precursors
The difference in ease of protoplast formation between members of the Actinoplanes genus and those of the Streptomyces genus and/or the Streptosporangiaceae was particularly evident in our experimental conditions. We reasoned that the differential response of actinomycetes of the Streptomycetaceae, Streptosporangiaceae and Micromonosporaceae to the treatments of either the digestion or the regeneration of their cell walls might be explained by investigating their cell wall structure. In a previous study,24 we reported in A. teichomyceticus the presence of cell wall precursors with structure UDP-N-glycolylmuramyl-Gly-D-Glu-mDap-D-Ala-D-Lac and UDP-N-glycolylmuramyl-Gly-D-Glu-mDap-D-Ala-D-Lac hydroxylated on the meso-Diaminopimelic acid. Previous findings also indicated the coexistence of these two peptidoglycan precursors in other Actinoplanes species and in Micromomospora species.32 For the model actinomycete S. coelicolor, Hong et al.33 reported instead a cell wall precursor with structure UDP-N-acetylmuramyl-L-Ala-D-Glu-LL-Dap-D-Ala-D-Ala. These basic observations strengthened the hypothesis that the difference observed in protoplast formation among our strains could be attributed to differences in the cell wall structure and precursors. To further get an insight into the possible structural relationship among strains displaying similar behavior in protoplast formation, we took advantage of an LC–MS method previously used in our laboratories24 for the analysis of cell wall cytoplasmatic precursors. Among the strains reported in this study, only Nonomuraea sp. ATCC 39727 is comparable in mycelium growth and ease of protoplast formation to S. coelicolor A3(2) (data not shown). Therefore, we analyzed Nonomuraea sp. ATCC 39727 with the aim of identifying similarities to the peptidoglycan precursor of S. coelicolor. In Nonomuraea sp. ATCC 39727, cell wall assembly is blocked by the addition of ramoplanin, which inhibits peptidoglycan transglycosylases and leads to an accumulation of cell wall precursors (which are undetectable in untreated cells because of their rapid assembly into the peptidoglycan).34 LC–MS was performed in Nonomuraea sp. ATCC 39727 on the ramoplanin-treated and -untreated cells, and peaks emerging in ramoplanin-treated cells were investigated. The comparison of LC–MS analyses between untreated (Figure 1a) and ramoplanin-treated (Figure 1b) cells showed in the latter case the occurrence of peaks eluting at 12.56 and 13.40 min. In correspondence of the peak eluting at 13.40 min, full scan mass spectrum in negative ion current showed quasi-molecular ions [M-H]−, and [M-2H]−2 at m/z 1192.3 and 595.7, respectively, corresponding to a calculated monoisotopic mass of 1193 (data not shown). This monoisotopic mass was consistent with the presence of a pentapeptide precursor with structure UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala-D-Ala, identical to the precursor described above for S. coelicolor33 except for the probable presence of meso-Diaminopimelic acid (as suggested by Goodfellow and Quintana35) instead of LL-Diaminopimelic acid. MS–MS analysis confirmed this identity (data not shown). Interestingly, full scan mass spectrum in negative and positive ion current of the peak eluting at 12.56 min (Figure 1b) showed quasi-molecular ions [M-H]−, [M-H]+ and [M-2H]−2 at m/z 1121.1, 1123.0 and 560.3, respectively, corresponding to a calculated monoisotopic mass of 1122 (Figures 1c and d). The structure attributable to the accumulated precursor was UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala and corresponded to the UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala-D-Ala depleted of the terminal D-Alanine (tetrapeptide peptidoglycan precursor). Additional evidence for the structure UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala was directly provided by the formation in the MS source of ions at m/z 719.2 (positive mode) corresponding to the loss of UDP and yielding N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala, and 403.0 (negative mode) corresponding to UDP. MS data are summarized in Figure 1e. It is worth noting that in the LC–MS chromatogram (Figure 1b) the peak corresponding to the tetrapeptide precursor was predominant on the pentapeptide.
Analysis of the peptidoglycan precursor of Nonomuraea sp. ATCC 39727. LC–MS analysis of ramoplanin-treated (b) cells showed the occurrence of peaks eluting at 12.56 and 13.40 min not detected in untreated cells (a). Full scan mass spectrum of the peak eluting at 13.40 min was consistent with the presence of a pentapeptide precursor with structure UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala-D-Ala (e). Full scan mass spectrum in positive (c) and negative (d) ion current of the peak eluting at 12.56 min showed quasi-molecular ions [M-H]+, [M-H]− and [M-2H]−2 at m/z 1123.0, 1121.1 and 560.3, respectively, corresponding to a calculated monoisotopic mass of 1122. The structure attributed to this precursor was UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala and corresponded to the tetrapeptide peptidoglycan precursor (e). The ion at m/z 403.0 corresponded to UDP formed directly in the MS source (d). The ion at m/z 719.2, formed directly in the MS source, corresponded to the loss of UDP from the UDP-N-acetylmuramyl-L-Ala-D-Glu-mDap-D-Ala (c). The masses corresponding to the observed fragment ions are reported above the arrows in (e).
Discussion
We have applied the technology developed for protoplast production in P. rosea to different actinomycete strains selected among producers of antibiotics of industrial interest. We have verified that the method works with the selected strains, even if different incubation times are required. As a rule, obtaining a good dispersed growth for each strain has been crucial for further cell wall digestion and protoplast formation. Standardizing mycelium dispersion (thus rendering the bacterial cell wall physically accessible to digestion enzymes) and determining the starting mycelium amount will allow better results. On the other hand, we observed that regeneration occurred with uneven efficiencies indicating that fine-tuning of the regeneration medium composition and the right degree of cell wall digestion are key factors for reversion to the normal mycelial growth.
On a molecular basis, our data suggested that cell wall precursors could influence the ease of protoplast formation. Our results with strains belonging to the Streptosporangiaceae show that the addition of mutanolysin (an autolysin produced by Streptomyces globisporus) enhances the digestion of their cell walls. A major difference of resistance to the digestion treatment in Actinoplanes spp. (Micromonosporaceae) can be explained by their modified peptidoglycan composition. Lysozymes are produced by phages, bacteria, fungi, invertebrates and vertebrates, and they differ in structure as well as in the specificity of their activity for a certain peptidoglycan type, for the presence or absence of secondary modifications on the glycan strands, or for either high-molecular weight or small fragments. Bacterial lysozymes, which usually act as autolysins, are ubiquitous, but only a few of them are well characterized with an experimentally proven specificity.10
It has been reported that modifications of the peptidoglycan such as N-deacetylation and O-acetylation of glycan strands in some bacterial species render them resistant to the hydrolytic activity of lysozymes. In Actinoplanes, the presence of a glycolyl residue (instead of acetate) on the amino group in the peptidoglycan precursor (N-glycolylation), could have a similar role in protection against lysozymes as have N-deacetylation and O-acetylation in other species.
Increased availability of lysozymes from different biological sources will allow in the future the targeted selection of the cell wall digesting enzyme for protoplast production in recalcitrant actinomycete strains.
Our work is going on to investigate the occurrence of tetrapeptide peptidoglycan precursor accumulation in Nonomuraea sp. and its role in cell wall biosynthesis, modification and resistance (Beltrametti F, unpublished results).
References
Lazzarini, A., Cavaletti, L., Toppo, G. & Marinelli, F. Rare genera of actinomycetes as potential producers of new antibiotics. Antonie Van Leeuwenhoek 79, 399–405 (2001).
Baltz, R. H. Genetic recombination in Streptomyces fradiae by protoplast fusion and cell regeneration. J. Gen. Microbiol. 107, 93–102 (1978).
Ogawa, H., Imai, S., Satoh, A. & Kojima, M. An improved method for the preparation of streptomycetes and Micromonospora protoplasts. J. Antibiot. 2, 184–186 (1983).
Hopwood, D. A. Genetic studies with bacterial protoplasts. Annu. Rev. Microbiol. 35, 237–272 (1981).
Stegmann, E., Pelzer, S., Wilken, K. & Wohlleben, W. Development of three different gene cloning systems for genetic investigation of the new species Amycolatopsis japonicum MG417-CF17, the ethylenediaminedisuccinic acid producer. J. Biotechnol. 92, 195–204 (2001).
Matsushima, P., McHenney, M. A. & Baltz, R. H. Efficient transformation of Amycolatopsis orientalis (Nocardia orientalis) protoplasts by Streptomyces plasmids. J. Bacteriol. 169, 2298–2300 (1987).
Rajnisz, A., Solecka, J. & Kurzatkowski, W. Properties of Saccharopolyspora erythraea strains after protoplast regeneration. Folia Microbiol. (Praha) 50, 13–18 (2005).
Palleroni, N. J. Genetic recombination in Actinoplanes brasiliensis by protoplast fusion. Appl. Environ. Microbiol. 45, 1865–1869 (1983).
Anderson, A. S. & Wellington, E. M. The taxonomy of Streptomyces and related genera. Int. J. Syst. Evol. Microbiol. 51, 797–814 (2001).
Vollmer, W., Blanot, D. & de Pedro, M. A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Vollmer, W. Structural variation in the glycan strands of bacterial peptidoglycan. FEMS Microbiol. Rev. 32, 287–306 (2008).
Beltrametti, F., Barucco, D., Rossi, R., Selva, E. & Marinelli, F. Protoplast fusion and gene recombination in the uncommon actinomycete Planobispora rosea producing GE2270. J. Antibiot. 60, 447–454 (2007).
Parenti, F., Beretta, G., Berti, M. & Arioli, V. Teichomycins, new antibiotics from Actinoplanes teichomyceticus Nov. Sp. I. Description of the producer strain, fermentation studies and biological properties. J. Antibiot. 31, 276–283 (1978).
Goldstein, B. P. et al. A40926, a new glycopeptide antibiotic with anti-Neisseria activity. Antimicrob. Agents Chemother. 31, 1961–1966 (1987).
Malabarba, A. & Goldstein, B. P. Origin, structure, and activity in vitro and in vivo of dalbavancin. J. Antimicrob. Chemother. 55, 15–20 (2005).
Parenti, F., Ciabatti, R., Cavalleri, B. & Kettenring, J. Ramoplanin: a review of its discovery and its chemistry. Drugs Exp. Clin. Res. 16, 451–455 (1990).
Yin, X. & Zabriskie, T. M. The enduracidin biosynthetic gene cluster from Streptomyces fungicidicus. Microbiology 152, 2969–2983 (2006).
Fang, X. et al. The mechanism of action of ramoplanin and enduracidin. Mol. Biosyst. 2, 69–76 (2006).
Parenti, F., Pagani, H. & Beretta, G. Gardimycin, a new antibiotic from Actinoplanes. I. Description of the producer strain and fermentation studies. J. Antibiot. 29, 501–506 (1976).
Lee, M. D. Antibiotics from Microbispora. U.S. 6551591, (2005).
Beltrametti, F., Lazzarini, A., Brunati, C., Selva, E. & Marinelli, F. Production of demannosyl-A40926 by a Nonomuraea sp. ATCC 39727 mutant strain. J. Antibiot. 56, 310–313 (2003).
Hobbs, G., Frazer, C. M., Gardner, C. J., Cullum, J. A. & Oliver, S. G. Dispersed growth of Streptomyces in liquid culture. Appl. Microbiol. Biotechnol. 31, 272–277 (1989).
Shirahama, T., Furumai, T. & Okanishi, M. A modified regeneration method for streptomycete protoplasts. Agric. Biol. Chem. 45, 1271–1273 (1981).
Beltrametti, F. et al. Resistance to glycopeptide antibiotics in the teicoplanin producer is mediated by van gene homologue expression directing the synthesis of a modified cell wall peptidoglycan. Antimicrob. Agents Chemother. 51, 1135–1141 (2007).
Le Rudulier, D., Strom, A. R., Dandekar, A. M., Smith, L. T. & Valentine, R. C. Molecular biology of osmoregulation. Science 224, 1064–1068 (1984).
Garcia-Dominguez, M., Martin, J. F., Mahro, B., Demain, A. L. & Liras, P. Efficient plasmid transformation of the beta-lactam producer Streptomyces clavuligerus. Appl. Environ. Microbiol. 53, 1376–1381 (1987).
Hopwood, D. A., Chater, K. F., Dowding, J. E. & Vivian, A. Advances in Streptomyces coelicolor genetics. Bacteriol. Rev. 37, 371–405 (1973).
Beltrametti, F. et al. Valine influences production and complex composition of glycopeptide antibiotic A40926 in fermentations of Nonomuraea sp. ATCC 39727. J. Antibiot. 57, 37–44 (2004).
Kieser, T., Bibb, M. J., Buttner, M. J., Chater, K. F. & Hopwood, D. A. Practical Streptomyces Genetics (The John Innes Foundation, Norwich, 2000).
Demain, A. L. & Adrio, J. L. Strain improvement for production of pharmaceuticals and other microbial metabolites by fermentation. Prog. Drug Res. 65, 251, 253–289 (2008).
Hopwood, D. A., Wright, H. M., Bibb, M. J. & Cohen, S. N. Genetic recombination through protoplast fusion in Streptomyces. Nature 268, 171–174 (1977).
Kawamoto, I., Oka, T. & Nara, T. Cell wall composition of Micromonospora olivoasterospora, Micromonospora sagamiensis, and related organisms. J. Bacteriol. 146, 527–534 (1981).
Hong, H. J., Hutchings, M. I., Hill, L. M. & Buttner, M. J. The role of the novel Fem protein VanK in vancomycin resistance in Streptomyces coelicolor. J. Biol. Chem. 280, 13055–13061 (2005).
Hu, Y., Helm, J. S., Chen, L., Ye, X. Y. & Walker, S. Ramoplanin inhibits bacterial transglycosylases by binding as a dimer to lipid II. J. Am. Chem. Soc. 125, 8736–8737 (2003).
Goodfellow, M. & Quintana, E. in The Prokaryotes A Handbook on the Biology of Bacteria. Volume 3: Archea, Bacteria, Firmicutes, Actinomycetes. The family Streptosporangiaceae (eds Dworkin, M., Falkow, S., Rosemberg, E. & Schleifer, K.H.) 725–753 (Springer Science+Business Media, New York, 2006).
Mertz, F. P. Planomonospora alba sp. nov. and Planomonospora sphaerica sp. nov., two new species isolated from soil by baiting techniques. Int. J. Syst. Bacteriol. 44, 274–281 (1994).
Acknowledgements
This work was supported by FAR 2005–2006–2007–2008 to Flavia Marinelli and by an MIUR fellowship to Giorgia Letizia Marcone. We thank Mervyn Bibb, John Innes Institute, Norwich, UK for the kind gift of microbial strains.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marcone, G., Carrano, L., Marinelli, F. et al. Protoplast preparation and reversion to the normal filamentous growth in antibiotic-producing uncommon actinomycetes. J Antibiot 63, 83–88 (2010). https://doi.org/10.1038/ja.2009.127
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2009.127
Keywords
This article is cited by
-
Improvement of the conjugation transfer of N. gerenzanensis based on the synergistic effect of quorum sensing and antibiotic interference
AMB Express (2023)
-
Establishment of protoplast preparation protocol and genetic transformation system for Sclerotiophoma versabilis
Journal of Plant Pathology (2023)
-
Establishment of a visual gene knockout system based on CRISPR/Cas9 for the rare actinomycete Nonomuraea gerenzanensis
Biotechnology Letters (2023)
-
Production enhancement of the glycopeptide antibiotic A40926 by an engineered Nonomuraea gerenzanensis strain
Biotechnology Letters (2022)
-
Highly efficient genome editing in N. gerenzanensis using an inducible CRISPR/Cas9–RecA system
Biotechnology Letters (2020)