Abstract
Although grazers have long been recognized as top–down architects of plant communities, animal roles in determining microbial community composition have seldom been examined, particularly in aboveground systems. To determine the extent to which an animal can shape microbial communities, we conducted a controlled mesocosm study in situ to see if introducing mycophageous tree snails changed phyllosphere fungal community composition relative to matched control mesocosms. Fungal community composition and change was determined by Illumina sequencing of DNA collected from leaf surfaces before snails were introduced, daily for 3 days and weekly for 6 weeks thereafter. Scanning electron microscopy was used to confirm that grazing had occurred, and we recorded 3.5 times more cover of fungal hyphae in control mesocosms compared with those containing snails. Snails do not appear to vector novel microbes and despite grazing, a significant proportion of the initial leaf phyllosphere persisted in the mesocosms. Within-mesocosm diversities of fungi were similar regardless of whether or not snails were added. The greatest differences between the snail-treated and control mesocosms was that grazed mesocosms showed greater infiltration of microbes that were not sampled when the experiment commenced and that the variance in fungal community composition (beta diversity) was greater between leaves in snail-treated mesocosms indicating increased community heterogeneity and ecosystem fragmentation.
Similar content being viewed by others
Introduction
Animal activity, such as grazing, may be as important a factor in determining environmental microbial community assembly as climatic, geographic, physical, nutrient or other chemical factors that are observed routinely in microbial ecology studies. If heterogeneity is defined as the extent to which species are aggregated (Taylor, 1961), then animals may either increase or decrease microbial community heterogeneity depending on context and the nature of the interaction in question (Wardle et al., 2004; Tylianakis et al., 2008). For example, animals may promote homogeneity among microbial communities by active gardening. This type of animal microbial mutualistic interaction has arisen multiple times in numerous microbial and animal lineages, including textbook examples in insects such as termites (Aanen et al., 2002), ambrosia beetles (Beaver, 1989; Farrell et al., 2001) and leaf cutter ants (Poulsen and Boomsma, 2005). By vectoring microbes into an ecosystem, animals shape microbial communities to become more similar between geographically distinct locations (Aizenberg-Gershtein et al., 2013; Lax et al., 2014). Animals may also indirectly perturb microbial composition by modifying substrate availability or site physiochemistry. For example by foraging or burrowing, mammals may enable an alternative state in which microbial substrates are simplified by mixing (Eldridge et al., 2015; Pansu et al., 2015). Alternatively, animals can eat the hosts of microbes making their substrate range more complex (Bråthen et al., 2015).
Less studied are cases where animals are directly impacting wild non-cultivated microbial communities via direct grazing of microbes. Analogous literature from vegetation ecology suggests that, at least for plant communities, the impacts of grazing depend largely on factors such as productivity, grazing pressure and on the degree of diet selectivity of the grazers (Proulx and Mazumder, 1998). At opposite ends of the spectrum, grazing may result in either reduced alpha diversity (by selecting for one or a few dense weedy species), or alternatively, animals may create gaps in microbial communities and facilitate novel introductions and variability at small spatial scales (Denslow, 1985). It is unknown how these dynamics relate to aboveground microbial systems in which diversity, growth and dispersal rates are elevated relative to plants.
Here we determine whether presence of mycophageous tree snails impacts phyllosphere microbial community homogeneity. Hawaiian tree snails were once species rich and abundant across the archipelago, but they have succumbed to habitat loss, predation by exotic animals and a European fad for collecting numerous beautiful shells into cabinets (Hadfield, 1986). Tree snails feed continuously and voraciously throughout the night (O'Rorke et al., 2016), scraping phyllosphere microbes off of the leaf surface with a specialized radula. Although caution must be taken when making generalizations from mesocosm studies, we tested whether the reintroduction of tree snails is likely to increase or decrease species diversity at a fine scale, here sampled at the scale of a leaf, by reducing dominant taxa or, alternatively, by promoting the growth of particular fungal species. We also assess if snail introduction will change fungal heterogeneity across a coarser grain, which is the range over which snails can graze (that is, by measuring the variance of local alpha diversity within mesocosms). To test this, we translocated tree snails to five mesocosms and measured changes in fungal community composition over time. These mesocosms were compared with five matched control mesocosms that remained free of snails. As many of the fungi consumed by snails are still viable after passage through their gastrointestinal tract (O'Rorke et al., 2016), we also hypothesized that tree snails engage in microbial ‘farming’ by vectoring preferred fungus into the phyllosphere through fecal deposits as is observed in other snail systems (Silliman and Newell, 2011).
Materials and methods
Study design
We collected individuals of the non-listed, endemic O’ahu tree-snail Auriculella diaphana (Acahtinellidae), at a mid-elevation site, from a non-native host plant Cestrum nocturnum (Solanaceae; night-blooming jasmine), upon which they subsist in the study area (Holland et al., 2016), and translocated them to Metrosideros polymorpha (Myrtaceae), a native Hawaiian tree species on which arboreal snails have been observed (Hadfield, 1986; Meyer et al., 2014; Price et al., 2017). It is highly likely that mycophageous organisms other than snails can potentially interact with the fungal operational taxonomic units (OTUs) inside either control or snail-treated mesocosms. Although the recipient site is located within the snails’ historical range (Pilsbry and Cooke, 1912), multiple surveys over the past three decades indicate that snails have been locally extirpated. Therefore, the new host plant had never hosted snails and is >2 km from the remnant snail population (study site location co-ordinates available from the Hawaii Department of Land and Natural Resources upon request). We placed 10 adult snails on living tree branches inside each of five mesocosms (50 snails total). Mesocosms were ~36 l cylinders constructed from fiberglass mesh sewn at the edges and reinforced with polyethelyne tape (Figure 1). The mesh size (1 mm2) reduced light to the branch by 30% (measured by lux, Light Sensor LS-BTA (Vernier, Beaverton, OR, USA)). We paired each of these snail-treated mesocosms with a mesocosm that was free of snails but otherwise identical, so that the experiment had a total of 10 mesocosms distributed into five blocks. Mesocosms blocks were located on the same tree (to minimize host genotype effects) and spaced at least 80 cm apart. Mesocosms were placed over leaves of the same age (new season leaves of <4 months). We transported snails to the mesocosms in sterile 10 cm polypropylene Petri dishes (Falcon, Corning, NY, USA). We collected snail feces deposited in transit (n=10) and stored them in hexadecyltrimethylammonium bromide (CTAB) buffer for subsequent DNA extraction and Illumina sequencing, these were used to determine if the snails do vector fungus into new habitats. Two vials of CTAB buffer were taken into the field, opened and then stored until the DNA was extracted for PCR amplification and sequencing, these served as negative controls for field contamination during the sampling process.
(a) We placed mycophageous tree snails into five mesh mesocosms of ~36 l. A matched snail-free control mesocosm was placed adjacent to each snail-treated mesocosm, making up ten mesocosms in total. (b) To determine how snails changed the fungal community composition of the phyllosphere we swabbed leaves before we introduced the snails, then daily (3 days) and weekly thereafter for 6 weeks. We extracted DNA from the swabs and Illumina sequenced the ITS1 intergenic region to identify which fungus were present at each time point. To confirm whether grazing by snails reduced microbial abundance we took scanning electron microscope images of leaf surfaces from (c) snail-treated and (d) control mesocosms on the final day of the experiment.
We sampled mesocosms at 1300 hours Hawaiian standard time before the snails were introduced (T0) and then near this time (±30 min) each day for the subsequent 3 days and then weekly thereafter for a total of 6 weeks. This duration is sufficient to detect community change in other studies (Lax et al., 2014). For each sampling event, we swabbed the entire top and bottom surfaces of three leaves (>4 cm in length) from each mesocosm and stored the swabs in CTAB buffer until DNA extraction and Illumina sequencing. Triplicate swabs of individual leaves (that is, biological replicates, not triplicate technical replicates of the same leaves) were taken from mesocosms at Tfinal, and T0 to determine extent of heterogeneity within mesocosms. Leaves were also collected at Tfinal and stored in scanning electron microscopy (SEM) fixative (2.5% gluteraldehyde buffered in 0.1 m sodium cacodylate) for SEM analysis.
Amplicon sequencing
We determined the composition of fungal communities on leaves via Illumina sequencing of DNA amplicons following the methods outlined in O’Rorke et al. (2015). Briefly, we extracted DNA from feces using the 96-well PowerPlant Pro DNA Isolation Kit (Mo Bio Laboratories, Jefferson City, MO, USA) and then PCR amplified for 20 cycles with primers that amplified the ITS1 region of fungi (Gardes and Bruns, 1993). The ITS1 region is well represented in public DNA sequence databases and has been demonstrated to work well in previous studies of the Hawaiian phyllosphere (Zimmerman and Vitousek, 2012; O’Rorke et al., 2015; Price et al., 2017) and the ITS1f primer is, in our experience, the only primer that will not co-amplify plant DNA when fungi are present in low biomass. The primers had a 5’ universal sequence as used in the standard Nextera protocol (Illumina, 2012; PCR master mixes and thermocycler protocol in Supplementary Materials). PCR amplicons were transferred to a second 22-cycle reaction in which unique barcodes were added to each sample, as well as the Illumina i5 and i7 adapters (Supplementary Data S1). Reactions were cleaned using a SPRI plate (Beckman Coulter, Brea, CA, USA) and Sera-Mag Speedbeads (FisherSci, Pittsburgh, PA, USA) in an amplicon:bead ratio of 1:0.7, were eluted in water, made equimolar in SequalPrep Normalization plates (Invitrogen, Grand Island, NY, USA) and subsequently pooled. Pooled samples were cleaned again with magnetic beads, and submitted to Beckman Coulter Genomics (Danvers, MA, USA) for sequencing on an Illumina MiSeq sequencer using the MiSeq Reagent v3 600-cycle chemistry (Illumina, San Diego, CA, USA). PCR-negative controls were also amplified and sequenced; contents of these controls were identical to the samples and were PCR amplified at the same time, except that only molecular grade water was added to the PCR reaction mix.
Microscopy
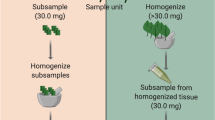
We used SEM to quantify differences between snail-treated and control mesocosms. We sectioned gluteraldehyde fixed leaves, then dehydrated them by serial washing in increasing concentrations of ethanol, critical point dried and then sputter coated with gold-palladium after mounting on metal stubs. Samples were viewed on a Hitachi S-4800 (Hitachi, Tokyo, Japan) field emission SEM at × 1400 magnification. SEM images were taken of the top and bottom of a leaf from each of the 10 mesocosms. Transects were run at the widest part of the leaf between the central vein (midrib) and the leaf’s edge at 1 mm intervals. We quantified filamentous microbes across these transects by measuring their surface area as a total of the proportion of pixels in Adobe Photoshop, and then converted this into area coverage (mm2).
Bioinformatics and analyses
Forward and reverse Illumina sequences were merged using PEAR (Zhang et al., 2013), demultiplexed in QIIME (Caporaso et al., 2010) and clustered into OTUs at 97% similarity using UPARSE (Edgar, 2013). Subsequent analyses were performed in R 3.1.3 (www.R-project.org). Subsampling was performed to standardize read number to the lowest number of reads that would enable the majority of samples to be included in the data set. The impact of grazing on the predictability and uniformity of fungal species abundance at a local scale was calculated using Shannon’s index H (Shannon and Weaver, 1949) in R using the vegan package, as was species richness (Dixon, 2009). Whether richness and H varied significantly between treatment and control groups was assessed by analysis of variance. We determined whether snails preferentially feed on particular fungus using Paine’s per capita effect index: (E–C)/(C × P), where E is the experimental food density, C is the control density and P refers to predator density (Paine, 1992). This index gives a value of –1 when a consumer has a strong interaction with a prey item (that is, removes it) relative to the control, a value of 0 for no interaction and any positive value is interpreted as the consumer favoring the continued existence or recruitment of a species.
Temporal analyses of microbial community composition
We hypothesized that the addition of grazers to an ecosystem will change the microbial community composition over time. Consequently, we tested whether the model y~time+treatment+(time × treatment) explained changes in OTU abundances by permutational analysis of variances after running the manyglm command in the mvabund package (Wang et al., 2012; Warton, 2008; Szöcs et al., 2015) to ascertain the effect of interaction term. The ‘treatment’ term refers to whether the mesocosm contained snails or not, but includes measurements made before introduction of snails. To reflect the repeated-measures experimental design, the null hypothesis was tested by permutation of OTU values at any time points within each mesocosm to see if the random order deviated from the actual order of OTU values (R script in Supplementary Files; Dixon, 2009). We also graph the raw compositional abundance data in ggplot (Warton, 2008; Wickham, 2009) to visualize how the mean–variance relationship of the fungus in snail-treated mesocosms differed from that of controls. The mean–variance relationship can be taken as an indicator of system heterogeneity, in which systems with greater intercepts and slopes can be considered to have greater heterogeneity (Taylor, 1961; Fairweather, 1988).
We used the Bayesian statistical program SourceTracker v 1.0 (Knights et al., 2011) to determine if the composition of fungal communities was consistent over time, if snail feces vector microbes into an environment, and if so, how long these microbes persist in the community. For this analysis, we included three initial groups of fungus: the initial phyllosphere (measured at T0 in all mesocosms before snails were introduced), fungus from snail feces (suggestive of snail ‘vectoring’) and unexplained variance attributable to the environment, aerobiota or sampling error. All subsequent samplings of the fungus community were then compared with these initial three groups to see which initial group became more abundant over time. In a community with high resistance (sensu Pimm, 1984; for a glossary refer de Vries and Shade, 2013), all composition would be identical to initial composition and would be classified as ‘phyllosphere’. If feces were a source of fungus, then ‘fecal’ OTUs would come to be a significant part of the microbiota and if the snails grazing increased the infiltration of microbes from the regional species pool, then this ‘environmental’ group would become more dominant in the snail-treated mesocosms than in the controls. A likelihood ratio test was used to evaluate if a mixed effect model that included ‘treatment’ significantly explained the data. In this model, ‘treatment’ was nested inside ‘block’, which was treated as a random factor.
Results
Sequencing results
Miseq amplicon sequencing yielded a total of 3 217 880 reads from this study that passed quality control (mean=20 364±1116 s.e.). As there were two samples that yielded fewer than 1000 reads (767 and 557 reads), all samples were randomly subsampled to a depth of 750 reads. The four negative PCR controls yielded 1, 1, 1 and 0 reads and controls for field collection and extraction negatives yielded 293 and 23 reads, respectively, suggesting that contamination did not contribute much noise to our data or analyses.
Temporal analyses of changes in microbial composition
Does snail treatment disturb the phyllosphere through time?
The interaction term snail treatment × time is significant (P<0.05) indicating that snails modify the abundances of fungal OTUs over time (Table 1). Generally, the changes to fungal OTUs in the snail-treated mesocosms were either small changes in abundances over time (Table 2 and Figure 2) or erratic invasions and there was greater heterogeneity in the snail-treated mesocosms (Figure 3). Paine’s per capita effect index demonstrated that no fungal OTUs had a strong negative or positive effect as a result of grazing (Table 2).
Raw abundance data over time. The abundances of the 17 most abundant fungal taxa from within the mesocosms (which together comprised two-thirds of the total read count) are shown. The plots are ordered by effect size, with OTU_5637, OTU_1 and OTU_10 having the greatest effect size under the treatment × time interaction and OTU_15, OTU_8 and OTU_5 having lower effects. Data are log transformed and lines are fitted by local polynomial regression (loss).
Sources of phyllosphere fungi and the heterogeneity of the assemblage. There was considerable community inertia as fungi persisted through the course of the experiment and were present in the initial phyllosphere. However, fungi from the environmental species pool (blue area of figure) also infiltrated the mesocosm, but this was more so in the snail-treated mesocosm, (a) than the control mesocosms (b). The microbes in the feces of snails was also determined before snails were introduced into the mesocosms to determine if snail feces would be a source for fungus being vectored into the new community. These fecal samples were not a significant part of the community assemblage, occurring in a large part in only one sample point in mesocosm G, where it is likely that a fecal sample adhering to a leaf as the commencement of the experiment was inadvertently sampled. (c) Despite the persistence of certain OTUs inside the mesocosms, the heterogeneity of these OTUs was greater in the snail-treated mesocosms, where heterogeneity is measured as degree of variance as a function of mean (as per Taylor, 1961), then snail-treated mesocosms all have greater heterogeneity than any control mesocosm, which is represented by the slopes and intercepts of all lines fitting the log (variance)~log (mean) being greater for the snail-treated mesocosms than control mesocosms.
Sources of microbes
The Bayesian software Sourcetracker (Knights et al., 2011) found that the composition of the fungal communities in the mesocosms was dominated by microbes that could be traced back to the initial phyllosphere community (Figures 3a and b). However, fungal OTUs from the environmental species pool and were significantly greater in snail-treated mesocosms than controls (χ2(1)=14.43, P=0.00015), contributing 25.5% (±3.2% s.e.) of fungus counted in the snail-treated mesocosm after 1 week and 11.3% (±1.1% s.e.) to the control after 1 week. Microbes assigned to feces were only a small component of the mesocosm assemblages with the exception of three mesocosms: G, I and J. Mesocosm G had an elevated proportion of fungus assigning to feces on days 10 and 17 (Figure 3).
End-point analyses: differences in absolute abundances and alpha diversity
Alpha diversity and evenness for fungi were relatively similar between mesocosms for snail treatments (H=3.37±0.05 s.e.) and control mesocosms (H=3.45±0.04 s.e.). At the end of the experiment, the mean Shannon alpha diversity of fungal mescosms was similar for snail treatments (3.36±0.11 s.e.) and controls (3.57±0.09 s.e.) and were not significantly different (F(1, 28)=1.41, P=0.25). However, mesocosms treated with snails had substantially greater variation in the s.d. of the mean alpha diversity when multiple samples were measured within the mesocosm (Figure 4). In other words, snail addition increased the patchiness of the relative abundances of microbes. Species richness did significantly decline in snail-grazed mesocosms (F(1, 28)=7.03, P=0.01, values in Table 3). Microscopy was used to compare the absolute cover of fungal hyphae between snail treatments and controls (Figure 5). Fungal hyphae was 3.59 times more dense on control mesocosm leaves than it was on snail leaves, indicating that snails significantly reduce fungal cover (Figure 5).
Snails increase within-system variability. Each mesocosm was sampled in triplicate before the experiment commenced (T0) and 6 weeks after the introduction of snails to half of the mesocosms. The within-mesocosm variability of fungal diversity increased if treated with snails, but slightly decreased in the no-snail control mesocosms over the time course of the experiment. Whiskers are 1.5 times the interquartile range±the first and third quartile.
Assessment of grazing impacts on microbial abundances using scanning electron microscopy. Fungal hyphal abundance is reduced by grazing as snail-grazed mesocosms (open boxes) have 3.5 times less fungal biomass than ungrazed controls (striped boxes). Abundances were determined by measuring the surface area occupied by fungal hyphae across transect of leaves at the widest part of the leaf, so that the number of counts was at least 12 along each transect. Whiskers are 1.5 times the interquartile range±the first and third quartile.
Discussion
Our study found that snail introductions impacted microbial communities by promoting infiltrations of fungal OTUs from the environmental species pool. We hypothesized that fungal community assemblages would be changed by mycophagous tree snails being reintroduced into a habitat. This was the case, because not only do fungal communities change over time, but communities with snails change differently compared with those without snails. It was not possible to predict a priori if the presence of snails would increase or decrease community homogeneity, because studies of other grazing systems suggest that snail grazing could do either. Our comparison of mesocosms treated with snails with those without snails demonstrated that snails alter microbial assembly by clearing fungal patches (which is likely to reduce resource limitation).
Spatial heterogeneity and patch size
In this study, the mean alpha diversity was the same for control and treatment mesocosms at the scale of the leaf, most likely due to changes in the relative abundances of fungal OTUs. However, this study found that there was much greater deviation from the mean of alpha diversity within mesocosms treated with snails compared with controls (Figure 4). This is explained by the way that disturbances disrupt community homogeneity and creates patches of habitat favorable to diverse environmental microbes found elsewhere (Pickett and White, 1985). Grazing snails rescale the size of patches in their environment, making them considerably smaller and more numerous than the single mesocosm inside which they are enclosed. By contrast, in the control mesocosms, the patch size remains identical to the mesocosm size as evidenced by high similarity of samples. The leaves grazed by snails do experience a small, but statistically significant, decrease in fungal richness (Table 3). A recent study examining the impacts of foraging animals on soil microbiology by Eldridge et al. (2015) found that there were no differences in fungal or bacterial richness between grazed and undisturbed sites, despite finding differences in beta diversity. The contrasting response of richness to grazing in this study is most likely a reflection of this study and the soil system having contrasting productivity or disturbance frequencies (Huston, 2014), but the both this study and Eldridge’s et al. (2015) indicate how animals can modify the structure of microbial communities at a broader scale.
Temporal heterogeneity
Stochasticity tends to dominate assembly processes before a habitat reaches carrying capacity, after which point deterministic environmental filters become more important for community assemblage (Fukami, 2010). The grazing action of snails prolongs this stochastic phase, and because snails reduce microbial biomass the mechanism for prolonging stochasticity is likely to be through competitor removal. Crowther et al. (2013) found that grazing of soil fungi by isopods removed the mycelia of dominant Basidomycetes and diversified the functional profile of the fungal assemblage.
Our results indicate snails do not target a particular fungal OTU as food, but instead have a minor impact on a broad range of OTUs (Table 2). Visual inspection of leaves at the close of the experiment also shows that snails clear the leaf surfaces in a general and not in a ‘picky’ manner. Consistent with our results (Figure 3c), Berlow (1999) argued that consumers that weakly interact with food species tend to increase the variance within the trophic group on which they predate. This is because consumers that strongly interact with prey groups should consistently remove the same food items and reduce variance (Berlow, 1999). Conversely, the absence of a predator can lead to a loss of diversity in the lower trophic levels because of the unchecked growth of individual prey species (Paine, 1966). The response of fungus to grazing, or snail selectivity require further research, but it does appear that snails do not specifically target dominant fungi (for example, isopods Crowther et al., 2013; and collembola Jørgensen et al., 2005) or uncompetitive fungi (for example, millipedes Crowther et al., 2011), but instead serve to clear areas of leaf surfaces and prolong the initial phase of assembly.
Snails increased the occurrence of fungal OTUs from the environmental species pool (Figures 3a and b), where that species pool consists of fungi from the aerobiota, endophytes or OTUs whose abundance was below detection at the sampling depth used in the study, diminishing the role that plant hosts may otherwise exert on phyllosphere communities. These spikes in abundance are likely to be the result of opportunistic fungi infiltrating spaces cleared by the snails, which is likely to account for the significant treatment × time interaction that was detected (Table 1). In a previous study conducted across the extent of the Waianae mountain range, we showed that the phyllosphere composition of trees containing Achatinella snails was more greatly influenced by location than by host tree identity (O'Rorke et al., 2015). Plant host identity strongly impacts phyllosphere community composition, and although phyllosphere communities are geographically structured, the plant host can be a stronger determining factor in phyllosphere composition (Kim et al., 2012)—even at the continental scale (Redford et al., 2010; Ludlow et al., 2016). The results of this study indicate that grazing pressure may reduce the specificity of the relationship between host plant and phyllosphere community as grazing keeps a high proportion of the microbial community in a sustained transitory state.
The similarity of microbial community composition in the phyllosphere over time is evidence for the selectivity of plants to determine which microbes persist (Lindow and Brandl, 2003), despite substantial removal of microbes by grazing. SourceTracker analyses show that there is a substantial consortium of fungi that are a persistent part of the phyllosphere in both treatments (Figure 3). It is plausible that some of the OTUs that persisted are epiphytes that also occur in the intercellular space inside leaves, which is consistent with a ‘refugia’ response to predation (Belyea and Lancaster, 1999). Phyllosphere microbes often originate from within the plant (Whipps et al., 2008), and if the cells that recolonize cleared leaf patches arise from within the leaf stomata, then this would explain some consistency in the assemblage composition.
Empirical studies of how grazing pressure affects richness (Proulx and Mazumder, 1998) have shown variable results. This variance has been explained, in part, by the nutrient status of the environment (Proulx and Mazumder, 1998), where decreased plant diversity is typical in grazed habitats with low nutrients. The phyllosphere is characteristically oligotrophic (Vorholt, 2012), so our finding that grazing does not impact alpha diversity contradicts this trend. That phyllosphere microbes are host associated is one important distinction between our study and the majority of those considering the grazing impacts of macro-organisms. In our case, the community composition of fungi may be constrained by the plant host, which maintains more homogenous diversity levels compared with an inert substrate. In this study, we controlled for the effects of plant genotype by using single host, but further research is required to determine whether the effects that we observe vary between different tree hosts. We also used a constant number of snails in each of our treatments, but it is likely that fungal relative abundances will respond differently under different grazing intensities (Grime, 1973; Connell, 1978; Huston, 2014). Therefore, further experiments are required to shed light on whether host association or other factors contribute to differences in grazing impacts on microbial assemblages.
Feces are not a source for novel OTUs
We hypothesized that snails are vectors for microbes, because they not only excrete viable fungi, but when they do so they deposit them within nutrient-rich feces that should facilitate rapid and vigorous growth (O'Rorke et al., 2015; Price et al., 2017). The results of this study do not support this hypothesis, and although there are brief pulses when the microbes found in translocated snail feces can be detected on the surfaces of leaves, these microbes subsequently become a minor component of the phyllosphere. This is surprising, but is concordant with our personal observations that snail feces persisted in snail mesocosms for weeks at a time and did not appear to be degraded by viable microbes within. The pulses when a fecal signature is detected in SourceTracker analyses may be the result of co-sampling feces along with the phyllosphere. That feces are not a substrate for rampant microbial growth could be due to physical properties of the feces. Tree snail feces are compacted with mucus, which is profoundly hydrophobic in A. diaphana (personal observation, see Supplementary Materials), and therefore not particularly conducive to microbial growth—despite containing viable microbes (O’Rorke et al., 2016). In addition, inspection of SEM images and leaf surfaces give no indication that A. diaphana snails damage the leaf cuticle when grazing microbial films. They are therefore very different from the marsh snail Littoraria irrorata, which damages live leaf surfaces of salt marsh cordgrass (Spartina alterniflora) when feeding and deposits fecal pellets on leaf wounds, encouraging fungal growth (Silliman and Newell, 2011).
The SourceTracker results of this study make it clear that snails are not homogenizing their environments by seeding microbes as we had hypothesized, but instead that snails increase the influence of environmental microbes via grazing. The net result of native tree snail grazing and feces deposition was enhancement of microbial diversity in the phyllosphere. Snails therefore change microbial communities by reducing the host plants’ top–down controls on the community structure while facilitating greater community turnover than in environments without snails.
Conclusion
A. diaphana grazing of vegetative fungi off leaf surfaces increases ecosystem patchiness and the incidence of colonization from the wider environmental species pool, but they do not seed fungus into the ecosystem through their feces. The disruptive manner in which these generalist feeders modify microbial assembly is therefore different from that observed in mycophageous animals that are specialists. We conclude that it is important for microbial ecologists to observe whether animals could account for any of the unexplained variance in their community assembly models and to observe if these animals are specialists, generalists or simply foragers that modify assembly processes. The results of this study are also in accord with other recent studies with a conservation focus (for example, Clarke et al., 2015) that point out that management of species of conservation interest needs to take into account how animal extinctions and reintroductions will impact on the composition and ecosystem services of the microbiota.
References
Aanen DK, Eggleton P, Rouland-Lefevre C, Guldberg-Froslev T, Rosendahl S, Boomsma JJ . (2002). The evolution of fungus-growing termites and their mutualistic fungal symbionts. Proc Natl Acad Sci USA 99: 14887–14892.
Aizenberg-Gershtein Y, Izhaki I, Halpern M . (2013). Do honeybees shape the bacterial community composition in floral nectar? PLoS One 8: e67556.
Beaver RA . (1989) Insect - Fungus relationships in the bark and ambrosia beetles. In : Wilding N, Collins NM, Hammond PM, Webber JF (eds). Insect - Fungus Interactions. Academic Press: New York, NY, USA, pp 121–143.
Belyea LR, Lancaster J . (1999). Assembly rules within a contingent ecology. Oikos 86: 402–416.
Berlow EL . (1999). Strong effects of weak interactions in ecological communities. Nature 398: 330–334.
Bråthen KA, Jahiri X, Jusdado JGH, Soininen EM, Jensen JB . (2015). Fungal endophyte diversity in tundra grasses increases by grazing. Fungal Ecol 17 (C): 41–51.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Clarke LJ, Weyrich LS, Cooper A . (2015). Reintroduction of locally extinct vertebrates impacts arid soil fungal communities. Mol Ecol 24: 3194–3205.
Connell JH . (1978). Diversity in tropical rain forests and coral reefs. Science 199: 1302–1310.
Crowther TW, Boddy L, Jones TH . (2011). Outcomes of fungal interactions are determined by soil invertebrate grazers. Ecol Lett 14: 1134–1142.
Crowther TW, Stanton DWG, Thomas SM, A'Bear AD, Hiscox J, Jones TH et al. (2013). Top-down control of soil fungal community composition by a globally distributed keystone consumer. Ecology 94: 2518–2528.
Denslow JS . (1985) Disturbance mediated coexistence of species. In: Pickett STA, White PS (eds). ‘The Ecology of Natural Disturbance and Patch Dynamics’. Academic Press: Orlando, FL, USA, pp 307–323.
Dixon P . (2009). VEGAN, a package of R functions for community ecology. J Vegetation Sci 14: 927–930.
Edgar RC . (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10: 996–998.
Eldridge DJ, Woodhouse JN, Curlevski NJA, Hayward M, Brown MV, Neilan BA . (2015). Soil-foraging animals alter the composition and co-occurrence of microbial communities in a desert shrubland. 9: 2671–2681.
Fairweather PG . (1988). Predation can increase variability in the abundance of prey on seashores. Oikos 53: 87–92.
Farrell BD, Sequeira AS, O'Meara BC, Normark BB, Chung JH, Jordal BH . (2001). The evolution of agriculture in beetles (Curculionidae: Scolytinae and Platypodinae). Evolution 55: 2011–2027.
Fukami T . (2010) Community assembly dynamics in space. In : Verboef HA, Morin PJ (eds). Community Ecology: Processes, Models, and Applications. Oxford University Press: London, UK, pp 45–54.
Gardes M, Bruns TD . (1993). ITS primers with enhanced specificity for basidiomycetes—application to the identification of mycorrhizae and rusts. Mol Ecol 2: 113–118.
Grime JP . (1973). Competitive exclusion in herbaceous vegetation. Nature 242: 344–347.
Hadfield MG . (1986). Extinction in Hawaiian achatinelline snails. Malacologia 27: 67–81.
Holland BS, Chiaverano LM, Howard CK . (2017). Diminished fitness in an endemic Hawaiian snail in nonnative host plants. Ethol Ecol Evol 29: 229–240.
Huston MA . (2014). Disturbance, productivity, and species diversity: empiricism vs. logic in ecological theory. Ecology 95: 2382–2396.
Illumina. (2012). Nextera DNA Sample Preparation Guide, 1–44.
Jørgensen HB, Johansson T, Canbäck B, Hedlund K, Tunlid A . (2005). Selective foraging of fungi by collembolans in soil. Biol Lett 1: 243–246.
Kim M, Singh D, Lai-Hoe A, Go R, Abdul Rahim R, Ainuddin AN et al. (2012). Distinctive phyllosphere bacterial communities in tropical trees. Microbial Ecol 63: 674–681.
Knights D, Kuczynski J, Charlson ES, Zaneveld J, Mozer MC, Collman RG et al. (2011). Bayesian community-wide culture-independent microbial source tracking. Nat Methods 8: 761–763.
Lax S, Smith DP, Hampton-Marcell J, Owens SM, Handley KM, Scott NM et al. (2014). Longitudinal analysis of microbial interaction between humans and the indoor environment. Science 345: 1048–1052.
Lindow SE, Brandl MT . (2003). Microbiology of the phyllosphere. Appl Environ Microbiol 69: 1875–1883.
Ludlow CL, Cromie GA, Garmendia-Torres C, Sirr A, Hays M, Field C et al. (2016). Independent origins of yeast associated with coffee and cacao fermentation. Curr Biol 26: 965–971.
Meyer WM, Gary DT, Yeung NW, Dirks C, Leung K, Leon JA et al. (2014). Native arboreal land snails in the Mt Kaala Natural Area Reserve, Oahu, Hawaii, have similar plant preferences: implications for conservation. J Molluscan Stud 80: 469–472.
O'Rorke R, Cobian GM, Holland BS, Price MR, Costello V, Amend AS . (2015). Dining local: the microbial diet of a snail that grazes microbial communities is geographically structured. Environ Microbiol 17: 1753–1764.
O’Rorke R, Holland BS, Cobian GM, Gaughen K, Amend AS . (2016). Biological conservation. Biol Conserv 198: 177–182.
Paine RT . (1966). Food web complexity and species diversity. Am Naturalist 100: 65–75.
Paine RT . (1992). Food-web analysis through field measurement of per capita interaction strength. Nature 355: 73–75.
Pansu J, Winkworth RC, Hennion F, Gielly L, Taberlet P, Choler P . (2015). Long-lasting modification of soil fungal diversity associated with the introduction of rabbits to a remote sub-Antarctic archipelago. Biol Lett 11: 20150408.
Pickett STA, White PS . (1985). Patch dynamics: a synthesis. In: Pickett STA, White PS (eds). The Ecology of Natural Disturbance and Patch Dynamics. Academic Press: New Brunswick, NJ, USA, pp 371–384.
Pilsbry HA, Cooke CM Jr . (1912–1914) Manual of Conchology. Structural and Systematic. With illustrations of the Species. Second Series: Pulmonata, Vol XXII Achatinellidae Academy of Natural Sciences: Philadelphia.
Pimm SL . (1984). The complexity and stability of ecosystems. Nature 307: 321–326.
Poulsen M, Boomsma JJ . (2005). Mutualistic fungi control crop diversity in fungus-growing ants. Science 307: 741–744.
Price MR, O'Rorke R, Amend AS, Hadfield MG . (2017). Diet selection at three spatial scales: implications for conservation of an endangered Hawaiian tree snail. Biotropica 49: 130–136.
Proulx M, Mazumder A . (1998). Reversal of grazing impact on plant species richness in nutrient-poor vs. nutrient-rich ecosystems. Ecology 79: 2581–2592.
Redford AJ, Bowers RM, Knight R, Linhart Y, Fierer N . (2010). The ecology of the phyllosphere: geographic and phylogenetic variability in the distribution of bacteria on tree leaves. Environ Microbiol 12: 2885–2893.
Shannon CE, Weaver W . (1949) The Mathematical Theory of Communication. The University of Illinois Press: Urbana, IL, USA, p 117.
Silliman BR, Newell SY . (2011). Fungal farming in a snail. Proc Natl Acad Sci USA 100: 15643–15648.
Szöcs E, Van den Brink PJ, Lagadic L, Caquet T, Roucaute M, Auber A et al. (2015). Analysing chemical-induced changes in macroinvertebrate communities in aquatic mesocosm experiments: a comparison of methods. Ecotoxicology 24: 760–769.
Taylor LR . (1961). Aggregation, mean and variance. Nature 189: 732–735.
Tylianakis JM, Didham RK, Bascompte J, Wardle DA . (2008). Global change and species interactions in terrestrial ecosystems. Ecol Lett 11: 1351–1363.
Vorholt JA . (2012). Microbial life in the phyllosphere. Nat Rev Microbiol 10: 828–840.
de Vries FT, Shade A . (2013). Controls on soil microbial community stability under climate change. Front Microbiol 4: 265.
Wardle DA, Bardgett RD, Klironomos JN, Setälä H, van der Putten WH, Wall DH . (2004). Ecological linkages between aboveground and belowground biota. Science 304: 1629–1633.
Wang Y, Naumann U, Wright ST, Warton DI . (2012). mvabund- an Rpackage for model-based analysis of multivariate abundance data. Methods Ecol Evol 3: 471–474.
Warton DI . (2008). Raw data graphing: an informative but under‐utilized tool for the analysis of multivariate abundances. Austral Ecol 33: 290–300.
Whipps JM, Hand P, Pink D, Bending GD . (2008). Phyllosphere microbiology with special reference to diversity and plant genotype. J Appl Microbiol 105: 1744–1755.
Wickham H . (2009) ggplot2: Elegant Graphics for Data Analysis. Springer: New York, NY, USA.
Zhang J, Kobert K, Flouri T, Stamatakis A . (2013). PEAR: a fast and accurate Illumina paired-end reAd mergeR. Bioinformatics 30: 614–620.
Zimmerman NB, Vitousek PM . (2012). Fungal endophyte communities reflect environmental structuring across a Hawaiian landscape. Proc Natl Acad Sci USA 109: 13022–13027.
Acknowledgements
We acknowledge the help of David Sischo, Jenny Prior and Cynthia King at the DNLR. Chloë Heiniemi, Talia Sellars, Casey Jones and Gerry Cobian for volunteering their time and help in the field, and Tina Carvahlo for electron microscopy at the BEMF at the University of Hawai’i. This project was funded through the US Army cooperative agreement W9126G-11-2-0066 with Pacific Cooperative Studies Unit, University of Hawaii through the Pacific International Center for High Technology Research, State of Hawaii, DLNR, Special Funds and NSF grant #1255972. FASTQ files have been deposited in NCBI’s short read archive (SRA) under project number PRJNA357010. All research was conducted under the Department of Land and Natural Resources Native Invertebrate Research and Collecting permit # FHM15-T&E-16.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
O'Rorke, R., Tooman, L., Gaughen, K. et al. Not just browsing: an animal that grazes phyllosphere microbes facilitates community heterogeneity. ISME J 11, 1788–1798 (2017). https://doi.org/10.1038/ismej.2017.52
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.52