Abstract
Members of the candidate phylum Hyd24-12 are globally distributed, but no genomic information or knowledge about their morphology, physiology or ecology is available. In this study, members of the Hyd24-12 lineage were shown to be present and abundant in full-scale mesophilic anaerobic digesters at Danish wastewater treatment facilities. In some samples, a member of the Hyd24-12 lineage was one of the most abundant genus-level bacterial taxa, accounting for up to 8% of the bacterial biomass. Three closely related and near-complete genomes were retrieved using metagenome sequencing of full-scale anaerobic digesters. Genome annotation and metabolic reconstruction showed that they are Gram-negative bacteria likely involved in acidogenesis, producing acetate and hydrogen from fermentation of sugars, and may play a role in the cycling of sulphur in the digesters. Fluorescence in situ hybridization revealed single rod-shaped cells dispersed within the flocs. The genomic information forms a foundation for a more detailed understanding of their role in anaerobic digestion and provides the first insight into a hitherto undescribed branch in the tree of life.
Similar content being viewed by others
Introduction
Production of methane by anaerobic digestion (AD) is widely used to convert organic waste into biogas and forms an important part of the transition from fossil fuel to sustainable energy production. The AD process is divided into four sequential steps that are performed by specialized microbes: hydrolysis, fermentation (acidogenesis), acetogenesis (dehydrogenation) and methanogenesis (acetoclastic or hydrogenotrophic) (Angenent et al., 2004). Hence, the overall function, stability and efficiency of the AD process are dependent on tightly coupled synergistic activities of the complex microbial communities (Schink, 1997; Weiland, 2010). However, the microbial communities in AD are still poorly understood, and relatively little is known about their diversity and function (Chouari et al., 2005; Werner et al., 2011; Sundberg et al., 2013; De Vrieze et al., 2015). In addition, most of the microorganisms have no pure culture representatives, and, given the synergistic interactions of members of the community, a reductionist approach to understand the ecology of the system is not possible (Kaeberlein et al., 2002; Fuhrman et al., 2015).
The AD environment also harbours extensive diversity of previously uncharacterized bacterial phyla, often known only by their 16S rRNA gene sequence, making it an ideal environment for the study of novel bacterial lineages (Guermazi et al., 2008; Pelletier et al., 2008; Limam et al., 2014; Sekiguchi et al., 2015). New developments in single-cell genomics and metagenomics have in recent years provided a glimpse into the ecology and evolution of many novel candidate phyla (Dinis et al., 2011; Albertsen et al., 2013; Rinke et al., 2013; Brown et al., 2015; Nobu et al., 2015; Sekiguchi et al., 2015). The genomes have enabled construction of metabolic models that attempt to explain the physiology of these organisms in detail. The genome-based models form the basis of more extensive investigations, such as in situ single-cell characterization, metatranscriptomics and proteomics (Koch et al., 2014).
In this study, extensive 16S rRNA gene amplicon sequencing was used to screen anaerobic digesters for the presence of members of the Hyd24-12 lineage, which remains one of the few known candidate phyla for which no genomic information is available with nothing known about their morphology, physiology or ecology (Rinke et al., 2013). Selected samples were subjected to metagenome sequencing and used for retrieval of three near-complete genomes of Hyd24-12 through differential coverage binning. The genomes were used for detailed metabolic reconstruction and design of oligonucleotide probes for the first in situ visualization of these hitherto unrecognized players in AD.
Materials and methods
Sample collection and storage
A total of 306 biomass samples were obtained from 29 anaerobic digesters at 17 Danish wastewater treatment facilities (see Supplementary Table S1). Most digesters were mesophilic (22), whereas 7 were thermophilic. A volume of 50 ml was sampled, homogenized and stored as 2 ml aliquots at −80 °C for DNA extraction.
For fluorescence in situ hybridization (FISH) analyses, diluted biomass samples (1:4 in 1 × phosphate-buffered saline) were fixed with 4% (w/v) paraformaldehyde and stored in 50% (v/v) ethanol/1 × phosphate-buffered saline solution at −20 °C, as previously described by Daims et al. (2005).
DNA extraction
DNA was extracted from anaerobic digester sludge using the FastDNA Spin kit for soil (MP Biomedicals, Santa Ana, CA, USA), following the standard protocol except for four times increased bead beating duration and a sludge input volume of 50 μl. These digester-sample-specific modifications to the protocol were found to provide the best trade-off between DNA yield/biomass and DNA integrity (Supplementary Figure S4).
Community profiling with 16S rRNA gene amplicon sequencing
Bacterial community profiling was carried out as recommended by Albertsen et al. (2015). The bacterial primers used were 27F (AGAGTTTGATCCTGGCTCAG) (Lane, 1991) and 534R (ATTACCGCGGCTGCTGG) (Muyzer et al., 1993), which amplify a DNA fragment of ~500 bp of the 16S rRNA gene (variable V1–V3 region). PCR amplification was performed using 1 × Platinum High fidelity buffer, 400 pm dNTP, 1.5 mm MgSO4, 2 mU Platinum Taq DNA Polymerase High Fidelity, 5 μm illumina barcoded V1–V3 adaptor mix (see Supplementary Data 1), and 10 ng template DNA. PCR conditions were 95 °C for 2 min, 30 cycles of 95 °C for 20 s, 56 °C for 30 s, 72 °C for 60 s, and a final step of elongation at 72 °C for 5 min. PCR products were purified using Agencourt AmpureXP (Beckman Coulter, Brea, CA, USA) with a ratio of 0.8 bead solution/PCR solution. Barcoded amplicons were pooled and paired-end sequenced on the Illumina MiSeq platform (v3 chemistry, 2 × 300 bp). The paired-end reads were trimmed using trimmomatic (v. 0.32) (Bolger et al., 2014) and then merged using FLASH (v. 1.2.11) (Magoč and Salzberg, 2011). The reads were screened for potential PhiX contamination using USEARCH (v. 7.0.1090) (Edgar, 2010). The reads were clustered into operational taxonomic units (OTUs, sequence identity ⩾97%) using USEARCH and subsequently classified using the RDP classifier (Wang et al., 2007) with the MiDAS database (v. 1.20) (McIlroy et al., 2015). Further processing was carried out in the R environment (v. 3.1.2) using the ampvis package (Albertsen et al., 2015) (v. 1.24.0), which wraps a number of packages including the phyloseq package (v. 1.8.2) (McMurdie and Holmes, 2013), ggplot2 (v. 1.0.1), reshape2 (v. 1.4.1) (Wickham, 2007), dplyr (v. 0.4.2), vegan (v. 2.3-0), knitr (v. 1.10.5), Biostrings (v. 2.36.1), data.table (v. 1.9.4), DESeq2 (v. 1.8.1) (Love et al., 2014), ggdendro (v. 0.1–15) and stringr (v. 1.0.0). The samples were subsampled to an even depth of 10 000 reads per sample, and the fraction of reads classified as Hyd24-12 was obtained. The survey data are available at the SRA with the accession IDs ERS861217-ERS861224.
In silico analysis of Hyd24-12 source locations
The Genbank IDs of the sequences classified as Hyd24-12 in SILVA (v. 121, 1982 sequences in total) (Quast et al., 2013) were used to download the corresponding Genbank files. The fields ‘isolation source’ and ‘PUBMED’ were extracted to classify the sequences as originating from either engineered or natural systems.
Metagenome sequencing, assembly and binning
Illumina TruSeq DNA PCR free libraries were prepared for DNA extracts from three of the mesophilic digesters (Supplementary Table S1) according to the manufacturer’s protocol and paired-end sequenced on the Illumina HiSeq 2000 platform (2 × 150 bp) and Illumina MiSeq platform (v3 chemistry, 2 × 300 bp). The metagenomic assembly and binning process was carried out as described by Albertsen et al. (2013) and detailed at ‘madsalbertsen.github.io/mmgenome/’. Unmerged reads were quality-trimmed and filtered using default settings in CLC Genomics Workbench (v. 7.5.1; CLC Bio, Aarhus, Denmark). The metagenomic reads were assembled separately for each plant using default settings in CLC Genomics Workbench. Reads were mapped to the assemblies using default settings in CLC Genomics Workbench. The assemblies and mappings were exported as .fasta and .sam files, respectively. The exported files and the mmgenome workflow script ‘data.generation.2.1.0.sh’ were used to generate the files necessary for the binning process. 16S rRNA gene sequences were extracted from the assemblies using ‘rRNA.sh’ and classified using the SINA Alignment service (SILVA v 121) (Pruesse et al., 2012); essential genes were called using Prodigal (Hyatt et al., 2010). Binning was carried out using differential coverage binning in the R environment (v. 3.1.2, R Core Team, 2016) using the R package ‘mmgenome’ (github.com/MadsAlbertsen/mmgenome v. 0.4.1) (Albertsen et al., 2013). The genome bins were checked for completeness, essential single copy genes and coverage distribution using CheckM (v. 0.9.7) (Parks et al., 2015) and the metrics in the mmgenome package. Average nucleotide identity between the genome bins was calculated using JSpecies (Richter and Rosselló-Móra, 2009), and CRISPR arrays were identified with CRT (v. 1.1) (Bland et al., 2007). The genome sequence data have been submitted to DDBJ/EMBL/GenBank databases under accession numbers LKHB00000000, LKHC00000000 and LKHD00000000.
Genome sequence-based phylogenetic analysis
The genomes were placed within the reference genome tree of CheckM (Parks et al., 2015) (v. 0.9.7) and subsequently visualized in ARB (Ludwig et al., 2004).
Phylogeny of the 16S rRNA gene and FISH probe design
Phylogenetic analysis and FISH probe design were performed with the ARB software package (Ludwig et al., 2004). Potential probes were assessed in silico with the mathFISH software (Yilmaz et al., 2011) for hybridization efficiencies of target and potentially weak, non-target matches (Yilmaz et al., 2011). Unlabelled helper probes (Fuchs et al., 2000) were designed for calculated inaccessible regions. Unlabelled competitor probes were designed for single-base mismatched non-target sequences (Manz et al., 1992). The Ribosomal Database Project (RDP) PROBE MATCH function (Cole et al., 2009) was used to identify non-target sequences with indels (McIlroy et al., 2011). Probe validation and optimization were based on generated formamide dissociation curves (Daims et al., 2005), where average relative fluorescent intensities, of at least 50 cells calculated with ImageJ software (National Institutes of Health, New York, NY, USA), were measured for varied hybridization buffer formamide concentrations in increments of 5% (v/v) over a range of 5–50% (v/v) (data not shown). Where available, weak base mismatch non-target axenic cultures were used for probe optimization, otherwise full-scale anaerobic digester sludge was used (Table 1).
FISH
FISH was performed essentially as described by Daims et al. (2005). Probes were applied, with recommended competitors and helpers, at the stringency conditions given in Table 1 or their original publications. The NON-EUB nonsense probe was used as a negative hybridization control (Wallner et al., 1993). Oligonucleotide probes were labelled on both the 3′ and 5′ ends with either 5(6)-carboxyfluorescein-N-hydroxysuccinimide ester (FLUOS) or with the sulphoindocyanine dyes (Cy3 and Cy5) (DOPE-FISH) (Stoecker et al., 2010). Microscopic analysis was performed with an Axioskop epifluorescence microscope (Carl Zeiss, Oberkochen, Germany).
Genome analysis
Genome annotation was performed in the ‘MicroScope’ annotation pipeline (Vallenet et al., 2009, 2013). Automatic annotations were manually curated for all genes described using the integrated bioinformatics tools and the proposed annotation rules, which include an amino acid identity of at least 40% to classify homologues and an identity of at least 25% with the support of conserved domains to determine putative homologues (Vallenet et al., 2009, 2013). The set of bioinformatics tools includes BlastP (Altschul et al., 1990) homology searches against the full non-redundant protein sequence databank UniProt (Uniprot Consortium, 2014) and against the well-annotated model organisms Escherichia coli K-12 and Bacillus subtilis 168 (Vallenet et al., 2013), enzymatic classifications based on COG (Tatusov et al., 2003), InterPro (Mitchell et al., 2015), FIGFam (Meyer et al., 2009) and PRIAM (Claudel-Renard et al., 2003) profiles, and prediction of protein localization using the TMHMM (Sonnhammer et al., 1998), SignalP (Bendtsen et al., 2004) and PSORTb (Gardy et al., 2005) tools. Synteny maps (i.e. conservation of local gene order) were used to validate the annotation of genes located within conserved operons (Vallenet et al., 2009). Metabolic pathways were subsequently identified with the assistance of the integrated MicroCyc (Vallenet et al., 2009) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases (Kanehisa et al., 2014).
Results and discussion
Survey of 16S rRNA genes of Hyd24-12 in anaerobic digesters
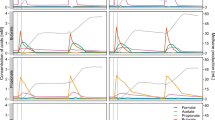
The survey of 22 full-scale mesophilic and 7 thermophilic anaerobic digesters from 17 Danish wastewater treatment plants over 3 years revealed that members of the Hyd24-12 lineage were stably present in most mesophilic but no thermophilic anaerobic digesters (Figure 1). In most mesophilic digesters, they were among the five most abundant bacterial OTUs and constituted around 1–3% and, in some cases, up to 8.2% of all sequenced bacterial reads (see Supplementary Figure S1). No 16S rRNA gene sequences from Hyd24-12 were detected in the incoming surplus sludge from the activated sludge treatment plants, which demonstrates that these bacteria were actively growing in the digesters. The other abundant bacterial phyla in the mesophilic digesters were Actinobacteria, Firmicutes, Chloroflexi, Synergistetes and Bacteroidetes (Figure 1). The best (LCA) classification is shown in Figure 1, but the lack of closely related organisms in the databases and a curated taxonomy hampers taxonomic classification for a number of the most abundant OTUs. In general, the abundance stability of these top genera was high, and that may be due to relatively similar growth conditions for all digesters: feed was primary sludge and surplus activated sludge, temperature in the interval 34–37 °C, pH 7.1–8.2 and total ammonium 0.57–1.1 g N/l (see Supplementary Table S1).
Heatmap of the 25 most abundant bacterial OTUs in mesophilic digesters at wastewater treatment plants along with their abundance in thermophilic digesters at 17 wastewater treatment plants. The OTU classified as belonging to the Hyd24-12 candidate phylum (purple) was detected exclusively in mesophilic reactors. Classification levels presented are phylum, class, order, family and genus and are separated by a semicolon. The field is empty where no classification at a given level could be provided. The abundance profiles show mean abundances for plants with more reactors (1–4 reactors at each WWTP) and 2–97 samples for each plant over 4 years (Supplementary Table S1). The OTUs are sorted on the basis of the mean abundance across the mesophilic samples.
In silico analysis of 16S rRNA gene sequences within Hyd24-12 of SILVA (Quast et al., 2013) from other surveys confirmed that members of Hyd24-12 are widespread in anaerobic environments. The sequences originate from 48 separate studies, with engineered systems such as anaerobic bioreactors accounting for 10 studies, and natural systems such as marine sediments, microbial mats in hydrogen, methane-rich waters and mud volcanoes accounting for 38 studies (see Supplementary References). Furthermore, the 48 studies show that members of Hyd24-12 are globally dispersed (Supplementary Figure S3 and Supplementary Table S2) and are potentially important in many microbial ecosystems besides ADs (Mills et al., 2005; Harris et al., 2012). Some of the surveys of full-scale anaerobic digesters detected some Hyd24-12 sequences (e.g., De Vrieze et al., 2015), while others did not (Sundberg et al., 2013). This was likely because they used the RDP database, where Hyd24-12 sequences are classified as ‘unclassified bacteria’.
Recovering genomic information from Hyd24-12
Three full-scale anaerobic digesters were sampled for metagenomic analyses. To ensure differential abundance of microorganisms needed to bin genomes based on coverage profiles (Albertsen et al., 2013), biomass samples were either taken from the sludge and foam layer of reactors or from the same reactor weeks apart. More than 50 gigabases of metagenomic data were generated, and population genomes were recovered by differential coverage binning (Albertsen et al., 2013) from each of the three plants (Table 2). The three population genomes were ~2.2 Mbp with a GC content of ~64%, and the completeness of the genomes were estimated by CheckM (Parks et al., 2015) to be between 86% and 91% with less than 2.2% estimated contamination (Table 2). However, the level of completeness may be underestimated, given that members of the Hyd24-12 are distantly related to other characterized organisms, and the genes used in the marker sets might be too divergent or simply not present (Rinke et al., 2013; Brown et al., 2015; Sekiguchi et al., 2015). The three genomes each contained a single rRNA operon and shared identical 16S rRNA gene sequences, which suggests that they belong to the same species (Yarza et al., 2014). The JSpecies program determined that these three genomes shared between 99.8% and 99.9% average nucleotide identity (ANIb), supporting the close taxonomic relationship observed from the 16S rRNA gene analysis (Kim et al., 2014). In order to further evaluate the similarity between the strains, the raw metagenome reads from each digester were mapped to the assembled Hyd24-12 genomes obtained from the other two digesters. Complete coverage of all genomes with the metagenome reads from the other digester revealed that the Hyd24-12 genomes were almost identical. This also indicates, along with the high ANIb, that the genomes are more complete than estimated in Table 2 by CheckM. Indeed, the data suggested that the three strains might actually be variants of the same strain with single-nucleotide polymorphisms only. This is very interesting as the digesters were from different parts of Denmark without any exchange of sludge or feed. This could indicate that they are highly adapted to the specific AD environment in this type of mesophilic digesters.
Hyd24-12 phylogeny, FISH probe design and morphology
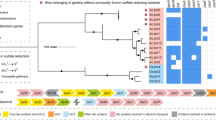
The 16S rRNA genes obtained have a sequence identity of 86% with the original clone Hyd24-12 sequence (AJ535232) (Knittel et al., 2003) and classify to the Hyd24-12-lineage (Figure 2a). Additional phylogenetic analyses, based on the genome sequence, placed the Hyd24-12 genomes within the Fibrobacteres-Chlorobi-Bacteroidetes superphylum (Figure 3). The Hyd24-12 genomes are distantly related to all currently available genomes, supporting its status as a novel phylum.
Phylogenetic analysis and design of FISH probes. (a) Maximum-likelihood (PhyML) 16S rRNA gene phylogenetic tree of target groups and selected related sequences (all >1200 bp). Phylogenetic classification is taken from the MiDAS database (Release 1.21), which is a version of the SILVA database (Release 119 NR99) (Quast et al., 2013) curated for activated sludge sequences (McIlroy et al., 2015). Clades of the Hyd24-12 lineage are shown in brackets. Probes covering clades are shown in red. The scale bar represents substitutions per nucleotide base. Bootstrap values from 100 re-samplings are indicated for branches when >50%. (b) Composite FISH micrographs of the B-1AC clade members in the Randers anaerobic digester sludge. B-1AC cells appear red (Hyd24-12_659, Cy3), other bacterial cells appear blue (EUBmix probe set (Amann et al., 1990; Daims et al., 1999), Cy5) and archaeal cells appear green (ARCH915 (Stahl and Amann, 1991), FLUOS).
Phylogenetic position of the Hyd24-12 genomes in the reference genome tree generated by CheckM. The CheckM tree is inferred from the concatenation of 43 conserved marker genes and incorporates 2052 finished and 3604 draft genomes from the IMG database (Parks et al., 2015).
Several FISH probes were designed to target different clades within the phylum. In the MiDAS taxonomy (v. 1.21) (McIlroy et al., 2015), a version of the SILVA taxonomy (Quast et al., 2013) that is curated for activated sludge-related organisms, the Hyd24-12 lineage is delineated into four clades, designated B-1AC, zEL51, Hyd-32 and B9.18. The Hyd24-12_468 and Hyd24-12_659 probes were designed to cover the B-1AC clade, which includes the Hyd24-12 genome sequences obtained in this study (Figure 2). The former probe covers almost all the B-1AC sequences, with the closest non-target sequence match having three internal base mismatches. The Hyd24-12_659 probe is less specific, having one perfectly matched non-target sequence and several with mismatches not covered by the competitor probes. Overlap in the coverage of these two probes, labelled with different fluorochromes, allows greater confidence in their specificity. A suitable probe to cover the entire Hyd24-12 lineage was not found. However, the Hyd24-12_731 and Hyd24-12_842 probes provide good coverage of the other sequences in the phylum (see Table 1). As sequences covered by these additional probes were not detected in the full-scale anaerobic digesters studied here, optimization and assessment of these probes were not pursued.
When applied to several full-scale anaerobic digester sludge samples, the Hyd24-12_468 and Hyd24-12_659 probes hybridized to small rods, approx. 2 × 0.4 μm in size, dispersed through the flocs (see Figure 2b). Good overlap was observed for these probes, supporting their specificity. Of the two probes, a much higher signal was observed for the Hyd24-12_659 probe. There was no observed overlap between the signal of two Hyd24-12 probes and the universal bacterial EUBmix probe set (see Figure 2b), which is supported by the absence of the target site for the probes of the latter in the Hyd24-12 sequences. Quantitative FISH was very difficult to carry out in the digesters due to high levels of background fluorescence. Instead, abundance estimates were carried out for the domains Bacteria, Archaea, Eukarya, and the Hyd24-12 lineage, based on read mapping from the PCR free metagenomes to the 16S rRNA genes of the MiDAS database. It showed that Archaea constituted 4–9% of the reads in sludge samples and 7–13% in foam samples. Reads from the Hyd24-12 lineage constituted 0.4–3.5% in the different samples (Supplementary Table S3).
Morphology and motility
The rod shape morphology of B-1AC clade organisms observed by FISH is supported by mreBCD and mrdAB operons in the Hyd24-12 genomes (see Supplementary Data 2). These operons encode proteins involved in the formation of membrane-bound actin filaments, which are essential for the biogenesis of rod-shape stabilizing peptidoglycans along the lateral cell wall of rod-shaped bacteria (Kruse et al., 2003, 2005; Osborn and Rothfield, 2007; Bendezú and de Boer, 2008).
The cell envelope characteristics of genome-sequenced bacteria can be determined based on PFAM protein families that are substantially enriched or depleted in archetypical monoderm lineages relative to archetypical diderm lineages (Albertsen et al., 2013). A search for such protein families in the Hyd24-12 genomes revealed an archetypical diderm cell envelope with lipopolysaccharides (see Supplementary Figure S2).
None of the Hyd24-12 genomes encode any flagella-related proteins, suggesting limited motility. However, genes associated with type IV pili were identified using the PilFind algorithm (see Supplementary Data 2) (Imam et al., 2011). These pili enable the bacteria to generate surface-associated twitching motility. This allows them to move effectively through environments that contain shear-thinning viscoelastic fluids, such as the extracellular polymeric substances of biofilms (Conrad et al., 2011; Jin et al., 2011). In addition to motility, type IV pili play a role in the attachment to living and non-living surfaces, including those of other bacteria (Giltner et al., 2012).
No genes associated with spore formation were detected in the Hyd24-12 genomes. This suggests that the Hyd24-12 genomes investigated represent non-sporulating bacteria.
Energy metabolism
The three genomes do not contain any genes for respiration with oxygen, nitrate/nitrite or Fe(III) and seem primarily to have a fermentative metabolism. However, the genomes indicate that the organisms may be able to use elemental sulphur as an electron acceptor, see below. The Hyd24-12 genomes encode a complete glycolysis pathway, along with the non-oxidative branch of the pentose phosphate pathway (Figure 4 and see Supplementary Data 2). This allows Hyd24-12 to potentially catabolize a wide range of hexoses and pentoses to pyruvate, thereby providing the cell with energy in the form of ATP and reducing equivalents in the form of NADH (Stincone et al., 2014). The sugars are probably obtained from the environment through a major facilitator superfamily transporter at the expense of the proton motive force (Madej, 2014; Wisedchaisri et al., 2014). The transporter does not share similarity (>30%) with any experimentally validated transporters, and it is therefore impossible to infer a specific substrate preference. It is known that primary sludge and activated sludge fed into the digesters contain many different polysaccharides (Raunkjaer et al., 1994; Frølund et al., 1996). No genes encoding for extracellular glycosylases were identified, which might indicate that Hyd24-12 is reliant on the hydrolytic action of other organisms present within the anaerobic digesters.
Metabolic model of Hyd24-12 species in mesophilic anaerobic digesters, based on the annotated genome sequences. Selected metabolic pathways important for the bacteria in the anaerobic digestion process are highlighted. Numbers correspond to annotated genes in Supplementary Data 1.
Hyd24-12 encodes for the complete pathway for glycogen biosynthesis and catabolism (Figure 4 and see Supplementary Data 2) (Preiss et al., 1983; Wilson et al., 2010). Hence, glycogen may serve as a carbon and energy storage which can be utilized to mitigate fluctuations in substrate availability. The Hyd24-12 genomes did not encode for pathways for other storage compounds such as trehalose or polyhydroxyalkanoates.
There are limited catabolic options for the pyruvate formed, for example, by glycolysis. The tri-carboxylic acid cycle of Hyd24-12 is incomplete (8 of 10 key enzymes are missing) and probably non-functional. However, pyruvate can be converted into acetyl-CoA by a pyruvate ferredoxin oxidoreductase, providing additional reducing equivalents in the form of reduced ferredoxin (Figure 4 and see Supplementary Data 2) (Menon and Ragsdale, 1997). Acetyl-CoA can then be converted into acetate by the action of phosphate acetyltransferase and acetate kinase, thus providing the bacterium with additional ATP (Latimer and Ferry, 1993; Mai and Adams, 1996).
All three Hyd24-12 genomes also encode for two aldehyde ferredoxin oxidoreductases (Figure 4 and see Supplementary Data 2). These may be used to oxidize formaldehyde and acetaldehyde to formate and acetate, respectively, providing the cell with energy in the form of additional reduced ferredoxin (Mukund and Adams, 1991). However, the enzyme may also be used in the reverse reaction to regenerate oxidized ferredoxin. The presence of a membrane-embedded, energy-conserving hydrogenase allows the cell to establish a proton motive force, based on the energy-rich reduced ferredoxin, which reduces H+ to H2 in the process (Strittmatter et al., 2009). The energy stored in the proton motive force may then be harvested through an ATP synthase to yield ATP.
High concentrations of H2 inhibit glycolysis and acidogenesis due to thermodynamic considerations (Huang et al., 2015). Hyd24-12 therefore needs a way to remove excess H2. This can be achieved by syntrophic association with other microorganisms, or internally by the action of a cytosolic hydrogenase, which couples the oxidation of H2 with the reduction of NAD+ (Figure 4 and see Supplementary Data 2). Alternatively, Hyd24-12 may employ a sulfhydrogenase to couple the oxidation of H2 to H+ with the reduction of elemental sulphur (S0) or polysulphide to hydrogen sulphide (H2S) as is seen for Pyrococcus furiosus (Mukund and Adams, 1991). The genomes do not indicate a potential for sulphate reduction. Elemental sulphur is continuously produced in the digesters because activated sludge fed into the digesters contains oxidized iron (Fe(III)), which in the presence of sulphide produces S0 and black iron sulphide (FeS) (Rasmussen and Nielsen, 1996; Nielsen et al., 2005; Omri et al., 2011). Sulphide is a normal compound in digesters and is produced from amino acids and reduction of sulphate. Notably, other studies have also detected members of the Hyd24-12 phylum in sulphur-rich environments such as hydrothermal vents, sulphur-rich springs and sediments (Elshahed et al., 2003; Schauer et al., 2011; Pjevac et al., 2014). Thus, Hyd24-12 related organisms potentially play a role in sulphur transformations in digesters and other environments. Such a role requires further investigation.
The Hyd24-12 genomes do not contain the genes required for fatty acid β-oxidation or for the catabolism of amino acids. Sugars are therefore considered the primary energy source of the Hyd24-12 in anaerobic digesters.
Whereas Hyd24-12 is able to take up carbon in the form of amino acids, carbohydrates, etc., it is unable to carry out fixation of CO2 as such genes are missing.
Amino acid and nitrogen metabolism
Based on the genome annotations, Hyd24-12 is only predicted to be able to synthesize few amino acids (glycine, serine, cysteine, threonine, asparagine, aspartate, glutamate and glutamine). Accordingly, Hyd24-12 might rely on amino acids present within the environment. As most amino acids are found as proteins, which cannot be taken up by the bacterium, Hyd24-12 needs a way to degrade these polymers, and this is achieved by the action of multiple extracellular proteases encoded in the genome, which are likely secreted in a Sec- or Tat-dependent mechanism (Natale et al., 2008) (see Supplementary Data 2). The cells may subsequently import the amino acids using ABC-transporters encoded in the genome. Owing to the lack of experimentally validated homologues from closely related species, it is not possible to predict the substrate specificity of these transporters. A reduced capacity of microorganisms for synthesizing amino acids is known from strict symbionts and, recently, also from a number of candidate phyla with very small genomes (<1 Mbp) (Brown et al., 2015). However, the relatively large size of the Hyd24-12 genomes (~2.2 Mbp) and their dispersed growth in the anaerobic sludge suggest that they are not strict symbionts.
Hyd24-12 does not have the necessary pathways for fixation of nitrogen. The nitrogen metabolism of Hyd24-12 is generally limited. Amino acids may also represent a source of nitrogen. However, nitrogen can also be obtained from ammonium assimilation via the glutamine synthetase/glutamate synthase pathways (Bravo and Mora, 1988).
Oxidative stress protection
The three Hyd24-12 genomes each contains a gene cluster encoding for a superoxide reductase, nitric oxide reductase and ferroxidase. These genes are probably involved in resistance against oxidative stress, and may allow the bacteria to survive in the presence of oxygen. However, 16S rRNA gene sequences from Hyd24-12 have only been observed in oxygen-depleted environments.
Ecological significance and concluding remarks
This study applied metagenomic sequencing to obtain genomes from the candidate phylum Hyd24-12 and provides the first morphological and physiological information for the lineage. Members of the phylum were shown to be very abundant and stably present in mesophilic anaerobic digesters, occasionally accounting for the most abundant OTU in the samples, but absent in thermophilic reactors. This indicates that they are likely to play a substantial role in the ecology of mesophilic AD systems at wastewater treatment plants fed with primary sludge and surplus activated sludge. Metabolic reconstruction based on the genomic information showed that members of Hyd24-12 are likely to be fermenters relying on simple sugars. In addition, they may also use elemental sulphur as an electron acceptor, thus forming part of the microbial cycling of sulphur in anaerobic systems and partly responsible for production of hydrogen sulphide. Sulphide is unwanted in the biogas due to toxicity and corrosion (Syed et al., 2006), but will also provide more elemental sulphur by reacting with incoming Fe(III). In that case, members of Hyd24-12 may compete with the methanogens for organics. An in silico investigation of environmental 16S rRNA gene surveys suggests that members of the phylum are present in anaerobic environments, often associated with sulphurous compounds and methane production, such as sediment mats and anaerobic bioreactors. The fact that the genomes are auxotrophic for several amino acids and lacking putative secreted glycoside hydrolases also indicates a strict reliance on other organisms for nutrients. The genomes generated in this study provide the foundation for future detailed analyses of members of the phylum, such as metatranscriptomics and metaproteomics. The design of FISH probes for the phylum also revealed their morphology and spatial arrangement in anaerobic digesters and will also facilitate future in situ investigations of the phylum in digesters and other environments.
Phylogenetic and genomic analyses of the three Hyd24-12 genomes classified them as a single species within a novel phylum located within the Fibrobacteres-Chlorobi-Bacteroidetes superphylum.
We propose the following taxonomic names for the novel genus and species of Hyd24-12:
-
‘Candidatus Fermentibacter’ gen. nov.
-
‘Candidatus Fermentibacter daniensis’ gen. et sp. nov.
Based on this, we propose the following names for the phylum, class, order, and family:
-
‘Candidatus Fermentibacteria’ phyl. nov.
-
‘Candidatus Fermentibacteria’ classis nov.
-
‘Candidatus Fermentibacterales’ ord. nov.
-
‘Candidatus Fermentibacteraceae’ fam. nov.
Etymology
Fermentibacter (Fer.men.ti.bac'ter. M.L. n. ferment -um to ferment, Gr. dim. n. bakterion a small rod, M.L. neut. n. Fermentibacter a small fermenting rod-shaped bacterium). Fermentibacter daniensis (da.ni.ensis. M.L. fem. adj. daniensis, pertaining to Dania, the Medieval Latin name for the country of Denmark, where the species was first discovered).
References
Albertsen M, Hugenholtz P, Skarshewski A, Nielsen KL, Tyson GW, Nielsen PH . (2013). Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat Biotechnol 31: 533–538.
Albertsen M, Karst SM, Ziegler AS, Kirkegaard RH, Nielsen PH . (2015). Back to basics—the influence of DNA extraction and primer choice on phylogenetic analysis of activated sludge communities. PLoS One 10: e0132783.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA . (1990). Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56: 1919–1925.
Angenent LT, Karim K, Al-Dahhan MH, Wrenn BA, Domíguez-Espinosa R . (2004). Production of bioenergy and biochemicals from industrial and agricultural wastewater. Trends Biotechnol 22: 477–485.
Bendezú FO, de Boer PAJ . (2008). Conditional lethality, division defects, membrane involution, and endocytosis in mre and mrd shape mutants of Escherichia coli. J Bacteriol 190: 1792–1811.
Bendtsen JD, Nielsen H, von Heijne G, Brunak S . (2004). Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340: 783–795.
Bland C, Ramsey TL, Sabree F, Lowe M, Brown K, Kyrpides NC et al. (2007). CRISPR recognition tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinformatics 8: 209.
Bolger AM, Lohse M, Usadel B . (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30: 2114–2120.
Bravo A, Mora J . (1988). Ammonium assimilation in Rhizobium phaseoli by the glutamine synthetase-glutamate synthase pathway. J Bacteriol 170: 980–984.
Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A et al. (2015). Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523: 208–211.
Chouari R, Le Paslier D, Dauga C, Daegelen P, Weissenbach J, Sghir A . (2005). Novel major bacterial candidate division within a municipal anaerobic sludge digester. Appl Environ Microbiol 71: 2145–2153.
Claudel-Renard C, Chevalet C, Faraut T, Kahn D . (2003). Enzyme-specific profiles for genome annotation: PRIAM. Nucleic Acids Res 31: 6633–6639.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Conrad JC, Gibiansky ML, Jin F, Gordon VD, Motto DA, Mathewson MA et al. (2011). Flagella and pili-mediated near-surface single-cell motility mechanisms in P. aeruginosa. Biophys J 100: 1608–1616.
Daims H, Brühl A, Amann R, Schleifer KH, Wagner M . (1999). The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22: 434–444.
Daims H, Stoecker K, Wagner M . (2005). Fluorescence in situ hybridization for the detection of prokaryotes. Mol Microb Ecol 213: 239.
De Vrieze J, Saunders AM, He Y, Fang J, Nielsen PH, Verstraete W et al. (2015). Ammonia and temperature determine potential clustering in the anaerobic digestion microbiome. Water Res 75: 312–323.
Dinis JM, Barton DE, Ghadiri J, Surendar D, Reddy K, Velasquez F et al. (2011). In search of an uncultured human-associated TM7 bacterium in the environment. PLoS One 6: 1–8.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Elshahed MS, Senko JM, Najar FZ, Kenton SM, Roe BA, Dewers TA et al. (2003). Bacterial diversity and sulfur cycling in a mesophilic sulfide-rich spring. Appl Environ Microbiol 69: 5609–5621.
Frølund B, Palmgren R, Keiding K, Nielsen PH . (1996). Extraction of extracellular polymers from activated sludge using a cation exchange resin. Water Res 30: 1749–1758.
Fuchs BM, Glockner FO, Wulf J, Amann R . (2000). Unlabeled helper oligonucleotides increase the in situ accessibility to 16S rRNA of fluorescently labeled oligonucleotide probes. Appl Environ Microbiol 66: 3603–3607.
Fuhrman JA, Cram JA, Needham DM . (2015). Marine microbial community dynamics and their ecological interpretation. Nat Rev Microbiol 13: 133–146.
Gardy JL, Laird MR, Chen F, Rey S, Walsh CJ, Ester M et al. (2005). PSORTb v.2.0: expanded prediction of bacterial protein subcellular localization and insights gained from comparative proteome analysis. Bioinformatics 21: 617–623.
Giltner CL, Nguyen Y, Burrows LL . (2012). Type IV pilin proteins: versatile molecular modules. Microbiol Mol Biol Rev 76: 740–772.
Guermazi S, Daegelen P, Dauga C, Rivière D, Bouchez T, Godon JJ et al. (2008). Discovery and characterization of a new bacterial candidate division by an anaerobic sludge digester metagenomic approach. Environ Microbiol 10: 2111–2123.
Harris JK, Gregory Caporaso J, Walker JJ, Spear JR, Gold NJ, Robertson CE et al. (2012). Phylogenetic stratigraphy in the Guerrero Negro hypersaline microbial mat. ISME J 7: 50–60.
Huang W, Wang Z, Zhou Y, Ng WJ . (2015). The role of hydrogenotrophic methanogens in an acidogenic reactor. Chemosphere 140: 40–46.
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ . (2010). Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11: 119.
Imam S, Chen Z, Roos DS, Pohlschröder M . (2011). Identification of surprisingly diverse type IV pili, across a broad range of Gram-positive bacteria. PLoS One 6: e28919.
Jin F, Conrad JC, Gibiansky ML, Wong GCL . (2011). Bacteria use type-IV pili to slingshot on surfaces. Proc Natl Acad Sci USA 108: 12617–12622.
Kaeberlein T, Lewis K, Epstein SS . (2002). Isolating ‘uncultivable’ microorganisms in pure culture in a simulated natural environment. Science 296: 1127–1129.
Kanehisa M, Goto S, Sato Y, Kawashima M, Furumichi M, Tanabe M . (2014). Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res 42: 199–205.
Kim M, Oh H-SS, Park S-CC, Chun J . (2014). Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64: 346–351.
Knittel K, Boetius A, Lemke A, Eilers H, Lochte K, Pfannkuche O et al. (2003). Activity, distribution, and diversity of sulfate reducers and other bacteria in sediments above gas hydrate (Cascadia Margin, Oregon). Geomicrobiol J 20: 269–294.
Koch H, Galushko A, Albertsen M, Schintlmeister A, Gruber-Dorninger C, Lucker S et al. (2014). Growth of nitrite-oxidizing bacteria by aerobic hydrogen oxidation. Science 345: 1052–1054.
Kruse T, Bork-Jensen J, Gerdes K . (2005). The morphogenetic MreBCD proteins of Escherichia coli form an essential membrane-bound complex. Mol Microbiol 55: 78–89.
Kruse T, Møller-Jensen J, Løbner-Olesen A, Gerdes K . (2003). Dysfunctional MreB inhibits chromosome segregation in Escherichia coli. EMBO J 22: 5283–5292.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds). Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons: West Sussex.
Latimer MT, Ferry JG . (1993). Cloning, sequence analysis, and hyperexpression of the genes encoding phosphotransacetylase and acetate kinase from Methanosarcina thermophila. J Bacteriol 175: 6822–6829.
Limam RD, Chouari R, Mazéas L, Wu T, Di, Li T, Grossin-Debattista J et al. (2014). Members of the uncultured bacterial candidate division WWE1 are implicated in anaerobic digestion of cellulose. MicrobiologyOpen 3: 157–167.
Love MI, Huber W, Anders S . (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15: 550.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar A et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Madej MG . (2014). Function, structure and evolution of the major facilitator superfamily: the LacY manifesto. Adv Biol 2014: 20.
Magoč T, Salzberg SL . (2011). FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27: 2957–2963.
Mai X, Adams MW . (1996). Purification and characterization of two reversible and ADP-dependent acetyl coenzyme A synthetases from the hyperthermophilic archaeon Pyrococcus furiosus. J Bacteriol 178: 5897–5903.
Manz W, Amann R, Ludwig W, Wagner M, Schleifer K-H . (1992). Phylogenetic oligodeoxynucleotide probes for the major subclasses of Proteobacteria: problems and solutions. Syst Appl Microbiol 15: 593–600.
McIlroy SJ, Saunders AM, Albertsen M, Nierychlo M, McIlroy B, Hansen AA et al. (2015). MiDAS: the field guide to the microbes of activated sludge. Database 2015: bav062.
McIlroy SJ, Tillett D, Petrovski S, Seviour RJ . (2011). Non-target sites with single nucleotide insertions or deletions are frequently found in 16S rRNA sequences and can lead to false positives in fluorescence in situ hybridization (FISH). Environ Microbiol 13: 33–47.
McMurdie PJ, Holmes S . (2013). phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8: e61217.
Menon S, Ragsdale SW . (1997). Mechanism of the clostridium thermoaceticum pyruvate: ferredoxin oxidoreductase: evidence for the common catalytic intermediacy of the hydroxyethylthiamine pyropyrosphate radical. Biochemistry 36: 8484–8494.
Meyer F, Overbeek R, Rodriguez A . (2009). FIGfams: yet another set of protein families. Nucleic Acids Res 37: 6643–6654.
Mills HJ, Martinez RJ, Story S, Sobecky PA . (2005). Characterization of microbial community structure in Gulf of Mexico gas hydrates: comparative analysis of DNA- and RNA-derived clone libraries. Appl Environ Microbiol 71: 3235–3247.
Mitchell A, Chang H-YH-Y, Daugherty L, Fraser M, Hunter S, Lopez R et al. (2015). The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43: D213–D221.
Mukund S, Adams MWW . (1991). The novel tungsten-iron-sulfur protein of the hyperthermophilic archaebacterium, Pyrococcus furiosus, is an aldehyde ferredoxin oxidoreductase: evidence for its participation in a unique glycolytic pathway. J Biol Chem 266: 14208–14216.
Muyzer G, De Waal EC, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
Natale P, Brüser T, Driessen AJM . (2008). Sec- and Tat-mediated protein secretion across the bacterial cytoplasmic membrane—distinct translocases and mechanisms. Biochim Biophys Acta 1778: 1735–1756.
Nielsen AH, Lens P, Vollertsen J, Hvitved-Jacobsen T . (2005). Sulfide-iron interactions in domestic wastewater from a gravity sewer. Water Res 39: 2747–2755.
Nobu MK, Narihiro T, Rinke C, Kamagata Y, Tringe SG, Woyke T et al. (2015). Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J 9: 1710–1722.
Omri I, Bouallagui H, Aouidi F, Godon JJ, Hamdi M . (2011). H2S gas biological removal efficiency and bacterial community diversity in biofilter treating wastewater odor. Bioresour Technol 102: 10202–10209.
Osborn MJ, Rothfield L . (2007). Cell shape determination in Escherichia coli. Curr Opin Microbiol 10: 606–610.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW . (2015). CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25: 1043–1055.
Pelletier E, Kreimeyer A, Bocs S, Rouy Z, Gyapay G, Chouari R et al. (2008). ‘Candidatus Cloacamonas acidaminovorans’: genome sequence reconstruction provides a first glimpse of a new bacterial division. J Bacteriol 190: 2572–2579.
Pjevac P, Kamyshny A, Dyksma S, Mussmann M . (2014). Microbial consumption of zero-valence sulfur in marine benthic habitats. Environ Microbiol 16: 3416–3430.
Preiss J, Yung S, Baecker PA . (1983). Regulation of bacterial glycogen synthesis. Mol Cell Biochem 57: 61–80.
Pruesse E, Peplies J, Glöckner FO . (2012). SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28: 1823–1829.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41: 590–596.
R Core Team. (2016) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, Available from https://www.R-project.org/.
Rasmussen H, Nielsen PH . (1996). Iron reduction in activated sludge measured with different extraction techniques. Water Res 30: 551–558.
Raunkjaer K, Hvitved-Jacobsen T, Nielsen PH . (1994). Measurement of pools of protein, carbohydrate and lipid in domestic wastewater. Water Res 28: 251–262.
Richter M, Rosselló-Móra R . (2009). Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106: 19126–19131.
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng J-F et al. (2013). Insights into the phylogeny and coding potential of microbial dark matter. Nature 499: 431–437.
Schauer R, Røy H, Augustin N, Gennerich H-H, Peters M, Wenzhoefer F et al. (2011). Bacterial sulfur cycling shapes microbial communities in surface sediments of an ultramafic hydrothermal vent field. Environ Microbiol 13: 2633–2648.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Sekiguchi Y, Ohashi A, Parks DH, Yamauchi T, Tyson GW, Hugenholtz P . (2015). First genomic insights into members of a candidate bacterial phylum responsible for wastewater bulking. Peer J 3: e740.
Sonnhammer EL, von Heijne G, Krogh A . (1998). A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6: 175–182.
Stahl DA, Amann R . (1991). Development and application of nucleic acid probes in bacterial systematics. In Stackebrandt E, Goodfellow M (eds) Nucleic Acid Techniques in Bacterial Systematics. John Wiley & Sons Ltd. Chichester: England, pp 205–248.
Stincone A, Prigione A, Cramer T, Wamelink MMC, Campbell K, Cheung E et al. (2014). The return of metabolism: biochemistry and physiology of the pentose phosphate pathway. Biol Rev Camb Philos Soc 90: 927–963.
Stoecker K, Dorninger C, Daims H, Wagner M . (2010). Double labeling of oligonucleotide probes for fluorescence in situ hybridization (DOPE-FISH) improves signal intensity and increases rRNA accessibility. Appl Environ Microbiol 76: 922–926.
Strittmatter AW, Liesegang H, Rabus R, Decker I, Amann J, Andres S et al. (2009). Genome sequence of Desulfobacterium autotrophicum HRM2, a marine sulfate reducer oxidizing organic carbon completely to carbon dioxide. Environ Microbiol 11: 1038–1055.
Sundberg C, Al-Soud Wa, Larsson M, Alm E, Yekta SS, Svensson BH et al. (2013). 454 Pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85: 612–626.
Syed M, Soreanu G, Falletta P, Beland M . (2006). Removal of hydrogen sulfide from gas streams using biological processes—a review. Can Biosyst Eng 48: 2.1–2.14.
Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin E V et al. (2003). The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4: 41.
Uniprot Consortium. (2014). Activities at the Universal Protein Resource (UniProt). Nucleic Acids Res 42: D191–D198.
Vallenet D, Belda E, Calteau A, Cruveiller S, Engelen S, Lajus A et al. (2013). MicroScope—an integrated microbial resource for the curation and comparative analysis of genomic and metabolic data. Nucleic Acids Res 41: D636–D647.
Vallenet D, Engelen S, Mornico D, Cruveiller S, Fleury L, Lajus A et al. (2009). MicroScope: a platform for microbial genome annotation and comparative genomics. Database (Oxford) 2009: bap021.
Wallner G, Amann R, Beisker W . (1993). Optimizing fluorescent in situ hybridization with rRNA targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry 14: 136–143.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Weiland P . (2010). Biogas production: current state and perspectives. Appl Microbiol Biotechnol 85: 849–860.
Werner JJ, Knights D, Garcia ML, Scalfone NB, Smith S, Yarasheski K et al. (2011). Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Natl Acad Sci USA 108: 4158–4163.
Wickham H . (2007). Reshaping data with the {reshape} package. J Stat Softw 21: 1–20.
Wilson WA, Roach PJ, Montero M, Baroja-Fernández E, Muñoz FJ, Eydallin G et al. (2010). Regulation of glycogen metabolism in yeast and bacteria. FEMS Microbiol Rev 34: 952–985.
Wisedchaisri G, Park M-S, Iadanza MG, Zheng H, Gonen T . (2014). Proton-coupled sugar transport in the prototypical major facilitator superfamily protein XylE. Nat Commun 5: 4521.
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer K-H et al. (2014). Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12: 635–645.
Yilmaz LS, Parnerkar S, Noguera DR . (2011). MathFISH, a web tool that uses thermodynamics-based mathematical models for in silico evaluation of oligonucleotide probes for fluorescence in situ hybridization. Appl Environ Microbiol 77: 1118–1122.
Acknowledgements
This study was supported by the Villum Foundation, the Grundfos Foundation, Aalborg University and Innovation Fund Denmark (NomiGas project).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Kirkegaard, R., Dueholm, M., McIlroy, S. et al. Genomic insights into members of the candidate phylum Hyd24-12 common in mesophilic anaerobic digesters. ISME J 10, 2352–2364 (2016). https://doi.org/10.1038/ismej.2016.43
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.43
This article is cited by
-
Anaerobic degradation of organic carbon supports uncultured microbial populations in estuarine sediments
Microbiome (2023)
-
Metabolic dependencies govern microbial syntrophies during methanogenesis in an anaerobic digestion ecosystem
Microbiome (2020)
-
Novel prosthecate bacteria from the candidate phylum Acetothermia
The ISME Journal (2018)
-
Metagenome, metatranscriptome, and metaproteome approaches unraveled compositions and functional relationships of microbial communities residing in biogas plants
Applied Microbiology and Biotechnology (2018)
-
Diversity of microbial carbohydrate-active enzymes in Danish anaerobic digesters fed with wastewater treatment sludge
Biotechnology for Biofuels (2017)