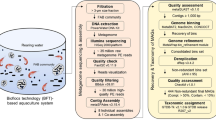
Abstract
Rapid gravity sand filtration is a drinking water production technology widely used around the world. Microbially catalyzed processes dominate the oxidative transformation of ammonia, reduced manganese and iron, methane and hydrogen sulfide, which may all be present at millimolar concentrations when groundwater is the source water. In this study, six metagenomes from various locations within a groundwater-fed rapid sand filter (RSF) were analyzed. The community gene catalog contained most genes of the nitrogen cycle, with particular abundance in genes of the nitrification pathway. Genes involved in different carbon fixation pathways were also abundant, with the reverse tricarboxylic acid cycle pathway most abundant, consistent with an observed Nitrospira dominance. From the metagenomic data set, 14 near-complete genomes were reconstructed and functionally characterized. On the basis of their genetic content, a metabolic and geochemical model was proposed. The organisms represented by draft genomes had the capability to oxidize ammonium, nitrite, hydrogen sulfide, methane, potentially iron and manganese as well as to assimilate organic compounds. A composite Nitrospira genome was recovered, and amo-containing Nitrospira genome contigs were identified. This finding, together with the high Nitrospira abundance, and the abundance of atypical amo and hao genes, suggests the potential for complete ammonium oxidation by Nitrospira, and a major role of Nitrospira in the investigated RSFs and potentially other nitrifying environments.
Similar content being viewed by others
Introduction
Microorganisms have an essential role in Earth’s global biogeochemical cycles, and they provide a multitude of nature’s ecosystem services. Microbes also dominate the technical systems used in wastewater treatment, drinking-water production, waste management and bioenergy generation (Rittmann, 2006). Although these technical systems have been in use for many years, the study of their microbial composition has only recently been initiated. Unraveling the microbial composition and processes occurring in these systems will not only help optimize the technical processes but can also lead to the discovery of novel pathways and organisms. In the era of next-generation sequencing techniques, metagenomics-based approaches have enabled the discovery of a suite of novel microbial pathways (Schleper et al., 2005; Bryant et al., 2007; Ettwig et al., 2010; Carrión et al., 2015) and the expansion of the tree of life (Brown et al., 2015). Some of these discoveries have been made in technical systems; for example, novel candidate phyla and expanded metabolic versatility have been identified in aerobic and anaerobic bioreactors (Wexler et al., 2005; Rinke et al., 2013; Nobu et al., 2015).
Rapid sand filters (RSFs), are engineered microbial systems used throughout the world for the production of drinking water from both surface and groundwater. With sufficiently high groundwater quality, rapid sand filtration—often in two serial filters—can be the sole treatment of groundwater before its distribution. Inlet water is initially aerated, which aims to physically remove compounds such as sulfide and methane and oxygenate the water. The main electron donors entering the serial filters are ammonia, reduced manganese and iron, small amounts of assimilable organic carbon, and residual methane and hydrogen sulfide (Tatari et al., 2013; Albers et al., 2015). Most of these compounds are oxidatively transformed by microbes in the RSFs (Lee et al., 2014). Hence, microorganisms colonize the filter material and its mineral coatings (Gülay et al., 2014). In contrast to other engineered microbial systems, such as wastewater treatment plants and anaerobic digesters, RSFs treating groundwater can be considered oligotrophic (with influent concentrations of individual constituents below mM) and receive predominantly inorganic electron donors, supporting chemolithotrophic metabolisms (Gülay et al., 2016).
Central metabolic functions in RSFs have previously been assigned to taxa based on their activity in other environments—Nitrosomonas, Nitrospira, Gallionella and Bacillus clades have been proposed as important ammonium (van der Wielen et al., 2009; Lautenschlager et al., 2014), nitrite (Gülay et al., 2014; Albers et al., 2015; LaPara et al., 2015), iron (Søgaard, 2001; de Vet et al., 2012; Li et al., 2013) and manganese oxidizers (Mouchet, 1992; Cerrato et al., 2010), respectively. Ammonia-oxidizing archaea (AOA) have been detected in RSFs at low abundance (van der Wielen et al., 2009; Albers et al., 2015), although in a few examples they outnumbered ammonia-oxidizing bacteria (AOB; Wang et al., 2014; Nitzsche et al., 2015). However, the total AOB and AOA densities are often insufficient to explain the high abundances of Nitrospira relative to the ammonium oxidizers (reviewed in Gülay et al., 2016). The role of other dominant clades within RSF microbial communities (for example, Acidobacteria, Actinobacteria, OD1 and Chloroflexi; Pinto et al., 2012) are, similarly, undescribed.
Our recent investigation into the microbial communities of several groundwater-fed RSFs (Gülay et al., 2016) revealed an astonishing and consistent abundance and microdiversity of Nitrospira spp. over all other taxa: up to 27% of all amplicons in pre-filters and up to 45% in after-filters were unambiguously identified as Nitrospira of lineages I, II and IV, although most phylotypes belonged to previously undescribed lineages. In addition, the Rhizobiales, Chloracidobacterium, Da023, Acidimicrobiales, Gemmatimonadales and Burkholderiales taxa were abundant, suggesting a central, yet to be identified, metabolic role for those taxa. Here, we apply shotgun metagenomic sequencing on replicate samples from an after-filter of one of the earlier described waterworks, where Nitrospira accounted for 65% of the total amplicon abundance (Gülay et al., 2016), to determine the putative metabolic role of the Nitrospira clade and to elucidate the potential functions of the dominant taxa in investigated RSFs. We focused on two primary questions: can Nitrospira use any other energy source beyond nitrite in the RSFs? And which taxa contribute the central functions in the RSF? The metagenomic data set was assembled and a microbial gene catalog was constructed describing the community metabolic potential. By reassembly of taxonomically clustered contigs, we were able to identify and obtain 14 near-complete draft genomes for members of several abundant taxa described in Gülay et al. (2016). A highly abundant composite Nitrospira genome was identified, which harbored genes for complete ammonium oxidation.
Materials and methods
Sample collection and extraction of DNA
The studied waterworks follows a treatment chain consisting of aeration and two sequential filtration steps. The aerated water flows downward through a first RSF, designed to remove ferrous iron, and then through a second RSF. DNA samples obtained from the study of Gülay et al. (2016) were subjected to shotgun metagenomic sequencing. Briefly, filter material samples were taken from Islevbro waterworks in Denmark and collected using a 60-cm-long core sampler 6 days after filter backwash. Core samples were obtained at three random positions in the filter to obtain biological replicates. Samples were extruded and sliced into five sections aseptically (0–5, 5–15, 15–25, 25–35 and 35–45 cm). DNA extracts from the top section (0–5 cm, ISLTop) were used to represent the uppermost microbial community. For each replicate, a composite DNA sample was generated by mixing equal volumes of DNA extracts of the other core sections (5–45 cm, ISLBulk) to represent the microbial community deeper in the filter. Thus, three samples from the top and three from the bulk of the filter were obtained.
Library preparation, sequencing and de novo assembly
DNA-shearing and library preparation were performed according to the NEXTflex Rapid DNA-Seq Kit, V13.08 (Bioo Scientific, Austin, TX, USA). Briefly, 250 ng genomic DNA was sheared with the Covaris E210 System using 10% duty cycle, intensity of 5, cycles per burst of 200 for 300 s to create 200-bp fragments. The samples were end-repaired and adenylated to produce an A-overhang. Adapters containing unique barcodes were ligated to the DNA. The samples were purified using bead-size selection for range ~300–400 bp with the Agencourt AMPure XP beads (Beckman Coulter, Beverly, MA, USA). The purified DNA libraries were amplified according to the manufacturer’s protocol: initial denaturation, (2 min, 98 °C), followed by 12 cycles of denaturation (98 °C, 30 s), annealing (65 °C, 30 s), extension (72 °C, 1 min) and final extension (72 °C, 4 min). DNA was quantified using NanoDrop ND-1000 UV-VIS Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA), and quality was checked on an Agilent 2100 Bioanalyzer using the High Sensitivity DNA kit (Agilent Technologies, Santa Clara, CA, USA). The DNA libraries were mixed in equimolar ratios. Sequencing was performed as a 100-bp pair-end run on HiSeq 2000 (Illumina Int., San Diego, CA, USA) at BGI (Copenhagen, Denmark). Trimmomatic v0.22 (Bolger et al., 2014) was used to remove adapters and trim the reads (threshold quality=15; minimum length=45). Quality control was carried out using FastQC (Babraham Bioinformatics (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/)). High-quality reads from each sample were assembled into scaftigs using IDBA-UD (Peng et al., 2012) with default parameters. High-quality reads of each sample have been submitted to MG-RAST using the ‘join fastq-formatted paired-reads’ option retaining the nonoverlapping reads. They can be found under accession numbers 4629971.3, 4622590.3, 4629689.3, 4631157.3, 4630162.3 and 4631739.3.
Construction of a non-redundant gene catalog and quantification of reference gene abundance
Gene calling on the assembled scaftigs was performed using the metagenome implementation (‘-p meta’) of Prodigal 2.50 (Hyatt et al., 2010). Predicted genes from all samples (2.85 m in total) were clustered using UCLUST (Edgar, 2010). Any two genes with greater than 90% nucleotide identity were clustered together, resulting in a set of 1 242 515 non-redundant genes. High-quality reads were mapped to the reference gene catalog using Burrows–Wheeler Aligner (Li and Durbin, 2010), BWA-MEM. Mapped reads were subsequently filtered by removing those with a poor alignment (−q30). The remaining mapped reads were used to form an abundance matrix of the number of reads mapped to each gene in every sample. When both read pairs mapped to the same gene, it was counted as a single hit. If each pair mapped to different genes then this is counted as a hit to each gene. The abundance matrix was normalized based on data set size and gene length.
Taxonomic and functional annotation of the gene catalog
Predicted coding sequences were annotated using USEARCH (Edgar, 2010) –ublast against the UniProt database (best hit with E<1e−5, Bitscore>60 and sequence similarity >30%). An amino-acid similarity of 65% or higher was used as a threshold for phylum-level assignment (Li et al., 2014). Clusters of orthologous groups (COG) and non-supervised orthologous groups (NOG) annotations of unassembled sequences were conducted in MG-RAST (E<1e−5, sequence similarity >30%; Meyer et al., 2008) and compared with 25 metagenomes randomly selected from MG-RAST from five pre-selected biomes (Supplementary Table S1).
Binning, composite genome assembly and annotation
For each sample, contigs larger than 1000 nucleotides were clustered into putative taxonomic groups based on pentanucleotide signatures using VizBin (Laczny et al., 2015). Contigs of the resultant clusters were used to recruit sequence reads from the original quality-filtered data set. Recruited reads in each cluster were de novo assembled as described above to construct composite genomes (CGs). Resultant reconstructed CGs were manually evaluated through contig depth and GC content to validate accurate bin segregation. Completeness and potential contamination of each CG was evaluated using 107 essential single-copy genes (Albertsen et al., 2013). When the same draft CG could be reconstructed from several samples the one best assembled, most complete, and with lowest contamination was retained. To improve assembly of higher-abundance organisms, taxonomic clusters were subsampled by segregation based on read depth and were subjected to de novo assembly. The best assemblies were selected based on N50, completeness and taxonomic affiliation of single-copy genes (for further details see Supplementary Information). CGs that contained at least 75% of the single-copy genes were further analyzed. Genomes were identified to appropriate taxonomic levels based on two approaches: (i) A set of 107 essential single-copy genes were blasted (BLASTP, E< 1e−5) against the NCBI-nr database with follow-up analysis in MEGAN (Huson and Weber, 2013) to identify the lowest common ancestor of the blast output, (ii) using the standard settings of PhyloSift (Darling et al., 2014). A taxon was assigned when at least 75% of the identified essential single-copy genes resulted in a concordant taxonomy. For genomic comparison between CGs and reference genomes, the average amino-acid identity (AAI; Konstantinidis and Tiedje, 2005) was calculated based on reciprocal best hits (two-way AAI) between two genomic data sets of proteins using the AAI calculator (http://enve-omics.ce.gatech.edu/aai/). The relative abundance of the organisms represented by the CGs was calculated by mapping the original, quality-filtered reads from each sample against the CG sequence. Gene calling and annotation of CGs was conducted as described above. Furthermore, to confirm protein functional assignment, Kyoto encyclopedia of genes and genomes (KEGG) annotations of the predicted proteins in each CG were obtained using the WebMGA server (Wu et al., 2011). Presence of glycoside hydrolases and extracellular peptidases was evaluated by dbCAN (E<1e−5 and cover fraction >0.4; Yin et al., 2012) and MEROPS (E<1e−10; Rawlings et al., 2014), respectively.
Phylogenetic analysis of ammonium oxidation genes
Predicted amino-acid sequences from the metagenomic samples and CGs were aligned with reference sequences using MUSCLE (Edgar, 2004) or T-Coffee (Notredame et al., 2000). Multiple alignments were manually revised before phylogenetic analysis. Bootstrapped maximum likelihood trees were constructed in MEGA6 using the Jones Taylor Thornton model with 100 replicates (Tamura et al., 2013).
Results and discussion
Metagenomic assembly and taxonomy
We sequenced whole-community DNA for six samples from a RSF (three samples from the top 5 cm of the filter, ISLTop and three composite samples representing filter depths from 5 to 45 cm, ISLBulk), generating an average of 2.6±0.1 Gbp of high-quality, paired-end sequence data per sample. The estimated abundance-weighted average coverage (Rodriguez-R and Konstantinidis, 2014) of the top and bulk metagenomes were 62% and 82%, respectively (Supplementary Figure S1), indicating that the majority of the microbial community was captured in the study. Reads from each data set were de novo assembled, producing an average of 293 391 contigs (Table 1). Prediction of coding sequences resulted in an average of 475 155 putative genes per data set, which were combined across the six samples to a non-redundant gene catalog of 1 242 515 genes. The annotation of the non-redundant gene catalog yielded a total of 768 197 genes with function assigned (Table 1).
Within all of the communities, Bacteria were by far the most abundant domain (ISLBulk, 97.85±0.23; ISLTop, 98.63±0.02). Archaea represented less than 1% of the filter community in both top and bulk samples (ISLBulk, 0.74±0.10; ISLTop, 0.43±0.02), consistent with previous studies examining the microbial communities in rapid gravity filtrations filters (Bai et al., 2013; Wang et al., 2014) and with a previous pyrosequencing analysis of the same community (Gülay et al., 2016). At the phylum level, Proteobacteria and Nitrospirae dominate bulk sample communities with a relative abundance of 18.93%±1.31% and 9.22%±4.72%, respectively; in the top of the filter these phyla were also dominant, with Nitrospirae most abundant (26.08%±0.94%), followed by Proteobacteria (14.47%±1.07%; Figure 1). These results are in-line with other observations of very high Nitrospira abundances (13% to 50% of all community 16S rRNA amplicons or clones) in similar oligotrophic water treatment and distribution systems (Martiny et al., 2005; White et al., 2012; LaPara et al., 2015; Gülay et al., 2016).
Average relative occurence (%) of the most abundant phyla in top 5 cm (ISLTop, outer circle) and lower regions of filter (ISLBulk, inner circle) based on best blast hits of predicted protein sequences from the gene catalog to the Uniprot database (65% amino-acid identity for phylum-level assignment).
ISLBulk samples had higher alpha diversity and evenness than ISLTop samples. The average Shannon index for ISLBulk was 6.02±0.40, whereas for ISLTop samples it was 4.37±0.10. On the other hand, the Pielou evenness was 0.71±0.05 in ISLBulk and 0.52±0.02 in ISLTop. Together, these results indicate a stratified distribution in the filter environment: the top-few centimeters of the filter are dominated by a few phylotypes—potentially involved in the removal of most of the groundwater contaminants (Gülay et al., 2016)—whereas the remainder of the filter contains a higher diversity of less dominant microbes.
The functional potential of the RSF communities
The functional potential of the RSF microbial communities was contrasted with 25 metagenomes from different natural and engineered biomes (Supplementary Table S1) by comparison of COG and NOG categories (Supplementary Figure S2). This analysis showed that the RSF microbial community has relatively high proportions of chaperones and genes involved in post-translational modification and protein turnover (category O) as well as coenzyme transport and metabolism (H). These two categories have been detected to be abundant in oligotrophic ecosystems in comparison with higher nutrient availability ecosystems (Ortiz et al., 2013). The abundance of chaperones may help bacteria to deal with low nutrient availability and have been related to environmental stress tolerance (Storz and Hengge, 2011). The high abundance of secondary metabolism genes (Q) together with the low number of genes associated with transcription (K) in the filter samples is consistent with another study in which these two categories differentiated oligotrophic from copiotrophic microorganisms (Lauro et al., 2009).
The abundance of transposases in the RSF metagenomes was striking: they accounted for 1.18%±0.09% and 1.49%±0.04% of the total annotated genes in the ISLBulk and ISLTop, respectively, substantially higher than in a study of 2137 complete genomes where the average transposase abundance was 0.83% (Aziz et al., 2010). The high number of transposases may be associated with the filter environment, as mobile genetic elements have been found with greater prevalence in surface-attached communities (Stewart, 2013; Madsen et al., 2012). Thus, transposases could provide the means by which phylogenetically distinct bacteria share genetic material, facilitating functional similarities as has been suggested in other studies (Hooper et al., 2009).
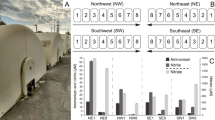
Potential for inorganic electron-donor usage by the microbial community was inferred from the relative abundance of specific marker genes (Figure 2). Ammonium and nitrite oxidation were the dominant potential electron-donor processes in ISLTop. In ISLBulk potentials for the oxidation of ammonium, nitrite, reduced sulfur species, manganese and, to a lesser extent, methane, iron and hydrogen oxidation were more equal in abundance. This is consistent with experimental observations that nitrification is a dominant biogeochemical process in the first few centimeters of these filters (Tatari et al., 2013; Lee et al., 2014). Given the abundance of the ammonium oxidation potential, we investigated the phylogenetic diversity of amoA (as marker gene of ammonium oxidation) including reference amoA sequences and amoA genes recovered from the gene catalog (Figure 3). The RSF metagenome amoA sequences separated into four different clusters. In all, 1% and 7% (ISLBulk and ISLTop, respectively) of amoA genes were related to a cluster of oligotrophic AOB Nitrosomonas spp. including strain Is79A3 and N. oligotropha. Ammonia-oxidizing archaea-associated amoA made up 1% of the amoA in the ISLBulk samples; however, this cluster was not detected in the top of the filter. amoA associated with heterotrophic ammonia oxidizers made up 46% and 12% of all amoA (ISLBulk and ISLTop, respectively). Finally, the majority of the amoA genes (52% of amoA genes of the ISLBulk and 81% in ISLTop) could not be assigned to any known ammonia-oxidizing prokaryote (AOP) clades (40% dissimilar to known AOB at the protein level). Representative pmoA reference sequences were included to reject the possible misclassification of amoA sequences with the marker gene of methane oxidation, as they share substantial sequence similarity (Holmes et al., 1995). Consistent clustering patterns were observed for other ammonium oxidation-related genes: amoB, amoC and hao genes clustered with the characterized AOP clades, as well as an additional novel clade (Supplementary Figure S3).
Log-transformed relative abundances of marker genes for carbon fixation pathways, electron donors and electron acceptors in top 5 cm (ISLTop) and lower regions of filter (ISLBulk). Marker genes used for carbon fixation pathways were RuBisCo (CBB cycle), pyruvate: ferredoxin oxidoreductase (reverse TCA cycle) and 4-hydrocybutyryl-CoA dehydratase (4-hydroxybutyrate cycle).
Phylogenetic reconstruction of amoA and pmoA reference protein sequences with putative amoA sequences recovered from the metagenomes (bold). Relative abundance of putative amoA clusters in the metagenomes is shown with colors corresponding to phylogenetic groups. Bootstrap support greater than 60 is indicated (based on 100 replicates).
In addition to potential electron-donor usage, we also investigated potential electron-acceptor usage (Figure 2). As expected from the predominantly aerobic conditions within the filters, cytochrome c oxidases involved in oxygen reduction accounted for around 80% of genes involved in electron-accepting processes. Genes involved in the reduction of nitrogen oxides (NO3− and NO2−) were also relatively abundant in the gene catalog, showing the capacity of certain members of the community to use alternate electron acceptors under anoxic or micro-oxic conditions.
As the majority of characterized nitrifiers are autotrophic, we examined the diversity and relative abundance of genes for carbon fixation pathways in the gene catalog. The reverse tricarboxylic acid cycle (rTCA) pathway was dominant (Figure 2). The rTCA pathway is not found in typical AOB and Alphaproteobacterial nitrite-oxidizing bacteria (both have the Calvin–Benson–Bassham (CBB) cycle (Badger and Bek, 2007)), but is characteristic of the Nitrospira genus. In fact, all marker genes of the rTCA cycle found in the catalog were taxonomically assigned to Nitrospira spp. Carbon fixation via the CBB cycle was the second most abundant. The CBB genes displayed a wider phylogenetic diversity. The 4-hydroxybutyrate cycle pathway was also detected but was rare (Figure 2). Genes of the acetyl-CoA or 3-hydroxypropionate carbon fixation pathway were not detected in the gene catalog. These results further underline the dominance of Nitrospira spp. among the autotrophic organisms in the examined filters.
Genome reconstruction and identification
Assembled contigs (length>1000 bp) from each sample were used to generate taxonomically restricted bins using pentanucleotide frequency (Laczny et al., 2015; Figure 4 and Supplementary Figure S4). Genomic coverage and GC content were analyzed to validate bin segregation. Fourteen near-complete (>75% of essential genes) draft genomes were reconstructed from the metagenomic data sets (Table 2). Reconstructed genomes were classified as Betaproteobacteria (CG5, CG13 and CG26), Alphaproteobacteria (CG3, CG18 and CG6), Acidobacteria (CG10 and CG15), Nitrospirae (CG24), Gammaproteobacteria (CG7), Gemmatimonadetes (CG33) and Planctomycetes (CG4). Two of the recovered genomes could not be classified to currently defined phyla (CG1 and CG2). Nitrospira CG24 and Acidobacteria CG15 represent bins containing more than one genome based on the presence of multiple single-copy genes (Supplementary Table S2). The difficulty to separate these genomes probably resides in the existence of microdiversity, which is a hindrance to genome reconstruction and segregation from metagenomic data (Wilmes et al., 2009).
Scatter plot of contigs (⩾1 kb) assembled from metagenomic reads from sample Top2. Individual points represent contigs, with point size proportional to the natural log of contig length, and opaqueness proportional to the natural log of contig coverage. The manually placed red polygons highlight selected clusters used for the assembly of CGs as marked. A star shape highlights contigs annotated to contain the amoA (red) and nxrB (yellow) genes.
Composite genomes were genetically compared with their most closely related reference genomes (Supplementary Table S3). On the basis of average AAI, just two of the genomes had values approaching the species level of 85% (Luo et al., 2014). CG7 had an AAI of 78%±17% with Methyloglobulus morosus, and CG26 had an AAI of 87%±15% with Nitrosomonas sp. IS79. This analysis suggests that most of the draft genomes described herein are not closely related to previously sequenced organisms.
The most abundant CG both in ISLTop and ISLBulk samples was Nitrospira CG24, with an average abundance of 29.59% and 9.57%, respectively. CG10, affiliated to Acidobacteria subdivision 4, had a relative read proportion of 2.45% in the top of the filter and 2.43% in bulk samples. Burkholderiales CG13 and Rhizobiales CG3 appeared with greater than 1% abundance in both ISLTop and ISLBulk. Acidobacteria CG15 bin accounted for 5.37% of the reads in ISLBulk and the 0.70% in ISLTop. In addition, Betaproteobacteria CG5 and a poorly classified CG (CG1) were present with greater than 2% abundance in the bulk samples (Table 2).
Composite-genome metabolic potential
The metabolic potential of CGs was examined, focusing on energy and carbon metabolism, including electron donor, electron acceptor and carbon fixation pathways (Supplementary Figure S5).
Ammonium and nitrite oxidation
The near-complete CG26 contained a complete ammonia monooxygenase (amoCAB) operon. However, the gene set lacked hao (hydroxylamine oxidase), possibly because the genome was not complete. The Nitrosomonas bin encoded genes for the CBB cycle. Furthermore, it harbored an aa3-type cytochrome oxidase and nitrite reductase (nirK). The closest relative to CG26 was Nitrosomonas sp. Is79 (AAI of 87%±15%), a strain adapted to low ammonium concentrations (Bollmann et al., 2013).
CG24, a Nitrospira cluster, composed of five genomes (Supplementary Table S2), contained nitrite oxidoreductase (nxr) genes as well as cytochrome bd-type oxidase involved in oxygen reduction. This bin also encoded for nitrite reductase (NO-forming). CG24 harbored genes for the reductive citric acid cycle (rTCA) involved in carbon fixation. In addition, it contained NAD+-dependent formate dehydrogenase and formate transporters, suggesting that it could oxidize formate coupled with the reduction of oxygen, nitrate or nitrite under anoxic conditions, as has been observed in other Nitrospira sp. (Koch et al., 2015). Another feature recently found in Nitrospira, which was also encoded in this bin, is urease which is involved in the hydrolysis of urea into ammonia and carbamate (Koch et al., 2015). CG24 contained urease subunits alpha, beta and gamma. Furthermore, it encoded chlorite dismutase, an enzyme involved in the conversion of chlorite to chloride and molecular oxygen. Surprisingly, Nitrospira CG24 also harbored three complete ammonia monooxygenase (amoCAB) operons. Considering the novelty of this finding, scaffolds were visualized for evenness of mapped read depth to identify potential chimeric regions using Integrative Genome Viewer (Thorvaldsdottir et al., 2013). The visualization revealed homogeneity of read depth with other genes in the same scaffold and with other scaffolds in the same bin (Supplementary Figure S6). These were the most abundant amo genes in the gene catalog; however, they shared low identity with those previously described in classical AOB (and are therefore referred to as atypical-AMOX genes (Figures 3 and 5 and Supplementary Figure S3)). In comparison, typical AOB amo genes (here referred to as typical-AMOX genes) accounted for a low percentage of total amo genes. The same pattern was observed for other genes involved in ammonia oxidation (Supplementary Table S4). To rule out potential contamination during genome binning, depth and GC content of atypical-AMOX genes and typical-AMOX genes were compared with average depth and GC content of Nitrospira CG24 and Nitrosomonas CG26 (Supplementary Figure S7). In both cases, the distribution of atypical-AMOX genes was identical to Nitrospira CG24 (Wilcoxon test, P-value>0.05) but different from Nitrosomonas CG26. The comparison was also performed with other draft genomes, but none of them presented similar distributions (Supplementary Figure S7). To further support these findings, the covariance between the abundance of genes of interest (Supplementary Table S4) in the gene catalog and investigated bins (CG24 and CG26) along the six samples was analyzed. All atypical-AMOX genes statistically correlated with Nitrospira CG24 (R2>0.97–0.99, P-value<0.001) but not with Nitrosomonas CG26 (R2>0.24–0.51, P-value>0.1; Supplementary Figure S7). Together, these results strongly support the finding that Nitrospira CG24 contains genes encoding for ammonia oxidation.
Nitrospira CG24 was compared with two other Nitrospira genomes based on average AAI. CG24 has 70% AAI with N. moscoviensis and 65% with ‘Candidatus Nitrospira defluvii’ (Supplementary Figure S8). These AAIs are far from the 85% species-level cutoff value but would comply with genus-level similarity (60–80%; Luo et al., 2014). Thus, CG24 would represent a different species of the Nitrospira genus. An amo operon-containing contig from CG24 was compared with amo operons and flanking regions from Nitrosomonas sp. IS79 (the reference genome most closely related to the AOB found in this RSF) and an AOB reference genome from another genus (Nitrosospira multiformis). Predicted amino-acid sequence identities were highest for amoC, which shared 70% and 71% identity with Nitrosomonas and Nitrosospira, respectively. Identity was lower for amoA and amoB, ranging from 56% to 59%, respectively (Figure 5). This is in contrast to the amo AAI between Nitrosomonas and Nitrosospira spp., which are upwards of 80%. These results suggest that, whereas these proteins still share a common function, they are derived from a more distantly related clade. Examination of the neighboring regions of the amo operon in CG24 revealed no similarity with neighboring regions in the Nitrosomonas and Nitrosospira genomes (Figure 5).
Methane oxidation
A reconstructed genome CG7 contained particulate methane monooxygenase (pmo) and methanol dehydrogenase (mdh). This draft genome possessed the electron transport chain as well as aa3-type and bd-type cytochrome oxidases. In relation to carbon metabolism, it encoded for the TCA cycle, pentose phosphate pathway and Entner–Doudoroff pathway. On the basis of this genetic content, CG7 likely thrives on residual methane present in the water.
Oxidation of reduced sulfur, manganese and iron
A genomic bin (CG13) associated with Burkholderiales encoded dissimilatory sulfite reductases (dsrA and dsrB) for sulfide oxidation as well as adenylylsulfate reductases (aprA and aprB) and sulfate adenylyltransferase (sat) for complete oxidation of sulfite to sulfate. CG13 carried genes for near-complete CBB cycle, including ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO) gene. CG13 contained cytochrome cbb3-type oxidase, periplasmic nitrate reductase (napA) and nitrite reductase (NO-forming). Furthermore, this genome contained mtrAB genes with homology (amino-acid sequence identity from 29% to 55%) to those found in lithotrophic iron-oxidizing bacteria (Jiao and Newman, 2007; Beckwith et al., 2015). Thus, CG13 could be involved in the oxidation of ferrous iron, although the presence of mtrABC (periplasmic and outer membrane cytochromes) could also indicate electron transfer to minerals (Weber et al., 2006; Gülay et al., 2014). The CG13 bin also contained multicopper oxidases homologous (60–75% at the protein level) to those found in the known manganese oxidizers Leptothrix cholodnii str. SP-6 and Pedomicrobium sp. ACM 3067, suggesting a potential capacity for manganese oxidation.
Rhizobiales CG3 encoded for two multicopper oxidases with sequence identity (59% to 75% at protein level) related to those found in the manganese oxidizer Aurantimonas manganoxydans SI85-9A1 (Dick et al., 2008). CG3 encoded for cytochrome cbb3-type oxidase and nitrate reductase (narG). Rhizobiales CG3 also harbored genes of the TCA cycle, pentose phosphate pathway (non-oxidative phase) and beta oxidation pathway. Moreover, this genetic bin possessed extracellular peptidase and glycoside hydrolases involved in protein and carbohydrate metabolism, respectively (Supplementary Figure S9). These genetic features together with the absence of any carbon fixation pathway suggest the potential use of Mn2+ as an energy source, coupled to organic compound degradation as a carbon source. This chemolithoheterotrophic behavior has previously been observed in other manganese oxidizers (Francis et al., 2001; Templeton et al., 2005).
Heterotrophy/organic carbon degradation
To explore the capability of the microbial community to degrade organic carbon present in the influent water as well as products derived from microbial growth and decay, we screened reconstructed genomes for carbon metabolism pathways, glycoside hydrolases, extracellular peptidases and sugar and amino-acid transporters. Most of CGs encoded for complete or near-complete TCA cycle, beta oxidation and glycolysis pathways. Partial or complete pentose phosphate pathways were also present in most of the CGs. On the other hand, the Entner–Doudoroff pathway was less abundant, being complete, or mostly complete in only four of the CGs. In relation to glycoside hydrolases, most of the CGs contained genes for the degradation of cellulose, peptidoglycan, N-acetylglucosamine and starch (Supplementary Figure S9). Glycoside hydrolases were particularly abundant and diverse in both Acidobacteria CG15 and CG10 and Planctomycetaceae CG4. Reconstructed genomes CG1 and Sphingomonas CG18 had glycoside hydrolases for the degradation of hemicellulose. Furthermore, extracellular peptidases were present in most of CGs. CG1, Gemmatimonadetes CG33 and Acidobacteria CG10 possessed several genes of different families of extracellular peptidases potentially involved in the decomposition of bacterial cell walls and proteins (Supplementary Figures S8 and S9). These observations, together with the presence of sugar (CG10, CG2, CG1 and CG4) and amino-acid (CG15 and CG3) transporters in several of the reconstructed genomes, indicate the potential of several of the CGs to degrade and uptake organic carbon in this system. In particular, the high diversity of genes involved in carbohydrate degradation and uptake in CG1 and both Acidobacteria CG15 and CG10 combined with their high abundance suggests that these organisms may have a key role in organic carbon flow in the filter.
Predicted metabolic and geochemical model
On the basis of metabolic reconstruction of CGs and the chemical characteristics of the RSF, a metabolic and geochemical model of the RSF community is presented (Figure 6). Reconstructed genome Methylococcaceae CG7 could be involved in methane oxidation, as methane may not be completely stripped during aeration. Burkholderiales CG13 harbors the metabolic capabilities to realize complete oxidation of sulfide to sulfate and may be also associated with autotrophic manganese and iron oxidation. The ability to oxidize Fe2+ and Mn2+ indiscriminately has also been observed in other bacteria (Gouzinis et al., 1998). Manganese could potentially also be oxidized by the heterotrophic Rhizobiales CG3. A large number of CGs encoded heterotrophic metabolisms, which are likely involved in the degradation of organic molecules present in the groundwater or the decay and metabolic by-products of autotrophic biomass. Ammonium in the filter could be oxidized by the typical autotrophic AOB Nitrosomonas CG26. The nitrite produced by CG26 could then be utilized by Nitrospira CG24. However, given its metabolic potential, the dominance of Nitrospira CG24 and the high abundance of Nitrospira-AMOX genes compared with other AMO genes, Nitrospira CG24 might mediate complete ammonium oxidation in this system. This would be in agreement with the suggestion that a microorganism that completely oxidizes ammonia to nitrate exists (Costa et al., 2006). The Nitrospira abundance observed in this study is consistent with earlier pyrosequencing and quantitative PCR-based analysis of the same system, where abundances of up to 65% and 18%, respectively, were measured, and Nitrospira 16S rRNA gene sequences were often up to two orders of magnitude more abundant than Nitrosospira and Nitrosomonas 16S rRNA sequences combined (Gülay et al., 2016). Further, ammonium oxidation Nitrospira activity may help to explain the high abundance of Nitrospira and the unusual Nitrospira/AOB ratios observed in other rapid gravity filters (Feng et al., 2012; Albers et al., 2015; Cai et al., 2015; LaPara et al., 2015). Whereas Nitrospira may be the primary ammonium oxidizer, the high abundance of heterotrophic amoA sequences, especially in the bulk samples, suggests that heterotrophic ammonium oxidation may also occur in the RSFs.
During the revision of this manuscript, evidence of complete ammonia oxidization (comammox) by Nitrospira organisms was described by others (Daims et al., 2015; van Kessel et al., 2015). Metagenomic evidence of comammox Nitrospira in a drinking water distribution system was reported by Pinto et al. (2016). Average AAI of CG24 was compared with the recently published comammox genomes: Nitrospira CG24 would represent a different species within the Nitrospira genus (AAI of 68%±16% with ‘Candidatus Nitrospira inopinata’, 71±16% ‘Candidatus Nitrospira nitrosa’, 73±17% ‘Candidatus Nitrospira nitrificans’ and 75±18% with the Nitrospira sp. genome from a drinking-water system) sharing the metabolic potential for complete ammonia oxidation. As CG24 consisted of more than a single genome, we investigated the heterogeneity within CG24. We examined the average number of essential single-copy genes and amo operons (one in recently published comammox genomes; Daims et al., 2015; Pinto et al., 2016; van Kessel et al., 2015) present in CG24. The Nitrospira bin CG24 has an average of 4.7 essential single-copy genes and three amo operons. On the basis of this, we predict that approximately three out of the five potential genomes represented by CG24 are complete ammonia oxidizers. Although we were unable to assign the comammox Nitrospira in this system to a specific lineage, previous analysis of this system revealed that the majority of Nitrospira sequences in this filter belong to novel lineages (Gülay et al., 2016). Thus, we predict that the comammox Nitrospira present in this system may belong to a novel lineage, and not to lineage II as for other described comammox Nitrospira. Further sequencing of the comammox Nitrospira in this system is required to elucidate the lineage of the comammox Nitrospira described herein. In conclusion, our metagenomic survey provides insights into the metabolic capabilities of the microbial communities in an oligotrophic engineered system. Genome reconstructions allowed us to predict the roles of dominant community members in nitrogen, carbon, manganese, iron, methane and sulfur cycling—the main biological processes occurring in the sand filters. The results of our analysis point toward the novel metabolic capability of complete ammonia oxidation in the genus Nitrospira, which was highly abundant and ubiquitous in the investigated RSFs.
The metabolic and geochemical model of the RSF microbial communities can be used to inform further modeling efforts with the ultimate goal of enhancing contaminant removal and improving the design or function of RSFs. Improved understanding of the function of novel taxa in RSFs may be of considerable utility in restoring malfunctioning filters where common microbiological data have, thus far, been insufficient to explain the observed patterns of contaminant removal. Transcriptomic analysis of the RSFs will help to validate the predictions made in this study and to elucidate the ecological importance of the various taxa and functions in RSFs.
References
Albers CN, Ellegaard-Jensen L, Harder CB, Rosendahl S, Knudsen BE, Ekelund F et al. (2015). Groundwater chemistry determines the prokaryotic community structure of waterworks sand filters. Environ Sci Technol 49: 839–846.
Albertsen M, Hugenholtz P, Skarshewski A, Nielsen KL, Tyson GW, Nielsen PH . (2013). Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat Biotechnol 31: 533–538.
Aziz RK, Breitbart M, Edwards RA . (2010). Transposases are the most abundant, most ubiquitous genes in nature. Nucleic Acids Res 38: 4207–4217.
Badger MR, Bek EJ . (2007). Multiple Rubisco forms in proteobacteria: their functional significance in relation to CO2 acquisition by the CBB cycle. J Exp Bot 59: 1525–1541.
Bai Y, Liu R, Liang J, Qu J . (2013). Integrated metagenomic and physiochemical analyses to evaluate the potential role of microbes in the sand filter of a drinking water treatment system. PLoS One 8: e61011.
Beckwith CR, Edwards MJ, Lawes M, Shi L, Butt JN, Richardson DJ et al. (2015). Characterization of MtoD from Sideroxydans lithotrophicus: a cytochrome c electron shuttle used in lithoautotrophic growth. Front Microbiol 6: 332.
Bolger AM, Lohse M, Usadel B . (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30: 2114–2120.
Bollmann A, Sedlacek CJ, Norton J, Laanbroek HJ, Suwa Y, Stein LY et al. (2013). Complete genome sequence of Nitrosomonas sp. Is79, an ammonia oxidizing bacterium adapted to low ammonium concentrations. Stand Genomic Sci 7: 469–482.
Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A et al. (2015). Unusual biology across a group comprising more than 15% of domain bacteria. Nature 523: 208–211.
Bryant DA, Costas AMG, Maresca JA, Chew AGM, Klatt CG, Bateson MM et al. (2007). Candidatus Chloracidobacterium thermophilum: an aerobic phototrophic Acidobacterium. Science 317: 523–526.
Cai Y, Li D, Liang Y, Luo Y, Zeng H, Zhang J . (2015). Effective start-up biofiltration method for Fe, Mn, and ammonia removal and bacterial community analysis. Bioresour Technol 176: 149–155.
Carrión O, Curson ARJ, Kumaresan D, Fu Y, Lang AS, Mercadé E et al. (2015). A novel pathway producing dimethylsulphide in bacteria is widespread in soil environments. Nat Commun 6: 6579.
Cerrato JM, Falkinham JO, Dietrich AM, Knocke WR, McKinney CW, Pruden A . (2010). Manganese-oxidizing and -reducing microorganisms isolated from biofilms in chlorinated drinking water systems. Water Res 44: 3935–3945.
Costa E, Pérez J, Kreft J-U . (2006). Why is metabolic labour divided in nitrification? Trends Microbiol 14: 213–219.
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M et al. (2015). Complete nitrification by Nitrospira bacteria. Nature 528: 504–509.
Darling AE, Jospin G, Lowe E, Matsen FA, Bik HM, Eisen JA . (2014). PhyloSift: phylogenetic analysis of genomes and metagenomes. PeerJ 2: e243.
de Vet WWJM, Dinkla IJT, Abbas BA, Rietveld LC, van Loosdrecht MCM . (2012). Gallionella spp. in trickling filtration of subsurface aerated and natural groundwater. Biotechnol Bioeng 109: 904–912.
Dick GJ, Podell S, Johnson HA, Rivera-Espinoza Y, Bernier-Latmani R, McCarthy JK et al. (2008). Genomic insights into Mn(II) oxidation by the marine alphaproteobacterium Aurantimonas sp. strain SI85-9A1. Appl Environ Microbiol 74: 2646–2658.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM et al. (2010). Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464: 543–548.
Feng S, Xie S, Zhang X, Yang Z, Ding W, Liao X et al. (2012). Ammonium removal pathways and microbial community in GAC-sand dual media filter in drinking water treatment. J Environ Sci (China) 24: 1587–1593.
Francis CA, Co EM, Tebo BM . (2001). Enzymatic manganese(II) oxidation by a marine alpha-proteobacterium. Society 67: 4024–4029.
Gouzinis A, Kosmidis N, Vayenas DV, Lyberatos G . (1998). Removal of Mn and simultaneous removal of NH3, Fe and Mn from potable water using a trickling filter. Water Res 32: 2442–2450.
Gülay A, Musovic S, Alberchtsen H-J, Al-Soud WA, Sørensen S, Smets BF . (2016). Ecological patterns, diversity and core taxa of microbial communities in groundwater- fed rapid gravity filters. ISME J e-pub ahead of print 8 March 2016; doi:10.1038/ismej.2016.16.
Gülay A, Tatari K, Musovic S, Mateiu RV, Albrechtsen H-J, Smets BF . (2014). Internal porosity of mineral coating supports microbial activity in rapid sand filters for groundwater treatment. Appl Environ Microbiol 80: 7010–7020.
Holmes AJ, Costello A, Lidstrom ME, Murrell JC . (1995). Evidence that particulate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol Lett 132: 203–208.
Hooper SD, Mavromatis K, Kyrpides NC . (2009). Microbial co-habitation and lateral gene transfer: what transposases can tell us. Genome Biol 10: R45.
Huson DH, Weber N . (2013). Microbial community analysis using MEGAN. Methods Enzymol 531: 465–485.
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ . (2010). Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11: 119.
Jiao Y, Newman DK . (2007). The pio operon is essential for phototrophic Fe(II) oxidation in Rhodopseudomonas palustris TIE-1. J Bacteriol 189: 1765–1773.
Koch H, Lücker S, Albertsen M, Kitzinger K, Herbold C, Spieck E et al. (2015). Expanded metabolic versatility of ubiquitous nitrite-oxidizing bacteria from the genus Nitrospira. Proc Natl Acad Sci USA 112: 11371–11376.
Konstantinidis KT, Tiedje JM . (2005). Towards a genome-based taxonomy for prokaryotes. J Bacteriol 187: 6258–6264.
Laczny CC, Sternal T, Plugaru V, Gawron P, Atashpendar A, Margossian H et al. (2015). VizBin - an application for reference-independent visualization and human-augmented binning of metagenomic data. Microbiome 3: 1.
LaPara TM, Hope Wilkinson K, Strait JM, Hozalski RM, Sadowksy MJ, Hamilton MJ . (2015). The bacterial communities of full-scale biologically active, granular activated carbon filters are stable and diverse and potentially contain novel ammonia-oxidizing microorganisms. Appl Environ Microbiol 81: 6864–6872.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S et al. (2009). The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci USA 106: 15527–15533.
Lautenschlager K, Hwang C, Ling F, Liu W-T, Boon N, Köster O et al. (2014). Abundance and composition of indigenous bacterial communities in a multi-step biofiltration-based drinking water treatment plant. Water Res 62: 40–52.
Lee CO, Boe-Hansen R, Musovic S, Smets BF, Albrechtsen H-J, Binning P . (2014). Effects of dynamic operating conditions on nitrification in biological rapid sand filters for drinking water treatment. Water Res 64C: 226–236.
Li H, Durbin R . (2010). Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26: 589–595.
Li J, Jia H, Cai X, Zhong H, Feng Q, Sunagawa S et al. (2014). An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32: 834–841.
Li X, Chu Z, Liu Y, Zhu M, Yang L, Zhang J . (2013). Molecular characterization of microbial populations in full-scale biofilters treating iron, manganese and ammonia containing groundwater in Harbin, China. Bioresour Technol 147: 234–239.
Luo C, Rodriguez-R LM, Konstantinidis KT . (2014). MyTaxa: An advanced taxonomic classifier for genomic and metagenomic sequences. Nucleic Acids Res 42: 1–12.
Madsen JS, Burmølle M, Hansen LH, Sørensen SJ . (2012). The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol Med Microbiol 65: 183–195.
Martiny AC, Albrechtsen H, Arvin E, Molin S . (2005). Identification of bacteria in biofilm and bulk water samples from a nonchlorinated model drinking water distribution system: detection of a large nitrite-oxidizing population associated with Nitrospira spp. Appl Environ Microbiol 71: 8611–8617.
Meyer F, Paarmann D, D’Souza M, Olson R, Glass E, Kubal M et al. (2008). The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9: 386.
Mouchet P . (1992). From conventional to biological removal of iron and manganese in France. J Am Water Work Assoc 84: 158–167.
Nitzsche KS, Weigold P, Lösekann-Behrens T, Kappler A, Behrens S . (2015). Microbial community composition of a household sand filter used for arsenic, iron, and manganese removal from groundwater in Vietnam. Chemosphere 138: 47–59.
Nobu MK, Narihiro T, Rinke C, Kamagata Y, Tringe SG, Woyke T et al. (2015). Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J 9: 1710–1722.
Notredame C, Higgins DG, Heringa J . (2000). T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302: 205–217.
Ortiz M, Legatzki A, Neilson JW, Fryslie B, Nelson WM, Wing RA et al. (2013). Making a living while starving in the dark: metagenomic insights into the energy dynamics of a carbonate cave. ISME J 8: 478–491.
Peng Y, Leung HCM, Yiu SM, Chin FYL . (2012). IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28: 1420–1428.
Pinto AJ, Marcus DN, Ijaz Z, Bautista-de los Santos QM, Dick GJ, Raskin L . (2016). Metagenomic evidence for the presence of comammox Nitrospira-like bacteria in a drinking water system. mSphere 1: e00054–15.
Pinto AJ, Xi C, Raskin L . (2012). Bacterial community structure in the drinking water microbiome is governed by filtration processes. Environ Sci Technol 46: 8851–8859.
Rawlings ND, Waller M, Barrett AJ, Bateman A . (2014). MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res 42: D503–D509.
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng J-F et al. (2013). Insights into the phylogeny and coding potential of microbial dark matter. Nature 499: 431–437.
Rittmann BE . (2006). Microbial ecology to manage processes in environmental biotechnology. Trends Biotechnol 24: 261–266.
Rodriguez-R LM, Konstantinidis KT . (2014). Estimating coverage in metagenomic data sets and why it matters. ISME J 8: 2349–2351.
Schleper C, Jurgens G, Jonuscheit M . (2005). Genomic studies of uncultivated archaea. Nat Rev Microbiol 3: 479–488.
Søgaard E . (2001). Conditions for biological precipitation of iron by Gallionella ferruginea in a slightly polluted ground water. Appl Geochemistry 16: 1129–1137.
Stewart FJ . (2013). Where the genes flow. Nat Geosci 6: 688–690.
Storz G, Hengge R . (2011) Bacterial Stress Responses. ASM Press: Washington, DC, USA.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S . (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30: 2725–2729.
Tatari K, Smets BF, Albrechtsen H-J . (2013). A novel bench-scale column assay to investigate site-specific nitrification biokinetics in biological rapid sand filters. Water Res 47: 6380–6387.
Templeton AS, Staudigel H, Tebo BM . (2005). Diverse Mn(II)-oxidizing bacteria isolated from submarine basalts at Loihi Seamount. Geomicrobiol J 22: 127–139.
Thorvaldsdottir H, Robinson JT, Mesirov JP . (2013). Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14: 178–192.
van der Wielen PWJJ, Voost S, van der Kooij D . (2009). Ammonia-oxidizing bacteria and archaea in groundwater treatment and drinking water distribution systems. Appl Environ Microbiol 75: 4687–4695.
van Kessel MAHJ, Speth DR, Albertsen M, Nielsen PH, Op den Camp HJM, Kartal B et al. (2015). Complete nitrification by a single microorganism. Nature 528: 555–559.
Wang H, Narihiro T, Straub AP, Pugh CR, Tamaki H, Moor JF et al. (2014). MS2 bacteriophage reduction and microbial communities in biosand filters. Environ Sci Technol 48: 6702–6709.
Weber KA, Achenbach LA, Coates JD . (2006). Microorganisms pumping iron: anaerobic microbial iron oxidation and reduction. Nat Rev Microbiol 4: 752–764.
Wexler M, Bond PL, Richardson DJ, Johnston AWB . (2005). A wide host-range metagenomic library from a waste water treatment plant yields a novel alcohol/aldehyde dehydrogenase. Environ Microbiol 7: 1917–1926.
White CP, Debry RW, Lytle DA . (2012). Microbial survey of a full-scale, biologically active filter for treatment of drinking water. Appl Environ Microbiol 78: 6390–6394.
Wilmes P, Simmons SL, Denef VJ, Banfield JF . (2009). The dynamic genetic repertoire of microbial communities. FEMS Microbiol Rev 33: 109–132.
Wu S, Zhu Z, Fu L, Niu B, Li W . (2011). WebMGA: a customizable web server for fast metagenomic sequence analysis. BMC Genomics 12: 444.
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y . (2012). dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40: W445–W451.
Acknowledgements
We thank K Amoako, MD Dalgaard, and BJR Cockx for supporting sample collection, library preparation and NG sequencing, and graphics. This research was financially supported by MERMAID (An initial training network funded by the People Programme - Marie Skłodowska-Curie Actions- of the European Union's Seventh Framework Programme FP7/2007-2013/ under REA grant agreement n°607492).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Author contributions
The original study design was conceived by AG, AP, TSP and BFS. Sampling and DNA extraction were performed by AG. AP performed bioinformatic analyses, supported by SJF, TSP and SR. AP and SJF lead interpretation of the results. AP drafted the manuscript; all authors contributed to its revision and completion, and approved the final submission.
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Palomo, A., Jane Fowler, S., Gülay, A. et al. Metagenomic analysis of rapid gravity sand filter microbial communities suggests novel physiology of Nitrospira spp.. ISME J 10, 2569–2581 (2016). https://doi.org/10.1038/ismej.2016.63
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.63
This article is cited by
-
Microbial tapestry of the Shulgan-Tash cave (Southern Ural, Russia): influences of environmental factors on the taxonomic composition of the cave biofilms
Environmental Microbiome (2023)
-
Net-spinning caddisflies create denitrifier-enriched niches in the stream microbiome
ISME Communications (2023)
-
A metagenomic catalog for exploring the plastizymes landscape covering taxa, genes, and proteins
Scientific Reports (2023)
-
Ecological distribution and function of comammox Nitrospira in the environment
Applied Microbiology and Biotechnology (2023)
-
Genomic profiling of Nitrospira species reveals ecological success of comammox Nitrospira
Microbiome (2022)